Regulation of Deubiquitinating Enzymes by Post-Translational Modifications
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Tyrosine Kinase – Role and Significance in Cancer
Int. J. Med. Sci. 2004 1(2): 101-115 101 International Journal of Medical Sciences ISSN 1449-1907 www.medsci.org 2004 1(2):101-115 ©2004 Ivyspring International Publisher. All rights reserved Review Tyrosine kinase – Role and significance in Cancer Received: 2004.3.30 Accepted: 2004.5.15 Manash K. Paul and Anup K. Mukhopadhyay Published:2004.6.01 Department of Biotechnology, National Institute of Pharmaceutical Education and Research, Sector-67, S.A.S Nagar, Mohali, Punjab, India-160062 Abstract Tyrosine kinases are important mediators of the signaling cascade, determining key roles in diverse biological processes like growth, differentiation, metabolism and apoptosis in response to external and internal stimuli. Recent advances have implicated the role of tyrosine kinases in the pathophysiology of cancer. Though their activity is tightly regulated in normal cells, they may acquire transforming functions due to mutation(s), overexpression and autocrine paracrine stimulation, leading to malignancy. Constitutive oncogenic activation in cancer cells can be blocked by selective tyrosine kinase inhibitors and thus considered as a promising approach for innovative genome based therapeutics. The modes of oncogenic activation and the different approaches for tyrosine kinase inhibition, like small molecule inhibitors, monoclonal antibodies, heat shock proteins, immunoconjugates, antisense and peptide drugs are reviewed in light of the important molecules. As angiogenesis is a major event in cancer growth and proliferation, tyrosine kinase inhibitors as a target for anti-angiogenesis can be aptly applied as a new mode of cancer therapy. The review concludes with a discussion on the application of modern techniques and knowledge of the kinome as means to gear up the tyrosine kinase drug discovery process. -

CYLD Is a Deubiquitinating Enzyme That Negatively Regulates NF-Kb
letters to nature 13. Schwartz, S. et al. Human–mouse alignments with BLASTZ. Genome Res 13, 103–107 (2003). necrosis factor receptors (TNFRs). Loss of the deubiquitinating 14. Schwartz, S. et al. MultiPipMaker and supporting tools: alignments and analysis of multiple genomic activity of CYLD correlates with tumorigenesis. CYLD inhibits DNA sequences. Nucleic Acids Res. 31, 3518–3524 (2003). 15.Murphy,W.J.et al. Resolution of the early placental mammal radiation using Bayesian phylogenetics. activation of NF-kB by the TNFR family members CD40, XEDAR Science 294, 2348–2351 (2001). and EDAR in a manner that depends on the deubiquitinating 16. Poux, C., Van Rheede, T., Madsen, O. & de Jong, W. W. Sequence gaps join mice and men: activity of CYLD. Downregulation of CYLD by RNA-mediated phylogenetic evidence from deletions in two proteins. Mol. Biol. Evol. 19, 2035–2037 (2002). 17. Huelsenbeck, J. P., Larget, B. & Swofford, D. A compound Poisson process for relaxing the molecular interference augments both basal and CD40-mediated activation clock. Genetics 154, 1879–1892 (2000). of NF-kB. The inhibition of NF-kBactivationbyCYLDis 18. Cooper, G. M. et al. Quantitative estimates of sequence divergence for comparative analyses of mediated, at least in part, by the deubiquitination and inacti- mammalian genomes. Genome Res. 13, 813–820 (2003). vation of TNFR-associated factor 2 (TRAF2) and, to a lesser 19. Siepel, A. & Haussler, D. Proc. 7th Annual Int. Conf. Research in Computational Molecular Biology (ACM, New York, 2003). extent, TRAF6. These results indicate that CYLD is a negative 20. Hardison, R. C. et al. Covariation in frequencies of substitution, deletion, transposition, and regulator of the cytokine-mediated activation of NF-kB that is recombination during eutherian evolution. -

A Drosophila Ortholog of the Human Cylindromatosis Tumor Suppressor
RESEARCH ARTICLE 2605 Development 134, 2605-2614 (2007) doi:10.1242/dev.02859 A Drosophila ortholog of the human cylindromatosis tumor suppressor gene regulates triglyceride content and antibacterial defense Theodore Tsichritzis1, Peer C. Gaentzsch3, Stylianos Kosmidis2, Anthony E. Brown3, Efthimios M. Skoulakis2, Petros Ligoxygakis3,* and George Mosialos1,4,* The cylindromatosis (CYLD) gene is mutated in human tumors of skin appendages. It encodes a deubiquitylating enzyme (CYLD) that is a negative regulator of the NF-B and JNK signaling pathways, in vitro. However, the tissue-specific function and regulation of CYLD in vivo are poorly understood. We established a genetically tractable animal model to initiate a systematic investigation of these issues by characterizing an ortholog of CYLD in Drosophila. Drosophila CYLD is broadly expressed during development and, in adult animals, is localized in the fat body, ovaries, testes, digestive tract and specific areas of the nervous system. We demonstrate that the protein product of Drosophila CYLD (CYLD), like its mammalian counterpart, is a deubiquitylating enzyme. Impairment of CYLD expression is associated with altered fat body morphology in adult flies, increased triglyceride levels and increased survival under starvation conditions. Furthermore, flies with compromised CYLD expression exhibited reduced resistance to bacterial infections. All mutant phenotypes described were reversible upon conditional expression of CYLD transgenes. Our results implicate CYLD in a broad range of functions associated with fat homeostasis and host defence in Drosophila. KEY WORDS: Cylindromatosis, Drosophila, Fat body, Host defense, NF-kappaB INTRODUCTION disease and it is required for the proper development of T Familial cylindromatosis is an autosomal-dominant predisposition lymphocytes in mice (Costello et al., 2005; Reiley et al., 2006). -

Principles of Protein Phosphorylation Biophysical Chemistry 1, Fall 2010
Principles of protein phosphorylation Biophysical Chemistry 1, Fall 2010 Signalling “cascades” Structural biology of phosphorylation Web assignment: http://pkr.genomics.purdue.edu Reversible protein phosphorylation Enzymatic reaction Posttranslational Control ΔG~12kcal/mol Kinases and phosphatases Phosphatase Kinase dephospho- phosphorylates rylates General Examples Signalling overview Cells are way too complex! MAPK/ERK Signaling Pathway RousExample:sarcoma virus Rous (RSV) sarcoma virus (RSV) gag - encodes capsid proteins pol - encodes reverse transcriptase env - encodes envelope proteins src - encodes a tyrosine kinase that attaches phosphate groups to the amino acid tyrosine in host cell proteins MutationsExample:, viruses Rous and cancer sarcoma virus (RSV) v-src lacks the C-terminal inhibitory phosphorylation site (tyrosine-527), and is therefore constitutively active as opposed to normal src (c-src) Continuous cell profileration tumor BiophysicsStructural of signalling Effect of Phosphorylation Phosphorylation is an important regulatory mechanism Can reversible attach/detach a phosphate and therefore switch “on”/”off” the function Effect of phosphorylation is manifold • Conformational change • Ordering/disordering • Electrostatic effects • Alternate binding behavior Signalling by reorientation Reorientation: A conformational switch Rmsd = 2.5Å DHP (red) and Z-Score = 4.6 DHPs74e (blue) (>3.6 same fold) Title: A phosphorylation-induced conformation change in dematin headpiece Author(s): Jiang ZHG, McKnight CJ Source: STRUCTURE Volume: 14 Issue: 2 Pages: 379-387 Published: FEB 2006 Order/disorderDisordering:transitions NtrC, a molecular switch upon phosphorylation Orange-yellow: unphosphorylated NtrC blue-cyan: phosphorylated NtrC Volkman et al., Science 2001, 291, 2429-33 Alternate Binding: SRC SH2 Src/SH2 interactions: binding vs release domain binding ExpectedCan conformationalwe understand effects (predict) the effect of phosphorylation Electrostatics Hydrogen bonding Size Mutational analysis Experimental data Lubman, O.Y. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Deubiquitinases in Cancer: New Functions and Therapeutic Options
Oncogene (2012) 31, 2373–2388 & 2012 Macmillan Publishers Limited All rights reserved 0950-9232/12 www.nature.com/onc REVIEW Deubiquitinases in cancer: new functions and therapeutic options JM Fraile1, V Quesada1, D Rodrı´guez, JMP Freije and C Lo´pez-Otı´n Departamento de Bioquı´mica y Biologı´a Molecular, Facultad de Medicina, Instituto Universitario de Oncologı´a, Universidad de Oviedo, Oviedo, Spain Deubiquitinases (DUBs) have fundamental roles in the Hunter, 2010). Consistent with the functional relevance ubiquitin system through their ability to specifically of proteases in these processes, alterations in their deconjugate ubiquitin from targeted proteins. The human structure or in the mechanisms controlling their genome encodes at least 98 DUBs, which can be grouped spatiotemporal expression patterns and activities cause into 6 families, reflecting the need for specificity in diverse pathologies such as arthritis, neurodegenerative their function. The activity of these enzymes affects the alterations, cardiovascular diseases and cancer. Accord- turnover rate, activation, recycling and localization ingly, many proteases are an important focus of of multiple proteins, which in turn is essential for attention for the pharmaceutical industry either as drug cell homeostasis, protein stability and a wide range of targets or as diagnostic and prognostic biomarkers signaling pathways. Consistent with this, altered DUB (Turk, 2006; Drag and Salvesen, 2010). function has been related to several diseases, including The recent availability of the genome sequence cancer. Thus, multiple DUBs have been classified as of different organisms has facilitated the identification oncogenes or tumor suppressors because of their regula- of their entire protease repertoire, which has been tory functions on the activity of other proteins involved in defined as degradome (Lopez-Otin and Overall, 2002). -

Design of a Highly Selective Quenched Activity-Based Probe
Article pubs.acs.org/JACS Design of a Highly Selective Quenched Activity-Based Probe and Its Application in Dual Color Imaging Studies of Cathepsin S Activity Localization † † † † ∥ Kristina Oresic Bender, Leslie Ofori, Wouter A. van der Linden, Elliot D. Mock, Gopal K. Datta, ∥ § † † ∥ # Somenath Chowdhury, Hao Li, Ehud Segal, Mateo Sanchez Lopez, Jonathan A. Ellman, , ⊥ † ‡ § † ⊥ Carl G. Figdor, Matthew Bogyo,*, , , and Martijn Verdoes*, , † ‡ § Departments of Pathology, Microbiology and Immunology, and Chemical and Systems Biology, Stanford University School of Medicine, Stanford, California 94305, United States ∥ Department of Chemistry, University of California-Berkeley, Berkeley, California 94720, United States ⊥ Department of Tumor Immunology, Radboud University Medical Center, Radboud Institute for Molecular Life Sciences, 6500 HB Nijmegen, The Netherlands *S Supporting Information ABSTRACT: The cysteine cathepsins are a group of 11 proteases whose function was originally believed to be the degradation of endocytosed material with a high degree of redundancy. However, it has become clear that these enzymes are also important regulators of both health and disease. Thus, selective tools that can discriminate between members of this highly related class of enzymes will be critical to further delineate the unique biological functions of individual cathepsins. Here we present the design and synthesis of a near-infrared quenched activity-based probe (qABP) that selectively targets cathepsin S which is highly expressed in immune cells. Importantly, this high degree of selectivity is retained both in vitro and in vivo. In combination with a new green-fluorescent pan-reactive cysteine cathepsin qABP we performed dual color labeling studies in bone marrow-derived immune cells and identified vesicles containing exclusively cathepsin S activity. -

Untying the Knot: Protein Quality Control in Inherited Cardiomyopathies
Pflügers Archiv - European Journal of Physiology https://doi.org/10.1007/s00424-018-2194-0 INVITED REVIEW Untying the knot: protein quality control in inherited cardiomyopathies Larissa M. Dorsch1 & Maike Schuldt1 & Dora Knežević1 & Marit Wiersma1 & Diederik W. D. Kuster1 & Jolanda van der Velden1 & Bianca J. J. M. Brundel1 Received: 30 July 2018 /Accepted: 6 August 2018 # The Author(s) 2018 Abstract Mutations in genes encoding sarcomeric proteins are the most important causes of inherited cardiomyopathies, which are a major cause of mortality and morbidity worldwide. Although genetic screening procedures for early disease detection have been improved significantly, treatment to prevent or delay mutation-induced cardiac disease onset is lacking. Recent findings indicate that loss of protein quality control (PQC) is a central factor in the disease pathology leading to derailment of cellular protein homeostasis. Loss of PQC includes impairment of heat shock proteins, the ubiquitin-proteasome system, and autophagy. This may result in accumulation of misfolded and aggregation-prone mutant proteins, loss of sarcomeric and cytoskeletal proteins, and, ultimately, loss of cardiac function. PQC derailment can be a direct effect of the mutation-induced activation, a compensa- tory mechanism due to mutation-induced cellular dysfunction or a consequence of the simultaneous occurrence of the mutation and a secondary hit. In this review, we discuss recent mechanistic findings on the role of proteostasis derailment in inherited cardiomyopathies, with special focus on sarcomeric gene mutations and possible therapeutic applications. Keywords Cardiomyopathy .Proteinqualitycontrol .Sarcomericmutation .Heatshockproteins .Ubiquitin-proteasomesystem . Autophagy Classification of cardiomyopathies occurring asymmetrically, and dilated CM (DCM), in which the presence of LV dilatation is accompanied by contractile Cardiomyopathies (CM) constitute one of the most common dysfunction [24]. -

Towards Therapy for Batten Disease
Towards therapy for Batten disease Mariana Catanho da Silva Vieira MRC Laboratory for Molecular Cell Biology University College London PhD Supervisor: Dr Sara E Mole A thesis submitted for the degree of Doctor of Philosophy University College London September 2014 Declaration I, Mariana Catanho da Silva Vieira, confirm that the work presented in this thesis is my own. Where information has been derived from other sources, I confirm that this has been indicated in the thesis. 2 Abstract The gene underlying the classic neurodegenerative lysosomal storage disorder (LSD) juvenile neuronal ceroid lipofuscinosis (JNCL) in humans, CLN3, encodes a polytopic membrane spanning protein of unknown function. Several studies using simpler models have been performed in order to further understand this protein and its pathological mechanism. Schizosaccharomyces pombe provides an ideal model organism for the study of CLN3 function, due to its simplicity, genetic tractability and the presence of a single orthologue of CLN3 (Btn1p), which exhibits a functional profile comparable to its human counterpart. In this study, this model was used to explore the effect of different mutations in btn1 as well as phenotypes arising from complete deletion of the gene. Different btn1 mutations have different effects on the protein function, underlining different phenotypes and affecting the levels of expression of Btn1p. So far, there is no cure for JNCL and therefore it is of great importance to identify novel lead compounds that can be developed for disease therapy. To identify these compounds, a drug screen with btn1Δ cells based on their sensitivity to cyclosporine A, was developed. Positive hits from the screen were validated and tested for their ability to rescue other specific phenotypes also associated with the loss of btn1. -

House Dust Mites: Ecology, Biology, Prevalence, Epidemiology and Elimination Muhammad Sarwar
Chapter House Dust Mites: Ecology, Biology, Prevalence, Epidemiology and Elimination Muhammad Sarwar Abstract House dust mites burrow cheerfully into our clothing, pillowcases, carpets, mats and furniture, and feed on human dead skin cells by breaking them into small particles for ingestion. Dust mites are most common in asthma allergens, and some people have a simple dust allergy, but others have an additional condition called atopic dermatitis, often stated to as eczema by reacting to mites with hideous itching and redness. The most common type of dust mites are Dermatophagoides farinae Hughes (American house dust mite) and Dermatophagoides pteronyssinus Trouessart (European house dust mite) of family Pyroglyphidae (Acari), which have been associated with dermatological and respiratory allergies in humans such as eczema and asthma. A typical house dust mite measures 0.2–0.3 mm and the body of mite has a striated cuticle. A mated female house dust mite can live up to 70 days and lays 60–100 eggs in the last 5 weeks of life, and an average life cycle is 65–100 days. In a 10-week life span, dust mite produces about 2000 fecal particles and an even larger number of partially digested enzyme-covered dust particles. They feed on skin flakes from animals, including humans and on some mold. Notably, mite’s gut contains potent digestive enzymes peptidase 1 that persist in their feces and are major induc- ers of allergic reactions, but its exoskeleton can also contribute this. Allergy testing by a physician can determine respiratory or dermatological symptoms to undergo allergen immunotherapy, by exposing to dust mite extracts for “training” immune system not to overreact. -

Pathways of Cellular Proteostasis in Aging and Disease
JCB: Review Pathways of cellular proteostasis in aging and disease Courtney L. Klaips, Gopal Gunanathan Jayaraj, and F. Ulrich Hartl Department of Cellular Biochemistry, Max Planck Institute of Biochemistry, Martinsried, Germany Ensuring cellular protein homeostasis, or proteostasis, re- control of abundance and subcellular localization, and finally, quires precise control of protein synthesis, folding, confor- disposal by degradation. A complex proteostasis network (PN) acts at each of these mational maintenance, and degradation. A complex and steps to maintain a balanced proteome linked by molecular adaptive proteostasis network coordinates these processes chaperones of different classes as central players. These factors with molecular chaperones of different classes and their ensure de novo folding in a crowded cellular environment and regulators functioning as major players. This network maintain proteins in a soluble, nonaggregated state. Moreover, in conditions that disfavor folding or solubility, certain chaper- serves to ensure that cells have the proteins they need ones act to target misfolded proteins for degradation or spatial while minimizing misfolding or aggregation events that sequestration, thus protecting the rest of the proteome from ab- are hallmarks of age-associated proteinopathies, includ- errant interactions (Balchin et al., 2016; Sontag et al., 2017). Here, we describe the major pathways of cellular pro- ing neurodegenerative disorders such as Alzheimer’s and teostasis and outline the challenges they face during aging and Parkinson’s diseases. It is now clear that the capacity of disease. We exemplify these processes using mainly the proteo- cells to maintain proteostasis undergoes a decline during stasis pathways operating in the cytosol, where most cellular aging, rendering the organism susceptible to these pa- proteins are produced. -
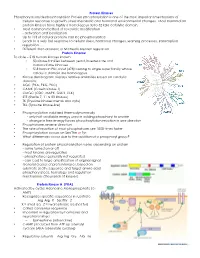
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off