Rotamer-Specific Statistical Potentials for Protein Structure Modeling
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Bioinformatics 1: Lecture 3
Bioinformatics 1: Lecture 3 •Pairwise alignment •Substitution •Dynamic Programming algorithm Scoring matrix To prepare an alignment, we first consider the score for aligning (associating) any one character of the first sequence with any one character of the second sequence. A A G A C G T T T A G A C T 0 0 1 0 0 1 0 0 0 0 1 1 0 1 0 0 0 0 0 1 0 0 0 0 1 0 0 0 0 0 Exact match 0 0 1 0 0 1 0 0 0 0 1/0 0 0 0 0 0 0 1 1 1 0 1 1 0 1 0 0 0 0 0 1 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 1 1 1 0 The cost of mutation is not a constant DNA: A change in the 3rd base in a codon, and sometimes the first base, sometimes conserves the amino acid. No selective pressure. Protein: A change in amino acids that are in the same chemical class conserve their chemical environment. For example: Lys to Arg is conservative because both a positively charged. Conservative amino acid changes N Lys <--> Arg C + N` N N` C N C + C C C N` C C C C O C O C C N N Ile <--> Leu C C C C C C C C O C O C C Ser <--> Thr Asp <--> Glu Asn <--> Gln If the “chemistry” of the sidechain is conserved, then the mutation is less likely to change structure/function. -

BASS: Approximate Search on Large String Databases
BASS: Approximate Search on Large String Databases Jiong Yang Wei Wang Philip Yu UIUC UNC Chapel Hill IBM [email protected] [email protected] [email protected] Abstract Similarity search on a string database can be classified into two categories: exact match and approximate match. The In this paper, we study the problem on how to build an index struc- search of exact match looks for substrings in the database, ture for large string databases to efficiently support various types of which is exactly identical to the query pattern while the search string matching without the necessity of mapping the substrings to of approximate match allows some types of imperfection such a numerical space (e.g., string B-tree and MRS-index) nor the re- as substitutions between certain symbols, some degree of mis- striction of in-memory practice (e.g., suffix tree and suffix array). alignment, and the presence of “wild-card” in the query pat- Towards this goal, we propose a new indexing scheme, BASS-tree, tern. We shall mention that supporting approximate match is to efficiently support general approximate substring match (in terms very important to many applications. For instance, biologists of certain symbol substitutions and misalignments) in sublinear time have observed that mutations between certain pair of amino on a large string database. The key idea behind the design is that all acids may occur at a noticeable probability in some proteins positions in each string are grouped recursively into a fully balanced and such a mutation usually does not alter the biological func- tree according to the similarities of the subsequent segments starting tion of the proteins. -

Computational Biology Lecture 8: Substitution Matrices Saad Mneimneh
Computational Biology Lecture 8: Substitution matrices Saad Mneimneh As we have introduced last time, simple scoring schemes like +1 for a match, -1 for a mismatch and -2 for a gap are not justifiable biologically, especially for amino acid sequences (proteins). Instead, more elaborated scoring functions are used. These scores are usually obtained as a result of analyzing chemical properties and statistical data for amino acids and DNA sequences. For example, it is known that same size amino acids are more likely to be substituted by one another. Similarly, amino acids with same affinity to water are likely to serve the same purpose in some cases. On the other hand, some mutations are not acceptable (may lead to demise of the organism). PAM and BLOSUM matrices are amongst results of such analysis. We will see the techniques through which PAM and BLOSUM matrices are obtained. Substritution matrices Chemical properties of amino acids govern how the amino acids substitue one another. In principle, a substritution matrix s, where sij is used to score aligning character i with character j, should reflect the probability of two characters substituing one another. The question is how to build such a probability matrix that closely maps reality? Different strategies result in different matrices but the central idea is the same. If we go back to the concept of a high scoring segment pair, theory tells us that the alignment (ungapped) given by such a segment is governed by a limiting distribution such that ¸sij qij = pipje where: ² s is the subsitution matrix used ² qij is the probability of observing character i aligned with character j ² pi is the probability of occurrence of character i Therefore, 1 qij sij = ln ¸ pipj This formula for sij suggests a way to constrcut the matrix s. -

B.Sc. (Hons.) Biotech BIOT 3013 Unit-5 Satarudra P Singh
B.Sc. (Hons.) Biotechnology Core Course 13: Basics of Bioinformatics and Biostatistics (BIOT 3013 ) Unit 5: Sequence Alignment and database searching Dr. Satarudra Prakash Singh Department of Biotechnology Mahatma Gandhi Central University, Motihari Challenges in bioinformatics 1. Obtain the genome of an organism. 2. Identify and annotate genes. 3. Find the sequences, three dimensional structures, and functions of proteins. 4. Find sequences of proteins that have desired three dimensional structures. 5. Compare DNA sequences and proteins sequences for similarity. 6. Study the evolution of sequences and species. Sequence alignments lie at the heart of all bioinformatics Definition of Sequence Alignment • Sequence alignment is the procedure of comparing two or more sequences by searching for a series of individual characters or character patterns that are in the same order in the sequences. LGPSSKQTGKGS - SRIWDN Global Alignment LN – ITKSAGKGAIMRLFDA --------TGKG --------- Local Alignment --------AGKG --------- • In global alignment, an attempt is made to align the entire sequences, as many characters as possible. • In local alignment, stretches of sequence with the highest density of matches are given the highest priority, thus generating one or more islands of matches in the aligned sequences. • Eg: problem of locating the famous TATAAT - box (a bacterial promoter) in a piece of DNA. Method for pairwise sequence Alignment: Dynamic Programming • Global Alignment: Needleman- Wunsch Algorithm • Local Alignment: Smith-Waterman Algorithm Needleman & Wunsch algorithm : Global alignment • There are three major phases: 1. initialization 2. Fill 3. Trace back. • Initialization assign values for the first row and column. • The score of each cell is set to the gap score multiplied by the distance from the origin. -

Parameter Advising for Multiple Sequence Alignment
PARAMETER ADVISING FOR MULTIPLE SEQUENCE ALIGNMENT by Daniel Frank DeBlasio Copyright c Daniel Frank DeBlasio 2016 A Dissertation Submitted to the Faculty of the DEPARTMENT OF COMPUTER SCIENCE In Partial Fulfillment of the Requirements For the Degree of DOCTOR OF PHILOSOPHY In the Graduate College THE UNIVERSITY OF ARIZONA 2016 2 THE UNIVERSITY OF ARIZONA GRADUATE COLLEGE As members of the Dissertation Committee, we certify that we have read the disser- tation prepared by Daniel Frank DeBlasio, entitled Parameter Advising for Multiple Sequence Alignment and recommend that it be accepted as fulfilling the dissertation requirement for the Degree of Doctor of Philosophy. Date: 15 April 2016 John Kececioglu Date: 15 April 2016 Alon Efrat Date: 15 April 2016 Stephen Kobourov Date: 15 April 2016 Mike Sanderson Final approval and acceptance of this dissertation is contingent upon the candidate's submission of the final copies of the dissertation to the Graduate College. I hereby certify that I have read this dissertation prepared under my direction and recommend that it be accepted as fulfilling the dissertation requirement. Date: 15 April 2016 Dissertation Director: John Kececioglu 3 STATEMENT BY AUTHOR This dissertation has been submitted in partial fulfillment of requirements for an advanced degree at the University of Arizona and is deposited in the University Library to be made available to borrowers under rules of the Library. Brief quotations from this dissertation are allowable without special permission, pro- vided that accurate acknowledgment of the source is made. Requests for permission for extended quotation from or reproduction of this manuscript in whole or in part may be granted by the copyright holder. -

Pairwise Statistical Significance of Local Sequence Alignment Using Sequence-Specific and Position-Specific Substitution Matrices
194 IEEE/ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 8, NO. 1, JANUARY/FEBRUARY 2011 Pairwise Statistical Significance of Local Sequence Alignment Using Sequence-Specific and Position-Specific Substitution Matrices Ankit Agrawal and Xiaoqiu Huang Abstract—Pairwise sequence alignment is a central problem in bioinformatics, which forms the basis of various other applications. Two related sequences are expected to have a high alignment score, but relatedness is usually judged by statistical significance rather than by alignment score. Recently, it was shown that pairwise statistical significance gives promising results as an alternative to database statistical significance for getting individual significance estimates of pairwise alignment scores. The improvement was mainly attributed to making the statistical significance estimation process more sequence-specific and database-independent. In this paper, we use sequence-specific and position-specific substitution matrices to derive the estimates of pairwise statistical significance, which is expected to use more sequence-specific information in estimating pairwise statistical significance. Experiments on a benchmark database with sequence-specific substitution matrices at different levels of sequence-specific contribution were conducted, and results confirm that using sequence-specific substitution matrices for estimating pairwise statistical significance is significantly better than using a standard matrix like BLOSUM62, and than database statistical significance estimates -
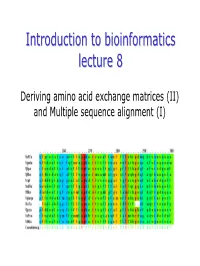
Deriving Amino Acid Exchange Matrices (II) and Multiple Sequence Alignment (I) Summarysummary Dayhoff’Sdayhoff’S PAMPAM--Matricesmatrices
IntroductionIntroduction toto bioinformaticsbioinformatics lecturelecture 88 Deriving amino acid exchange matrices (II) and Multiple sequence alignment (I) SummarySummary Dayhoff’sDayhoff’s PAMPAM--matricesmatrices Derived from global alignments of closely related sequences. Matrices for greater evolutionary distances are extrapolated from those for lesser ones. The number with the matrix (PAM40, PAM100) refers to the evolutionary distance; greater numbers are greater distances. Several later groups have attempted to extend Dayhoff's methodology or re-apply her analysis using later databases with more examples. Extensions of Dayhoff’s methodology: > Jones, Thornton and coworkers used the same methodology as Dayhoff but with modern databases (CABIOS 8:275). > Gonnett and coworkers (Science 256:1443) used a slightly different (but theoretically equivalent) methodology. > Henikoff & Henikoff (Proteins 17:49) compared these two newer versions of the PAM matrices with Dayhoff's originals. TheThe BLOSUMBLOSUM matricesmatrices ((BLOcksBLOcks SUbstitutionSUbstitution Matrix)Matrix) The BLOSUM series of matrices were created by Steve Henikoff and colleagues (PNAS 89:10915). Derived from local, un-gapped alignments of distantly related sequences. All matrices are directly calculated; no extrapolations are used. Again: the observed frequency of each pair is compared to the expected frequency (which is essentially the product of the frequencies of each residue in the dataset). Then: Log-odds matrix. TheThe BlocksBlocks DatabaseDatabase The Blocks Database contains multiple alignments of conserved regions in protein families. Blocks are multiply aligned un-gapped segments corresponding to the most highly conserved regions of proteins. The blocks for the BLOCKS database are made automatically by looking for the most highly conserved regions in groups of proteins represented in the PROSITE database. -

Pairwise Sequence Alignment Algorithm by a New Measure Based
Hara et al. BMC Bioinformatics 2010, 11:235 http://www.biomedcentral.com/1471-2105/11/235 SOFTWARE Open Access MTRAP: Pairwise sequence alignment algorithm by a new measure based on transition probability between two consecutive pairs of residues Toshihide Hara*, Keiko Sato, Masanori Ohya Abstract Background: Sequence alignment is one of the most important techniques to analyze biological systems. It is also true that the alignment is not complete and we have to develop it to look for more accurate method. In particular, an alignment for homologous sequences with low sequence similarity is not in satisfactory level. Usual methods for aligning protein sequences in recent years use a measure empirically determined. As an example, a measure is usually defined by a combination of two quantities (1) and (2) below: (1) the sum of substitutions between two residue segments, (2) the sum of gap penalties in insertion/deletion region. Such a measure is determined on the assumption that there is no an intersite correlation on the sequences. In this paper, we improve the alignment by taking the correlation of consecutive residues. Results: We introduced a new method of alignment, called MTRAP by introducing a metric defined on compound systems of two sequences. In the benchmark tests by PREFAB 4.0 and HOMSTRAD, our pairwise alignment method gives higher accuracy than other methods such as ClustalW2, TCoffee, MAFFT. Especially for the sequences with sequence identity less than 15%, our method improves the alignment accuracy significantly. Moreover, we also showed that our algorithm works well together with a consistency-based progressive multiple alignment by modifying the TCoffee to use our measure. -

Scoring Matrices for Sequence Comparisons
Scoring Matrices for Sequence Comparisons DEKM book Notes from Dr. Bino John and Dr. Takis Benos 1 Why compare sequences? • Given a new sequence, infer its funcHon based on similarity to another sequence • Find important molecular regions – conserved across species 2 Sequence -> Structure -> FuncHon MALRAK… SEQUENCE ? FUNCTION Cytochrome protein STRUCTURE 3 Important molecular regions conserved across species Human (C11A_HUMAN; P05108) vs. pig (C11A_PIG; P10612) Query: 1 MLAKGLPPRSVLVKGYQTFLSAPREGLGRLRVPTGEGAGISTRSPRPFNEIPSPGDNGWL 60 MLA+GL RSVLVKG Q FLSAPRE G RV TGEGA IST++PRPF+EIPSPGDNGW+ Sbjct: 1 MLARGLALRSVLVKGCQPFLSAPRECPGHPRVGTGEGACISTKTPRPFSEIPSPGDNGWI 60 Query: 61 NLYHFWRETGTHKVHLHHVQNFQKYGPIYREKLGNVESVYVIDPEDVALLFKSEGPNPER 120 NLY FW+E GT K+H HHVQNFQKYGPIYREKLGN+ESVY+IDPEDVALLFK EGPNPER Sbjct: 61 NLYRFWKEKGTQKIHYHHVQNFQKYGPIYREKLGNLESVYIIDPEDVALLFKFEGPNPER 120 Query: 121 FLIPPWVAYHQYYQRPIGVLLKKSAAWKKDRVALNQEVMAPEATKNFLPLLDAVSRDFVS 180 + IPPWVAYHQ+YQ+P+GVLLKKS AWKKDR+ LN EVMAPEA KNF+PLLD VS+DFV Sbjct: 121 YNIPPWVAYHQHYQKPVGVLLKKSGAWKKDRLVLNTEVMAPEAIKNFIPLLDTVSQDFVG 180 Query: 181 VLHRRIKKAGSGNYSGDISDDLFRFAFESITNVIFGERQGMLEEVVNPEAQRFIDAIYQM 240 VLHRRIK+ GSG +SGDI +DLFRFAFESITNVIFGER GMLEE+V+PEAQ+FIDA+YQM Sbjct: 181 VLHRRIKQQGSGKFSGDIREDLFRFAFESITNVIFGERLGMLEEIVDPEAQKFIDAVYQM 240 Query: 241 FHTSVPMLNLPPDLFRLFRTKTWKDHVAAWDVIFSKADIYTQNFYWELRQKGSVHHDYRG 300 FHTSVPMLNLPPDLFRLFRTKTW+DHVAAWD IF+KA+ YTQNFYW+LR+K ++Y G Sbjct: 241 FHTSVPMLNLPPDLFRLFRTKTWRDHVAAWDTIFNKAEKYTQNFYWDLRRKRE-FNNYPG 299 Query: 301 MLYRLLGDSKMSFEDIKANVTEMLAGGVDTTSMTLQWHLYEMARNLKVQDMLRAEVLAAR -

Multiple Sequence Alignment
Bioinformatics Algorithms Multiple Sequence Alignment David Hoksza http://siret.ms.mff.cuni.cz/hoksza Outline • Motivation • Algorithms • Scoring functions • exhaustive • multidimensional dynamic programming • heuristics • progressive alignment • iterative alignment/refinement • block(local)-based alignment Multiple sequence alignment (MSA) • Goal of MSA is to find “optimal” mapping of a set of sequences • Homologous residues (originating in the same position in a common ancestor) among a set of sequences are aligned together in columns • Usually employs multiple pairwise alignment (PA) computations to reveal the evolutionarily equivalent positions across all sequences Motivation • Distant homologues • faint similarity can become apparent when present in many sequences • motifs might not be apparent from pairwise alignment only • Detection of key functional residues • amino acids critical for function tend to be conserved during the evolution and therefore can be revealed by inspecting sequences within given family • Prediction of secondary/tertiary structure • Inferring evolutionary history 4 Representation of MSA • Column-based representation • Profile representation (position specific scoring matrix) • Sequence logo Manual MSA • High quality MSA can be carried out automatic MSA algorithms by hand using expert knowledge • specific columns • BAliBASE • highly conserved residues • https://lbgi.fr/balibase/ • buried hydrophobic residues • PROSITE • secondary structure (especially in RNA • http://prosite.expasy.org/ alignment) • Pfam • expected -

Practical Considerations of Working with Sequencing Data File Types
Practical considerations of working with sequencing data File Types • Fastq ->aligner -> reference(genome) coordinates • Coordinate files • SAM/BAM – most complete, contains all of the info in fastq and more! • Bedgraph – read density along the genome • Bed file –Read density reported in large continuous intervals • Genes/transcript and transcript structure • Transcription factor binding regions • If someone does a sequencing experiment usually one of these is available and deposited in a public database SAM/BAM Viewing genome coordinate files with IGV • Integrated Genome Browser • Cross-platform application • Knows about common genomes • Genome version is important! Different assemblies • Genome coordinates different between genome assemblies • Differences accumulate over chromosome length • You have to know which assembly was used • Sequencing files are non- randomly distributed relative to genes • RNAseq—should align with exons • TF binding sites—biased towards promoter regions Converting coordinates • UCSC liftOver -- converts genome coordinates • Convert from one assembly to another • Cross organism conversion • Mammals/vertebrates Sequence Alignment To do: • Global alignment • Local alignment • Scoring • Gaps • Scoring matrices • Database Search • Statistical Significance • Multiple Sequence alignment Why compare sequences • Given a new sequence, infer its function based on similarity to another sequence • Find important molecular regions – conserved across species • Determine 3d structure with homology modeling • Homologs-sequences that descended from a common ancestral sequence • Orthologs- separated by speciation • Paralogs separated by duplication in a single genome • Basic unit of protein homology is a sufficient functional unit—typically much smaller than a whole gene DNA vs Protein alignments • Protein coding • Typically compared in amino acid space • Amino acid change slower than nucleotides • Some nucleotides can change without any change to a.a. -

Amino Acid Substitution Matrices from Protein Blocks (Amino Add Sequence/Alignment Algorithms/Data Base Srching) STEVEN HENIKOFF* and JORJA G
Proc. Natl. Acad. Sci. USA Vol. 89, pp. 10915-10919, November 1992 Biochemistry Amino acid substitution matrices from protein blocks (amino add sequence/alignment algorithms/data base srching) STEVEN HENIKOFF* AND JORJA G. HENIKOFF Howard Hughes Medical Institute, Basic Sciences Division, Fred Hutchinson Cancer Research Center, Seattle, WA 98104 Communicated by Walter Gilbert, August 28, 1992 (receivedfor review July 13, 1992) ABSTRACT Methods for alignment of protein sequences new sequence and every other sequence in the block. For typically measure similait by using a substitution matrix with example, if the residue of the new sequence that aligns with scores for all possible exchanges of one amino acid with the first column of the first block is A and the column has 9 another. The most widely used matrices are based on the A residues and 1 S residue, then there are 9 AA matches and Dayhoff model of evolutionary rates. Using a different ap- 1 AS mismatch. This procedure is repeated for all columns of proach, we have derived substitution matrices from about 2000 all blocks with the summed results stored in a table. The new blocks of aligned sequence segments characterizing more than sequence is added to the group. For another new sequence, 500 groups of related proteins. This led to marked improve- the same procedure is followed, summing these numbers with ments in alignments and in searches using queries from each of those already in the table. Notice that successive addition of the groups. each sequence to the group leads to a table consisting of counts of all possible amino acid pairs in a column.