Computational Biology Lecture 8: Substitution Matrices Saad Mneimneh
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Optimal Matching Distances Between Categorical Sequences: Distortion and Inferences by Permutation Juan P
St. Cloud State University theRepository at St. Cloud State Culminating Projects in Applied Statistics Department of Mathematics and Statistics 12-2013 Optimal Matching Distances between Categorical Sequences: Distortion and Inferences by Permutation Juan P. Zuluaga Follow this and additional works at: https://repository.stcloudstate.edu/stat_etds Part of the Applied Statistics Commons Recommended Citation Zuluaga, Juan P., "Optimal Matching Distances between Categorical Sequences: Distortion and Inferences by Permutation" (2013). Culminating Projects in Applied Statistics. 8. https://repository.stcloudstate.edu/stat_etds/8 This Thesis is brought to you for free and open access by the Department of Mathematics and Statistics at theRepository at St. Cloud State. It has been accepted for inclusion in Culminating Projects in Applied Statistics by an authorized administrator of theRepository at St. Cloud State. For more information, please contact [email protected]. OPTIMAL MATCHING DISTANCES BETWEEN CATEGORICAL SEQUENCES: DISTORTION AND INFERENCES BY PERMUTATION by Juan P. Zuluaga B.A. Universidad de los Andes, Colombia, 1995 A Thesis Submitted to the Graduate Faculty of St. Cloud State University in Partial Fulfillment of the Requirements for the Degree Master of Science St. Cloud, Minnesota December, 2013 This thesis submitted by Juan P. Zuluaga in partial fulfillment of the requirements for the Degree of Master of Science at St. Cloud State University is hereby approved by the final evaluation committee. Chairperson Dean School of Graduate Studies OPTIMAL MATCHING DISTANCES BETWEEN CATEGORICAL SEQUENCES: DISTORTION AND INFERENCES BY PERMUTATION Juan P. Zuluaga Sequence data (an ordered set of categorical states) is a very common type of data in Social Sciences, Genetics and Computational Linguistics. -

Bioinformatics 1: Lecture 3
Bioinformatics 1: Lecture 3 •Pairwise alignment •Substitution •Dynamic Programming algorithm Scoring matrix To prepare an alignment, we first consider the score for aligning (associating) any one character of the first sequence with any one character of the second sequence. A A G A C G T T T A G A C T 0 0 1 0 0 1 0 0 0 0 1 1 0 1 0 0 0 0 0 1 0 0 0 0 1 0 0 0 0 0 Exact match 0 0 1 0 0 1 0 0 0 0 1/0 0 0 0 0 0 0 1 1 1 0 1 1 0 1 0 0 0 0 0 1 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 1 1 1 0 The cost of mutation is not a constant DNA: A change in the 3rd base in a codon, and sometimes the first base, sometimes conserves the amino acid. No selective pressure. Protein: A change in amino acids that are in the same chemical class conserve their chemical environment. For example: Lys to Arg is conservative because both a positively charged. Conservative amino acid changes N Lys <--> Arg C + N` N N` C N C + C C C N` C C C C O C O C C N N Ile <--> Leu C C C C C C C C O C O C C Ser <--> Thr Asp <--> Glu Asn <--> Gln If the “chemistry” of the sidechain is conserved, then the mutation is less likely to change structure/function. -

BASS: Approximate Search on Large String Databases
BASS: Approximate Search on Large String Databases Jiong Yang Wei Wang Philip Yu UIUC UNC Chapel Hill IBM [email protected] [email protected] [email protected] Abstract Similarity search on a string database can be classified into two categories: exact match and approximate match. The In this paper, we study the problem on how to build an index struc- search of exact match looks for substrings in the database, ture for large string databases to efficiently support various types of which is exactly identical to the query pattern while the search string matching without the necessity of mapping the substrings to of approximate match allows some types of imperfection such a numerical space (e.g., string B-tree and MRS-index) nor the re- as substitutions between certain symbols, some degree of mis- striction of in-memory practice (e.g., suffix tree and suffix array). alignment, and the presence of “wild-card” in the query pat- Towards this goal, we propose a new indexing scheme, BASS-tree, tern. We shall mention that supporting approximate match is to efficiently support general approximate substring match (in terms very important to many applications. For instance, biologists of certain symbol substitutions and misalignments) in sublinear time have observed that mutations between certain pair of amino on a large string database. The key idea behind the design is that all acids may occur at a noticeable probability in some proteins positions in each string are grouped recursively into a fully balanced and such a mutation usually does not alter the biological func- tree according to the similarities of the subsequent segments starting tion of the proteins. -

Sequence Motifs, Correlations and Structural Mapping of Evolutionary
Talk overview • Sequence profiles – position specific scoring matrix • Psi-blast. Automated way to create and use sequence Sequence motifs, correlations profiles in similarity searches and structural mapping of • Sequence patterns and sequence logos evolutionary data • Bioinformatic tools which employ sequence profiles: PFAM BLOCKS PROSITE PRINTS InterPro • Correlated Mutations and structural insight • Mapping sequence data on structures: March 2011 Eran Eyal Conservations Correlations PSSM – position specific scoring matrix • A position-specific scoring matrix (PSSM) is a commonly used representation of motifs (patterns) in biological sequences • PSSM enables us to represent multiple sequence alignments as mathematical entities which we can work with. • PSSMs enables the scoring of multiple alignments with sequences, or other PSSMs. PSSM – position specific scoring matrix Assuming a string S of length n S = s1s2s3...sn If we want to score this string against our PSSM of length n (with n lines): n alignment _ score = m ∑ s j , j j=1 where m is the PSSM matrix and sj are the string elements. PSSM can also be incorporated to both dynamic programming algorithms and heuristic algorithms (like Psi-Blast). Sequence space PSI-BLAST • For a query sequence use Blast to find matching sequences. • Construct a multiple sequence alignment from the hits to find the common regions (consensus). • Use the “consensus” to search again the database, and get a new set of matching sequences • Repeat the process ! Sequence space Position-Specific-Iterated-BLAST • Intuition – substitution matrices should be specific to sites and not global. – Example: penalize alanine→glycine more in a helix •Idea – Use BLAST with high stringency to get a set of closely related sequences. -

Development of Novel Classical and Quantum Information Theory Based Methods for the Detection of Compensatory Mutations in Msas
Development of novel Classical and Quantum Information Theory Based Methods for the Detection of Compensatory Mutations in MSAs Dissertation zur Erlangung des mathematisch-naturwissenschaftlichen Doktorgrades ”Doctor rerum naturalium” der Georg-August-Universität Göttingen im Promotionsprogramm PCS der Georg-August University School of Science (GAUSS) vorgelegt von Mehmet Gültas aus Kirikkale-Türkei Göttingen, 2013 Betreuungsausschuss Professor Dr. Stephan Waack, Institut für Informatik, Georg-August-Universität Göttingen. Professor Dr. Carsten Damm, Institut für Informatik, Georg-August-Universität Göttingen. Professor Dr. Edgar Wingender, Institut für Bioinformatik, Universitätsmedizin, Georg-August-Universität Göttingen. Mitglieder der Prüfungskommission Referent: Prof. Dr. Stephan Waack, Institut für Informatik, Georg-August-Universität Göttingen. Korreferent: Prof. Dr. Carsten Damm, Institut für Informatik, Georg-August-Universität Göttingen. Korreferent: Prof. Dr. Mario Stanke, Institut für Mathematik und Informatik, Ernst Moritz Arndt Universität Greifswald Weitere Mitglieder der Prüfungskommission Prof. Dr. Edgar Wingender, Institut für Bioinformatik, Universitätsmedizin, Georg-August-Universität Göttingen. Prof. Dr. Burkhard Morgenstern, Institut für Mikrobiologie und Genetik, Abteilung für Bioinformatik, Georg-August- Universität Göttingen. Prof. Dr. Dieter Hogrefe, Institut für Informatik, Georg-August-Universität Göttingen. Prof. Dr. Wolfgang May, Institut für Informatik, Georg-August-Universität Göttingen. Tag der mündlichen -

3D Representations of Amino Acids—Applications to Protein Sequence Comparison and Classification
Computational and Structural Biotechnology Journal 11 (2014) 47–58 Contents lists available at ScienceDirect journal homepage: www.elsevier.com/locate/csbj 3D representations of amino acids—applications to protein sequence comparison and classification Jie Li a, Patrice Koehl b,⁎ a Genome Center, University of California, Davis, 451 Health Sciences Drive, Davis, CA 95616, United States b Department of Computer Science and Genome Center, University of California, Davis, One Shields Ave, Davis, CA 95616, United States article info abstract Available online 6 September 2014 The amino acid sequence of a protein is the key to understanding its structure and ultimately its function in the cell. This paper addresses the fundamental issue of encoding amino acids in ways that the representation of such Keywords: a protein sequence facilitates the decoding of its information content. We show that a feature-based representa- Protein sequences tion in a three-dimensional (3D) space derived from amino acid substitution matrices provides an adequate Substitution matrices representation that can be used for direct comparison of protein sequences based on geometry. We measure Protein sequence classification the performance of such a representation in the context of the protein structural fold prediction problem. Fold recognition We compare the results of classifying different sets of proteins belonging to distinct structural folds against classifications of the same proteins obtained from sequence alone or directly from structural information. We find that sequence alone performs poorly as a structure classifier.Weshowincontrastthattheuseofthe three dimensional representation of the sequences significantly improves the classification accuracy. We conclude with a discussion of the current limitations of such a representation and with a description of potential improvements. -

B.Sc. (Hons.) Biotech BIOT 3013 Unit-5 Satarudra P Singh
B.Sc. (Hons.) Biotechnology Core Course 13: Basics of Bioinformatics and Biostatistics (BIOT 3013 ) Unit 5: Sequence Alignment and database searching Dr. Satarudra Prakash Singh Department of Biotechnology Mahatma Gandhi Central University, Motihari Challenges in bioinformatics 1. Obtain the genome of an organism. 2. Identify and annotate genes. 3. Find the sequences, three dimensional structures, and functions of proteins. 4. Find sequences of proteins that have desired three dimensional structures. 5. Compare DNA sequences and proteins sequences for similarity. 6. Study the evolution of sequences and species. Sequence alignments lie at the heart of all bioinformatics Definition of Sequence Alignment • Sequence alignment is the procedure of comparing two or more sequences by searching for a series of individual characters or character patterns that are in the same order in the sequences. LGPSSKQTGKGS - SRIWDN Global Alignment LN – ITKSAGKGAIMRLFDA --------TGKG --------- Local Alignment --------AGKG --------- • In global alignment, an attempt is made to align the entire sequences, as many characters as possible. • In local alignment, stretches of sequence with the highest density of matches are given the highest priority, thus generating one or more islands of matches in the aligned sequences. • Eg: problem of locating the famous TATAAT - box (a bacterial promoter) in a piece of DNA. Method for pairwise sequence Alignment: Dynamic Programming • Global Alignment: Needleman- Wunsch Algorithm • Local Alignment: Smith-Waterman Algorithm Needleman & Wunsch algorithm : Global alignment • There are three major phases: 1. initialization 2. Fill 3. Trace back. • Initialization assign values for the first row and column. • The score of each cell is set to the gap score multiplied by the distance from the origin. -

Parameter Advising for Multiple Sequence Alignment
PARAMETER ADVISING FOR MULTIPLE SEQUENCE ALIGNMENT by Daniel Frank DeBlasio Copyright c Daniel Frank DeBlasio 2016 A Dissertation Submitted to the Faculty of the DEPARTMENT OF COMPUTER SCIENCE In Partial Fulfillment of the Requirements For the Degree of DOCTOR OF PHILOSOPHY In the Graduate College THE UNIVERSITY OF ARIZONA 2016 2 THE UNIVERSITY OF ARIZONA GRADUATE COLLEGE As members of the Dissertation Committee, we certify that we have read the disser- tation prepared by Daniel Frank DeBlasio, entitled Parameter Advising for Multiple Sequence Alignment and recommend that it be accepted as fulfilling the dissertation requirement for the Degree of Doctor of Philosophy. Date: 15 April 2016 John Kececioglu Date: 15 April 2016 Alon Efrat Date: 15 April 2016 Stephen Kobourov Date: 15 April 2016 Mike Sanderson Final approval and acceptance of this dissertation is contingent upon the candidate's submission of the final copies of the dissertation to the Graduate College. I hereby certify that I have read this dissertation prepared under my direction and recommend that it be accepted as fulfilling the dissertation requirement. Date: 15 April 2016 Dissertation Director: John Kececioglu 3 STATEMENT BY AUTHOR This dissertation has been submitted in partial fulfillment of requirements for an advanced degree at the University of Arizona and is deposited in the University Library to be made available to borrowers under rules of the Library. Brief quotations from this dissertation are allowable without special permission, pro- vided that accurate acknowledgment of the source is made. Requests for permission for extended quotation from or reproduction of this manuscript in whole or in part may be granted by the copyright holder. -

Pairwise Statistical Significance of Local Sequence Alignment Using Sequence-Specific and Position-Specific Substitution Matrices
194 IEEE/ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 8, NO. 1, JANUARY/FEBRUARY 2011 Pairwise Statistical Significance of Local Sequence Alignment Using Sequence-Specific and Position-Specific Substitution Matrices Ankit Agrawal and Xiaoqiu Huang Abstract—Pairwise sequence alignment is a central problem in bioinformatics, which forms the basis of various other applications. Two related sequences are expected to have a high alignment score, but relatedness is usually judged by statistical significance rather than by alignment score. Recently, it was shown that pairwise statistical significance gives promising results as an alternative to database statistical significance for getting individual significance estimates of pairwise alignment scores. The improvement was mainly attributed to making the statistical significance estimation process more sequence-specific and database-independent. In this paper, we use sequence-specific and position-specific substitution matrices to derive the estimates of pairwise statistical significance, which is expected to use more sequence-specific information in estimating pairwise statistical significance. Experiments on a benchmark database with sequence-specific substitution matrices at different levels of sequence-specific contribution were conducted, and results confirm that using sequence-specific substitution matrices for estimating pairwise statistical significance is significantly better than using a standard matrix like BLOSUM62, and than database statistical significance estimates -
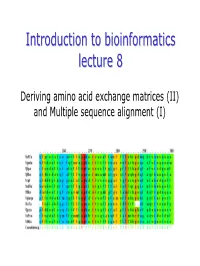
Deriving Amino Acid Exchange Matrices (II) and Multiple Sequence Alignment (I) Summarysummary Dayhoff’Sdayhoff’S PAMPAM--Matricesmatrices
IntroductionIntroduction toto bioinformaticsbioinformatics lecturelecture 88 Deriving amino acid exchange matrices (II) and Multiple sequence alignment (I) SummarySummary Dayhoff’sDayhoff’s PAMPAM--matricesmatrices Derived from global alignments of closely related sequences. Matrices for greater evolutionary distances are extrapolated from those for lesser ones. The number with the matrix (PAM40, PAM100) refers to the evolutionary distance; greater numbers are greater distances. Several later groups have attempted to extend Dayhoff's methodology or re-apply her analysis using later databases with more examples. Extensions of Dayhoff’s methodology: > Jones, Thornton and coworkers used the same methodology as Dayhoff but with modern databases (CABIOS 8:275). > Gonnett and coworkers (Science 256:1443) used a slightly different (but theoretically equivalent) methodology. > Henikoff & Henikoff (Proteins 17:49) compared these two newer versions of the PAM matrices with Dayhoff's originals. TheThe BLOSUMBLOSUM matricesmatrices ((BLOcksBLOcks SUbstitutionSUbstitution Matrix)Matrix) The BLOSUM series of matrices were created by Steve Henikoff and colleagues (PNAS 89:10915). Derived from local, un-gapped alignments of distantly related sequences. All matrices are directly calculated; no extrapolations are used. Again: the observed frequency of each pair is compared to the expected frequency (which is essentially the product of the frequencies of each residue in the dataset). Then: Log-odds matrix. TheThe BlocksBlocks DatabaseDatabase The Blocks Database contains multiple alignments of conserved regions in protein families. Blocks are multiply aligned un-gapped segments corresponding to the most highly conserved regions of proteins. The blocks for the BLOCKS database are made automatically by looking for the most highly conserved regions in groups of proteins represented in the PROSITE database. -

Assume an F84 Substitution Model with Nucleotide Frequ
Exercise Sheet 5 Computational Phylogenetics Prof. D. Metzler Exercise 1: Assume an F84 substitution model with nucleotide frequencies (πA; πC ; πG; πT ) = (0:2; 0:3; 0:3; 0:2), a rate λ = 0:1 of “crosses” and a rate µ = 0:2 of “bullets” (see lecture). (a) Assume that the nucleotide at some site is A at time t. Calculate the probabilities that the nucleotide is A, C or G at time t + 0:2. (b) Assume the nucleotide distribution in a genomic region is (0:1; 0:2; 0:3; 0:4) at time t, but from this time on the genomic region evolves according to the model above. Calculate the expectation values for the nucleotide distributions at time points t + 0:2 and t + 2. Exercise 2: Calculate rate matrix for the nucleotide substution process for which the substitution matrix for time t is 0 1−e−t=10 21+9e−t=10−30e−t=5 1−e−t=10 1 PA!A(t) 10 70 5 B 2−2e−t=10 3−3et=10 3+7e−t=10−10e−t=5 C B PC!C (t) C S(t) = B 5 10 15 C ; B 14+6e−t=10−20e−t=5 1−e−t=10 1−e−t=10 C B PG!G(t) C @ 35 10 5 A 2−2e−t=10 3+7e−t=10−10e−t=5 3−3e−t=10 5 30 10 PT !T (t) where the diagonal entries PA!A(t), PC!C (t), PG!G(t) and PT !T (t) are the values that fulfill that each row sum is 1 in the matrix S(t). -

Pairwise Sequence Alignment Algorithm by a New Measure Based
Hara et al. BMC Bioinformatics 2010, 11:235 http://www.biomedcentral.com/1471-2105/11/235 SOFTWARE Open Access MTRAP: Pairwise sequence alignment algorithm by a new measure based on transition probability between two consecutive pairs of residues Toshihide Hara*, Keiko Sato, Masanori Ohya Abstract Background: Sequence alignment is one of the most important techniques to analyze biological systems. It is also true that the alignment is not complete and we have to develop it to look for more accurate method. In particular, an alignment for homologous sequences with low sequence similarity is not in satisfactory level. Usual methods for aligning protein sequences in recent years use a measure empirically determined. As an example, a measure is usually defined by a combination of two quantities (1) and (2) below: (1) the sum of substitutions between two residue segments, (2) the sum of gap penalties in insertion/deletion region. Such a measure is determined on the assumption that there is no an intersite correlation on the sequences. In this paper, we improve the alignment by taking the correlation of consecutive residues. Results: We introduced a new method of alignment, called MTRAP by introducing a metric defined on compound systems of two sequences. In the benchmark tests by PREFAB 4.0 and HOMSTRAD, our pairwise alignment method gives higher accuracy than other methods such as ClustalW2, TCoffee, MAFFT. Especially for the sequences with sequence identity less than 15%, our method improves the alignment accuracy significantly. Moreover, we also showed that our algorithm works well together with a consistency-based progressive multiple alignment by modifying the TCoffee to use our measure.