MDCK) Cells Grown Under Differentiation Conditions
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Cytochrome P450 Enzymes in Oxygenation of Prostaglandin Endoperoxides and Arachidonic Acid
Comprehensive Summaries of Uppsala Dissertations from the Faculty of Pharmacy 231 _____________________________ _____________________________ Cytochrome P450 Enzymes in Oxygenation of Prostaglandin Endoperoxides and Arachidonic Acid Cloning, Expression and Catalytic Properties of CYP4F8 and CYP4F21 BY JOHAN BYLUND ACTA UNIVERSITATIS UPSALIENSIS UPPSALA 2000 Dissertation for the Degree of Doctor of Philosophy (Faculty of Pharmacy) in Pharmaceutical Pharmacology presented at Uppsala University in 2000 ABSTRACT Bylund, J. 2000. Cytochrome P450 Enzymes in Oxygenation of Prostaglandin Endoperoxides and Arachidonic Acid: Cloning, Expression and Catalytic Properties of CYP4F8 and CYP4F21. Acta Universitatis Upsaliensis. Comprehensive Summaries of Uppsala Dissertations from Faculty of Pharmacy 231 50 pp. Uppsala. ISBN 91-554-4784-8. Cytochrome P450 (P450 or CYP) is an enzyme system involved in the oxygenation of a wide range of endogenous compounds as well as foreign chemicals and drugs. This thesis describes investigations of P450-catalyzed oxygenation of prostaglandins, linoleic and arachidonic acids. The formation of bisallylic hydroxy metabolites of linoleic and arachidonic acids was studied with human recombinant P450s and with human liver microsomes. Several P450 enzymes catalyzed the formation of bisallylic hydroxy metabolites. Inhibition studies and stereochemical analysis of metabolites suggest that the enzyme CYP1A2 may contribute to the biosynthesis of bisallylic hydroxy fatty acid metabolites in adult human liver microsomes. 19R-Hydroxy-PGE and 20-hydroxy-PGE are major components of human and ovine semen, respectively. They are formed in the seminal vesicles, but the mechanism of their biosynthesis is unknown. Reverse transcription-polymerase chain reaction using degenerate primers for mammalian CYP4 family genes, revealed expression of two novel P450 genes in human and ovine seminal vesicles. -

EXTENDED CARRIER SCREENING Peace of Mind for Planned Pregnancies
Focusing on Personalised Medicine EXTENDED CARRIER SCREENING Peace of Mind for Planned Pregnancies Extended carrier screening is an important tool for prospective parents to help them determine their risk of having a child affected with a heritable disease. In many cases, parents aren’t aware they are carriers and have no family history due to the rarity of some diseases in the general population. What is covered by the screening? Genomics For Life offers a comprehensive Extended Carrier Screening test, providing prospective parents with the information they require when planning their pregnancy. Extended Carrier Screening has been shown to detect carriers who would not have been considered candidates for traditional risk- based screening. With a simple mouth swab collection, we are able to test for over 419 genes associated with inherited diseases, including Fragile X Syndrome, Cystic Fibrosis and Spinal Muscular Atrophy. The assay has been developed in conjunction with clinical molecular geneticists, and includes genes listed in the NIH Genetic Test Registry. For a list of genes and disorders covered, please see the reverse of this brochure. If your gene of interest is not covered on our Extended Carrier Screening panel, please contact our friendly team to assist you in finding a gene test panel that suits your needs. Why have Extended Carrier Screening? Extended Carrier Screening prior to pregnancy enables couples to learn about their reproductive risk and consider a complete range of reproductive options, including whether or not to become pregnant, whether to use advanced reproductive technologies, such as preimplantation genetic diagnosis, or to use donor gametes. -

Transcriptomic Characterization of Fibrolamellar Hepatocellular
Transcriptomic characterization of fibrolamellar PNAS PLUS hepatocellular carcinoma Elana P. Simona, Catherine A. Freijeb, Benjamin A. Farbera,c, Gadi Lalazara, David G. Darcya,c, Joshua N. Honeymana,c, Rachel Chiaroni-Clarkea, Brian D. Dilld, Henrik Molinad, Umesh K. Bhanote, Michael P. La Quagliac, Brad R. Rosenbergb,f, and Sanford M. Simona,1 aLaboratory of Cellular Biophysics, The Rockefeller University, New York, NY 10065; bPresidential Fellows Laboratory, The Rockefeller University, New York, NY 10065; cDivision of Pediatric Surgery, Department of Surgery, Memorial Sloan-Kettering Cancer Center, New York, NY 10065; dProteomics Resource Center, The Rockefeller University, New York, NY 10065; ePathology Core Facility, Memorial Sloan-Kettering Cancer Center, New York, NY 10065; and fJohn C. Whitehead Presidential Fellows Program, The Rockefeller University, New York, NY 10065 Edited by Susan S. Taylor, University of California, San Diego, La Jolla, CA, and approved September 22, 2015 (received for review December 29, 2014) Fibrolamellar hepatocellular carcinoma (FLHCC) tumors all carry a exon of DNAJB1 and all but the first exon of PRKACA. This deletion of ∼400 kb in chromosome 19, resulting in a fusion of the produced a chimeric RNA transcript and a translated chimeric genes for the heat shock protein, DNAJ (Hsp40) homolog, subfam- protein that retains the full catalytic activity of wild-type PKA. ily B, member 1, DNAJB1, and the catalytic subunit of protein ki- This chimeric protein was found in 15 of 15 FLHCC patients nase A, PRKACA. The resulting chimeric transcript produces a (21) in the absence of any other recurrent mutations in the DNA fusion protein that retains kinase activity. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Synonymous Single Nucleotide Polymorphisms in Human Cytochrome
DMD Fast Forward. Published on February 9, 2009 as doi:10.1124/dmd.108.026047 DMD #26047 TITLE PAGE: A BIOINFORMATICS APPROACH FOR THE PHENOTYPE PREDICTION OF NON- SYNONYMOUS SINGLE NUCLEOTIDE POLYMORPHISMS IN HUMAN CYTOCHROME P450S LIN-LIN WANG, YONG LI, SHU-FENG ZHOU Department of Nutrition and Food Hygiene, School of Public Health, Peking University, Beijing 100191, P. R. China (LL Wang & Y Li) Discipline of Chinese Medicine, School of Health Sciences, RMIT University, Bundoora, Victoria 3083, Australia (LL Wang & SF Zhou). 1 Copyright 2009 by the American Society for Pharmacology and Experimental Therapeutics. DMD #26047 RUNNING TITLE PAGE: a) Running title: Prediction of phenotype of human CYPs. b) Author for correspondence: A/Prof. Shu-Feng Zhou, MD, PhD Discipline of Chinese Medicine, School of Health Sciences, RMIT University, WHO Collaborating Center for Traditional Medicine, Bundoora, Victoria 3083, Australia. Tel: + 61 3 9925 7794; fax: +61 3 9925 7178. Email: [email protected] c) Number of text pages: 21 Number of tables: 10 Number of figures: 2 Number of references: 40 Number of words in Abstract: 249 Number of words in Introduction: 749 Number of words in Discussion: 1459 d) Non-standard abbreviations: CYP, cytochrome P450; nsSNP, non-synonymous single nucleotide polymorphism. 2 DMD #26047 ABSTRACT Non-synonymous single nucleotide polymorphisms (nsSNPs) in coding regions that can lead to amino acid changes may cause alteration of protein function and account for susceptivity to disease. Identification of deleterious nsSNPs from tolerant nsSNPs is important for characterizing the genetic basis of human disease, assessing individual susceptibility to disease, understanding the pathogenesis of disease, identifying molecular targets for drug treatment and conducting individualized pharmacotherapy. -

Colorectal Cancer and Omega Hydroxylases
1 The differential expression of omega-3 and omega-6 fatty acid metabolising enzymes in colorectal cancer and its prognostic significance Abdo Alnabulsi1,2, Rebecca Swan1, Beatriz Cash2, Ayham Alnabulsi2, Graeme I Murray1 1Pathology, School of Medicine, Medical Sciences and Nutrition, University of Aberdeen, Aberdeen, AB25, 2ZD, UK. 2Vertebrate Antibodies, Zoology Building, Tillydrone Avenue, Aberdeen, AB24 2TZ, UK. Address correspondence to: Professor Graeme I Murray Email [email protected] Phone: +44(0)1224 553794 Fax: +44(0)1224 663002 Running title: omega hydroxylases and colorectal cancer 2 Abstract Background: Colorectal cancer is a common malignancy and one of the leading causes of cancer related deaths. The metabolism of omega fatty acids has been implicated in tumour growth and metastasis. Methods: This study has characterised the expression of omega fatty acid metabolising enzymes CYP4A11, CYP4F11, CYP4V2 and CYP4Z1 using monoclonal antibodies we have developed. Immunohistochemistry was performed on a tissue microarray containing 650 primary colorectal cancers, 285 lymph node metastasis and 50 normal colonic mucosa. Results: The differential expression of CYP4A11 and CYP4F11 showed a strong association with survival in both the whole patient cohort (HR=1.203, 95% CI=1.092-1.324, χ2=14.968, p=0.001) and in mismatch repair proficient tumours (HR=1.276, 95% CI=1.095-1.488, χ2=9.988, p=0.007). Multivariate analysis revealed that the differential expression of CYP4A11 and CYP4F11 was independently prognostic in both the whole patient cohort (p = 0.019) and in mismatch repair proficient tumours (p=0.046). Conclusions: A significant and independent association has been identified between overall survival and the differential expression of CYP4A11 and CYP4F11 in the whole patient cohort and in mismatch repair proficient tumours. -

Differential Expression of Hydroxyurea Transporters in Normal and Polycythemia Vera Hematopoietic Stem and Progenitor Cell Subpopulations
Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2021 Differential expression of hydroxyurea transporters in normal and polycythemia vera hematopoietic stem and progenitor cell subpopulations Tan, Ge ; Meier-Abt, Fabienne Abstract: Polycythemia vera (PV) is a myeloproliferative neoplasm marked by hyperproliferation of the myeloid lineages and the presence of an activating JAK2 mutation. Hydroxyurea (HU) is a standard treat- ment for high-risk patients with PV. Because disease-driving mechanisms are thought to arise in PV stem cells, effective treatments should target primarily the stem cell compartment. We tested for theantipro- liferative effect of patient treatment with HU in fluorescence-activated cell sorting-isolated hematopoietic stem/multipotent progenitor cells (HSC/MPPs) and more committed erythroid progenitors (common myeloid/megakaryocyte-erythrocyte progenitors [CMP/MEPs]) in PV using RNA-sequencing and gene set enrichment analysis. HU treatment led to significant downregulation of gene sets associated with cell proliferation in PV HSCs/MPPs, but not in PV CMP/MEPs. To explore the mechanism underlying this finding, we assessed for expression of solute carrier membrane transporters, which mediate trans- membrane movement of drugs such as HU into target cells. The active HU uptake transporter OCTN1 was upregulated in HSC/MPPs compared with CMP/MEPs of untreated patients with PV, and the HU diffusion facilitator urea transporter B (UTB) was downregulated in HSC/MPPs compared withCM- P/MEPs in all patient and control groups tested. These findings indicate a higher accumulation ofHU within PV HSC/MPPs compared with PV CMP/MEPs and provide an explanation for the differential effects of HU in HSC/MPPs and CMP/MEPs of patients with PV. -

Expression of SLC2A9 Isoforms in the Kidney and Their Localization in Polarized Epithelial Cells
Expression of SLC2A9 Isoforms in the Kidney and Their Localization in Polarized Epithelial Cells Toru Kimura1, Michi Takahashi1, Kunimasa Yan2, Hiroyuki Sakurai1* 1 Department of Pharmacology and Toxicology, Kyorin University School of Medicine, Mitaka, Tokyo, Japan, 2 Department of Pediatrics, Kyorin University School of Medicine, Mitaka, Tokyo, Japan Abstract Background: Many genome-wide association studies pointed out that SLC2A9 gene, which encodes a voltage-driven urate transporter, SLC2A9/GLUT9 (a.k.a. URATv1), as one of the most influential genes for serum urate levels. SLC2A9 is reported to encode two splice variants: SLC2A9-S (512 amino acids) and SLC2A9-L (540 amino acids), only difference being at their N- termini. We investigated isoform-specific localization of SLC2A9 in the human kidney and role of N-terminal amino acids in differential sorting in vitro. Methodology/Principal Findings: Isoform specific antibodies against SLC2A9 were developed and human kidney sections were stained. SLC2A9-S was expressed in the apical side of the collecting duct while SLC2A9-L was expressed in the basolateral side of the proximal tubule. GFP fused SLC2A9s were expressed in MDCK cells and intracellular localization was observed. SLC2A9-S was expressed at both apical and basolateral membranes, whereas SLC2A9-L was expressed only at the basolateral membrane. Although SLC2A9-L has a putative di-leucine motif at 33th and 34th leucine, deletion of the motif or replacement of leucine did not affect its subcellular localization. When up to 16 amino acids were removed from the N- terminal of SLC2A9-S or when up to 25 amino acids were removed from the N-terminal of SLC2A9-L, there was no change in their sorting. -

2.1 Drosophila Melanogaster
Overend, Gayle (2010) Drosophila as a model for the Anopheles Malpighian tubule. PhD thesis, University of Glasgow. http://theses.gla.ac.uk/1604/ Copyright and moral rights for this thesis are retained by the author A copy can be downloaded for personal non-commercial research or study, without prior permission or charge This thesis cannot be reproduced or quoted extensively from without first obtaining permission in writing from the Author The content must not be changed in any way or sold commercially in any format or medium without the formal permission of the Author When referring to this work, full bibliographic details including the author, title, awarding institution and date of the thesis must be given Glasgow Theses Service http://theses.gla.ac.uk/ [email protected] Drosophila as a model for the Anopheles Malpighian tubule A thesis submitted for the degree of Doctor of Philosophy at the University of Glasgow Gayle Overend Integrative and Systems Biology Faculty of Biomedical and Life Sciences University of Glasgow Glasgow G11 6NU UK August 2009 2 The research reported within this thesis is my own work except where otherwise stated, and has not been submitted for any other degree. Gayle Overend 3 Abstract The insect Malpighian tubule is involved in osmoregulation, detoxification and immune function, physiological processes which are essential for insect development and survival. As the Malpighian tubules contain many ion channels and transporters, they could be an effective tissue for targeting with novel pesticides to control populations of Diptera. Many of the insecticide compounds used to control insect pest species are no longer suited to their task, and so new means of control must be found. -

Taqman® Openarray® Pharmacogenomics (Pgx) Panel
PRODUCT OVERVIEW TaqMan® OpenArray® Pharmacogenomics (PGx) Panel TaqMan® OpenArray® Pharmacogenomics (PGx) Panel Genotyping analysis of drug metabolism enzymes and associated transport proteins The TaqMan® OpenArray® Trusted TaqMan® performance samples, studies, and labs. Assays Pharmacogenomics (PGx) Panel is Each TaqMan® DME Genotyping were selected for optimal relevance a powerful tool to help researchers Assay contains two allele-specific to current pharmacogenomics study human genetic variation probes and a primer pair to detect studies and organized for a simplified in relation to drug action and its the specific SNP target. Both the workflow. potential application to medical probes and primers uniquely align ® ® treatment. within the genome, enabling the The TaqMan OpenArray PGx TaqMan® genotyping technology to Panel provides valuable data for ® ® The TaqMan OpenArray PGx provide superior specificity. It is this the study of drug interactions in Panel was developed for quick and specificity that allows these assays several research areas. Targeted easy screening of known high- to detect targets residing in highly genes relate to areas of study such value target genes associated with homologous gene families that may as cardiovascular (CYP2D6, CYP2C19, drug metabolism enzymes and include pseudogenes. TaqMan® Drug NAT1, NAT2), analgesics (CYP2C9, associated transport proteins. Metabolism Enzyme Genotyping CYP2D6), rheumatology (CYP2C9, The panel consists of 158 drug Assays were developed using a high TPMT), neurology (CYP2C19, CYP2D6), metabolism enzyme (DME) assays level of bioinformatics and wet-lab and musculoskeletal (CYP2C19). A derived from the PharmaADME stringency. All assays have passed total of 29 genes are covered across Core Marker Set (Table 1). The performance tests involving 180 158 unique assays. -

ABCB6 Is a Porphyrin Transporter with a Novel Trafficking Signal That Is Conserved in Other ABC Transporters Yu Fukuda University of Tennessee Health Science Center
University of Tennessee Health Science Center UTHSC Digital Commons Theses and Dissertations (ETD) College of Graduate Health Sciences 12-2008 ABCB6 Is a Porphyrin Transporter with a Novel Trafficking Signal That Is Conserved in Other ABC Transporters Yu Fukuda University of Tennessee Health Science Center Follow this and additional works at: https://dc.uthsc.edu/dissertations Part of the Chemicals and Drugs Commons, and the Medical Sciences Commons Recommended Citation Fukuda, Yu , "ABCB6 Is a Porphyrin Transporter with a Novel Trafficking Signal That Is Conserved in Other ABC Transporters" (2008). Theses and Dissertations (ETD). Paper 345. http://dx.doi.org/10.21007/etd.cghs.2008.0100. This Dissertation is brought to you for free and open access by the College of Graduate Health Sciences at UTHSC Digital Commons. It has been accepted for inclusion in Theses and Dissertations (ETD) by an authorized administrator of UTHSC Digital Commons. For more information, please contact [email protected]. ABCB6 Is a Porphyrin Transporter with a Novel Trafficking Signal That Is Conserved in Other ABC Transporters Document Type Dissertation Degree Name Doctor of Philosophy (PhD) Program Interdisciplinary Program Research Advisor John D. Schuetz, Ph.D. Committee Linda Hendershot, Ph.D. James I. Morgan, Ph.D. Anjaparavanda P. Naren, Ph.D. Jie Zheng, Ph.D. DOI 10.21007/etd.cghs.2008.0100 This dissertation is available at UTHSC Digital Commons: https://dc.uthsc.edu/dissertations/345 ABCB6 IS A PORPHYRIN TRANSPORTER WITH A NOVEL TRAFFICKING SIGNAL THAT -
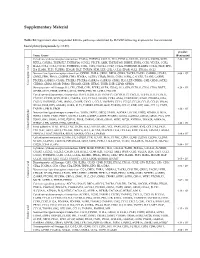
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R,