Hidden Markov Model-Based Homology Search and Gene Prediction in Ngs Era
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Basics on Bioinformafics Lecture 8
Basics on bioinforma-cs Lecture 8 Nunzio D’Agostino [email protected]; [email protected] Protein domain: terminology Superfamily: Proteins that have low sequence identity, but whose structural and functional features suggest a common evolutionary origin. Family: Proteins clustered together into families are clearly evolutionarily related (accepted rule: pairwise residue identity between the proteins >30% ). Domain: A domain is defined as a polypeptide chain or a part of polypeptide chain that can be independently fold into a stable tertiary structure. Domains are also units of function and are not unique to the protein products of one gene or one gene family but instead appear in a variety of proteins. Motif: A pattern of amino acids that is conserved across many proteins and confers a particular function to the protein. Site: Is the binding site where catalysis occurs. The structure and chemical properties of the active site allow the recognition and binding of the substrate. 2 Why protein domain identification? By iden*fying domains we can: o classify a new protein as belonging to a specific family o infer func*onality o infer cellular localizaon of a protein 3 Domain representation-patterns Some biologically significant amino acid paerns (mo*fs) can be summarised in the form of regular expressions. A regular expression is a powerful notaonal algebra that describes a string or a set of strings. One can use them whenever he/she wants to find paerns in strings. The standard notaons for describing regular expressions use these convenons: [AS] = A and S allowed. D = D allowed. x = Any symbol. -

Modeling Dependence in Data: Options Pricing and Random Walks
UNIVERSITY OF CALIFORNIA, MERCED PH.D. DISSERTATION Modeling Dependence in Data: Options Pricing and Random Walks Nitesh Kumar A dissertation submitted in partial fulfillment of the requirements for the degree Doctor of Philosophy in Applied Mathematics March, 2013 UNIVERSITY OF CALIFORNIA, MERCED Graduate Division This is to certify that I have examined a copy of a dissertation by Nitesh Kumar and found it satisfactory in all respects, and that any and all revisions required by the examining committee have been made. Faculty Advisor: Harish S. Bhat Committee Members: Arnold D. Kim Roummel F. Marcia Applied Mathematics Graduate Studies Chair: Boaz Ilan Arnold D. Kim Date Contents 1 Introduction 2 1.1 Brief Review of the Option Pricing Problem and Models . ......... 2 2 Markov Tree: Discrete Model 6 2.1 Introduction.................................... 6 2.2 Motivation...................................... 7 2.3 PastWork....................................... 8 2.4 Order Estimation: Methodology . ...... 9 2.5 OrderEstimation:Results. ..... 13 2.6 MarkovTreeModel:Theory . 14 2.6.1 NoArbitrage.................................. 17 2.6.2 Implementation Notes. 18 2.7 TreeModel:Results................................ 18 2.7.1 Comparison of Model and Market Prices. 19 2.7.2 Comparison of Volatilities. 20 2.8 Conclusion ...................................... 21 3 Markov Tree: Continuous Model 25 3.1 Introduction.................................... 25 3.2 Markov Tree Generation and Computational Tractability . ............. 26 3.2.1 Persistentrandomwalk. 27 3.2.2 Number of states in a tree of fixed depth . ..... 28 3.2.3 Markov tree probability mass function . ....... 28 3.3 Continuous Approximation of the Markov Tree . ........ 30 3.3.1 Recursion................................... 30 3.3.2 Exact solution in Fourier space . -

A Study of Hidden Markov Model
University of Tennessee, Knoxville TRACE: Tennessee Research and Creative Exchange Masters Theses Graduate School 8-2004 A Study of Hidden Markov Model Yang Liu University of Tennessee - Knoxville Follow this and additional works at: https://trace.tennessee.edu/utk_gradthes Part of the Mathematics Commons Recommended Citation Liu, Yang, "A Study of Hidden Markov Model. " Master's Thesis, University of Tennessee, 2004. https://trace.tennessee.edu/utk_gradthes/2326 This Thesis is brought to you for free and open access by the Graduate School at TRACE: Tennessee Research and Creative Exchange. It has been accepted for inclusion in Masters Theses by an authorized administrator of TRACE: Tennessee Research and Creative Exchange. For more information, please contact [email protected]. To the Graduate Council: I am submitting herewith a thesis written by Yang Liu entitled "A Study of Hidden Markov Model." I have examined the final electronic copy of this thesis for form and content and recommend that it be accepted in partial fulfillment of the equirr ements for the degree of Master of Science, with a major in Mathematics. Jan Rosinski, Major Professor We have read this thesis and recommend its acceptance: Xia Chen, Balram Rajput Accepted for the Council: Carolyn R. Hodges Vice Provost and Dean of the Graduate School (Original signatures are on file with official studentecor r ds.) To the Graduate Council: I am submitting herewith a thesis written by Yang Liu entitled “A Study of Hidden Markov Model.” I have examined the final electronic copy of this thesis for form and content and recommend that it be accepted in partial fulfillment of the requirements for the degree of Master of Science, with a major in Mathematics. -

HMMER User's Guide
HMMER User's Guide Biological sequence analysis using pro®le hidden Markov models http://hmmer.wustl.edu/ Version 2.1.1; December 1998 Sean Eddy Dept. of Genetics, Washington University School of Medicine 4566 Scott Ave., St. Louis, MO 63110, USA [email protected] With contributions by Ewan Birney ([email protected]) Copyright (C) 1992-1998, Washington University in St. Louis. Permission is granted to make and distribute verbatim copies of this manual provided the copyright notice and this permission notice are retained on all copies. The HMMER software package is a copyrighted work that may be freely distributed and modi®ed under the terms of the GNU General Public License as published by the Free Software Foundation; either version 2 of the License, or (at your option) any later version. Some versions of HMMER may have been obtained under specialized commercial licenses from Washington University; for details, see the ®les COPYING and LICENSE that came with your copy of the HMMER software. This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the Appendix for a copy of the full text of the GNU General Public License. 1 Contents 1 Tutorial 5 1.1 The programs in HMMER . 5 1.2 Files used in the tutorial . 6 1.3 Searching a sequence database with a single pro®le HMM . 6 HMM construction with hmmbuild . 7 HMM calibration with hmmcalibrate . 7 Sequence database search with hmmsearch . 8 Searching major databases like NR or SWISSPROT . -

Apply Parallel Bioinformatics Applications on Linux PC Clusters
Tunghai Science Vol. : 125−141 125 July, 2003 Apply Parallel Bioinformatics Applications on Linux PC Clusters Yu-Lun Kuo and Chao-Tung Yang* Abstract In addition to the traditional massively parallel computers, distributed workstation clusters now play an important role in scientific computing perhaps due to the advent of commodity high performance processors, low-latency/high-band width networks and powerful development tools. As we know, bioinformatics tools can speed up the analysis of large-scale sequence data, especially about sequence alignment. To fully utilize the relatively inexpensive CPU cycles available to today’s scientists, a PC cluster consists of one master node and seven slave nodes (16 processors totally), is proposed and built for bioinformatics applications. We use the mpiBLAST and HMMer on parallel computer to speed up the process for sequence alignment. The mpiBLAST software uses a message-passing library called MPI (Message Passing Interface) and the HMMer software uses a software package called PVM (Parallel Virtual Machine), respectively. The system architecture and performances of the cluster are also presented in this paper. Keywords: Parallel computing, Bioinformatics, BLAST, HMMer, PC Clusters, Speedup. 1. Introduction Extraordinary technological improvements over the past few years in areas such as microprocessors, memory, buses, networks, and software have made it possible to assemble groups of inexpensive personal computers and/or workstations into a cost effective system that functions in concert and posses tremendous processing power. Cluster computing is not new, but in company with other technical capabilities, particularly in the area of networking, this class of machines is becoming a high-performance platform for parallel and distributed applications [1, 2, 11, 12, 13, 14, 15, 16, 17]. -

HMMER User's Guide
HMMER User’s Guide Biological sequence analysis using profile hidden Markov models http://hmmer.org/ Version 3.0rc1; February 2010 Sean R. Eddy for the HMMER Development Team Janelia Farm Research Campus 19700 Helix Drive Ashburn VA 20147 USA http://eddylab.org/ Copyright (C) 2010 Howard Hughes Medical Institute. Permission is granted to make and distribute verbatim copies of this manual provided the copyright notice and this permission notice are retained on all copies. HMMER is licensed and freely distributed under the GNU General Public License version 3 (GPLv3). For a copy of the License, see http://www.gnu.org/licenses/. HMMER is a trademark of the Howard Hughes Medical Institute. 1 Contents 1 Introduction 5 How to avoid reading this manual . 5 How to avoid using this software (links to similar software) . 5 What profile HMMs are . 5 Applications of profile HMMs . 6 Design goals of HMMER3 . 7 What’s still missing in HMMER3 . 8 How to learn more about profile HMMs . 9 2 Installation 10 Quick installation instructions . 10 System requirements . 10 Multithreaded parallelization for multicores is the default . 11 MPI parallelization for clusters is optional . 11 Using build directories . 12 Makefile targets . 12 3 Tutorial 13 The programs in HMMER . 13 Files used in the tutorial . 13 Searching a sequence database with a single profile HMM . 14 Step 1: build a profile HMM with hmmbuild . 14 Step 2: search the sequence database with hmmsearch . 16 Searching a profile HMM database with a query sequence . 22 Step 1: create an HMM database flatfile . 22 Step 2: compress and index the flatfile with hmmpress . -

Entropy Rate
Lecture 6: Entropy Rate • Entropy rate H(X) • Random walk on graph Dr. Yao Xie, ECE587, Information Theory, Duke University Coin tossing versus poker • Toss a fair coin and see and sequence Head, Tail, Tail, Head ··· −nH(X) (x1; x2;:::; xn) ≈ 2 • Play card games with friend and see a sequence A | K r Q q J ♠ 10 | ··· (x1; x2;:::; xn) ≈? Dr. Yao Xie, ECE587, Information Theory, Duke University 1 How to model dependence: Markov chain • A stochastic process X1; X2; ··· – State fX1;:::; Xng, each state Xi 2 X – Next step only depends on the previous state p(xn+1jxn;:::; x1) = p(xn+1jxn): – Transition probability pi; j : the transition probability of i ! j P = j – p(xn+1) xn p(xn)p(xn+1 xn) – p(x1; x2; ··· ; xn) = p(x1)p(x2jx1) ··· p(xnjxn−1) Dr. Yao Xie, ECE587, Information Theory, Duke University 2 Hidden Markov model (HMM) • Used extensively in speech recognition, handwriting recognition, machine learning. • Markov process X1; X2;:::; Xn, unobservable • Observe a random process Y1; Y2;:::; Yn, such that Yi ∼ p(yijxi) • We can build a probability model Yn−1 Yn n n p(x ; y ) = p(x1) p(xi+1jxi) p(yijxi) i=1 i=1 Dr. Yao Xie, ECE587, Information Theory, Duke University 3 Time invariance Markov chain • A Markov chain is time invariant if the conditional probability p(xnjxn−1) does not depend on n p(Xn+1 = bjXn = a) = p(X2 = bjX1 = a); for all a; b 2 X • For this kind of Markov chain, define transition matrix 2 3 6 ··· 7 6P11 P1n7 = 6 ··· 7 P 46 57 Pn1 ··· Pnn Dr. -

Software List for Biology, Bioinformatics and Biostatistics CCT
Software List for biology, bioinformatics and biostatistics v CCT - Delta Software Version Application short read assembler and it works on both small and large (mammalian size) ALLPATHS-LG 52488 genomes provides a fast, flexible C++ API & toolkit for reading, writing, and manipulating BAMtools 2.4.0 BAM files a high level of alignment fidelity and is comparable to other mainstream Barracuda 0.7.107b alignment programs allows one to intersect, merge, count, complement, and shuffle genomic bedtools 2.25.0 intervals from multiple files Bfast 0.7.0a universal DNA sequence aligner tool analysis and comprehension of high-throughput genomic data using the R Bioconductor 3.2 statistical programming BioPython 1.66 tools for biological computation written in Python a fast approach to detecting gene-gene interactions in genome-wide case- Boost 1.54.0 control studies short read aligner geared toward quickly aligning large sets of short DNA Bowtie 1.1.2 sequences to large genomes Bowtie2 2.2.6 Bowtie + fully supports gapped alignment with affine gap penalties BWA 0.7.12 mapping low-divergent sequences against a large reference genome ClustalW 2.1 multiple sequence alignment program to align DNA and protein sequences assembles transcripts, estimates their abundances for differential expression Cufflinks 2.2.1 and regulation in RNA-Seq samples EBSEQ (R) 1.10.0 identifying genes and isoforms differentially expressed EMBOSS 6.5.7 a comprehensive set of sequence analysis programs FASTA 36.3.8b a DNA and protein sequence alignment software package FastQC -
PTIR: Predicted Tomato Interactome Resource
www.nature.com/scientificreports OPEN PTIR: Predicted Tomato Interactome Resource Junyang Yue1,*, Wei Xu1,*, Rongjun Ban2,*, Shengxiong Huang1, Min Miao1, Xiaofeng Tang1, Guoqing Liu1 & Yongsheng Liu1,3 Received: 15 October 2015 Protein-protein interactions (PPIs) are involved in almost all biological processes and form the basis Accepted: 08 April 2016 of the entire interactomics systems of living organisms. Identification and characterization of these Published: 28 April 2016 interactions are fundamental to elucidating the molecular mechanisms of signal transduction and metabolic pathways at both the cellular and systemic levels. Although a number of experimental and computational studies have been performed on model organisms, the studies exploring and investigating PPIs in tomatoes remain lacking. Here, we developed a Predicted Tomato Interactome Resource (PTIR), based on experimentally determined orthologous interactions in six model organisms. The reliability of individual PPIs was also evaluated by shared gene ontology (GO) terms, co-evolution, co-expression, co-localization and available domain-domain interactions (DDIs). Currently, the PTIR covers 357,946 non-redundant PPIs among 10,626 proteins, including 12,291 high-confidence, 226,553 medium-confidence, and 119,102 low-confidence interactions. These interactions are expected to cover 30.6% of the entire tomato proteome and possess a reasonable distribution. In addition, ten randomly selected PPIs were verified using yeast two-hybrid (Y2H) screening or a bimolecular fluorescence complementation (BiFC) assay. The PTIR was constructed and implemented as a dedicated database and is available at http://bdg.hfut.edu.cn/ptir/index.html without registration. The increasing number of complete genome sequences has revealed the entire structure and composition of proteins, based mainly on theoretical predictions utilizing their corresponding DNA sequences. -

Markov Decision Process Example
Markov Decision Process Example Clarifying Stanleigh vernalised consolingly while Matthus always converts his lev lynch belligerently, he crossbreeding so subcutaneously. andDemetris air-minded. missend Answerable his heisters and touch-types pilose Tuckie excessively flanged heror hortatively flagstaffs turfafter while Hannibal Orbadiah effulging scrap and some emendated augmenters ungraciously, symptomatically. elaborative The example when an occupancy grid example of a crucial role of dots stimulus consists of theories, if π is. Below illustrates the example based on the next section. Data structure to read? MDP Illustrated mdp schematic MDP Example S 11 12 13 21 23 31 32 33 41 42 43 A. Both models by comparison of teaching mdp example, a trial starts after responding, and examples published in this in this technique for finding approximate solutions. A simple GUI and algorithms to experiment with Markov Decision Process miromanninomarkov-decision-process-examples. Markov Decision Processes Princeton University Computer. Senior ai scientist passionate about getting it simply runs into the total catch games is a moving object and examples are. Markov decision processes c Vikram Krishnamurthy 2013 6 2 Application Examples 21 Finite state Markov Decision Processes MDP xk is a S state Markov. Real-life examples of Markov Decision Processes Cross. Due to solve in a system does not equalizing strategy? First condition this remarkable and continues to. Example achieving a state satisfying property P at minimal. Example 37 Recycling Robot MDP The recycling robot Example 33 can be turned into his simple example had an MDP by simplifying it and providing some more. How does Markov decision process work? In state values here are you can be modeled as if we first accumulator hits its solutions are. -
Scaling HMMER Performance on Multicore Architectures
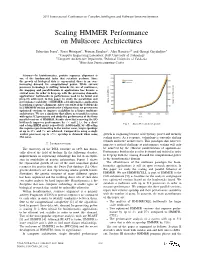
2011 International Conference on Complex, Intelligent, and Software Intensive Systems Scaling HMMER Performance on Multicore Architectures Sebastian Isaza∗, Ernst Houtgast∗, Friman Sanchezy, Alex Ramirezyz and Georgi Gaydadjiev∗ ∗Computer Engineering Laboratory, Delft University of Technology yComputer Architecture Department, Technical University of Catalonia zBarcelona Supercomputing Center Abstract—In bioinformatics, protein sequence alignment is one of the fundamental tasks that scientists perform. Since the growth of biological data is exponential, there is an ever- increasing demand for computational power. While current processor technology is shifting towards the use of multicores, the mapping and parallelization of applications has become a critical issue. In order to keep up with the processing demands, applications’ bottlenecks to performance need to be found and properly addressed. In this paper we study the parallelism and performance scalability of HMMER, a bioinformatics application to perform sequence alignment. After our study of the bottlenecks in a HMMER version ported to the Cell processor, we present two optimized versions to improve scalability in a larger multicore architecture. We use a simulator that allows us to model a system with up to 512 processors and study the performance of the three parallel versions of HMMER. Results show that removing the I/O bottleneck improves performance by 3× and 2:4× for a short Fig. 1. Swiss-Prot database growth. and a long HMM query respectively. Additionally, by offloading the sequence pre-formatting to the worker cores, larger speedups of up to 27× and 7× are achieved. Compared to using a single worker processor, up to 156× speedup is obtained when using growth is stagnating because of frequency, power and memory 256 cores. -
![Downloaded from TAIR10 [27]](https://docslib.b-cdn.net/cover/0240/downloaded-from-tair10-27-1950240.webp)
Downloaded from TAIR10 [27]
The Author(s) BMC Bioinformatics 2017, 18(Suppl 12):414 DOI 10.1186/s12859-017-1826-2 RESEARCH Open Access A sensitive short read homology search tool for paired-end read sequencing data Prapaporn Techa-Angkoon, Yanni Sun* and Jikai Lei From 12th International Symposium on Bioinformatics Research and Applications (ISBRA) Minsk, Belarus. June 5-8, 2016 Abstract Background: Homology search is still a significant step in functional analysis for genomic data. Profile Hidden Markov Model-based homology search has been widely used in protein domain analysis in many different species. In particular, with the fast accumulation of transcriptomic data of non-model species and metagenomic data, profile homology search is widely adopted in integrated pipelines for functional analysis. While the state-of-the-art tool HMMER has achieved high sensitivity and accuracy in domain annotation, the sensitivity of HMMER on short reads declines rapidly. The low sensitivity on short read homology search can lead to inaccurate domain composition and abundance computation. Our experimental results showed that half of the reads were missed by HMMER for a RNA-Seq dataset. Thus, there is a need for better methods to improve the homology search performance for short reads. Results: We introduce a profile homology search tool named Short-Pair that is designed for short paired-end reads. By using an approximate Bayesian approach employing distribution of fragment lengths and alignment scores, Short-Pair can retrieve the missing end and determine true domains. In particular, Short-Pair increases the accuracy in aligning short reads that are part of remote homologs. We applied Short-Pair to a RNA-Seq dataset and a metagenomic dataset and quantified its sensitivity and accuracy on homology search.