Leiomodin-3 Dysfunction Results in Thin Filament Disorganization and Nemaline Myopathy
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Compound Heterozygosity for Novel Truncating Variants in the LMOD3 Gene As the Cause of Polyhydramnios in Two Successive Fetuses
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Online Research @ Cardiff CASE REPORT published: 13 September 2019 doi: 10.3389/fgene.2019.00835 Compound Heterozygosity for Novel Truncating Variants in the LMOD3 Gene as the Cause of Polyhydramnios in Two Successive Fetuses Ye Wang 1, Caixia Zhu 1, Liu Du 2, Qiaoer Li 3, Mei-Fang Lin 2, Claude Férec 4,5, David N. Cooper 6, Jian-Min Chen 4† and Yi Zhou 1*† 1 Fetal Medicine Center, Department of Obstetrics and Gynecology, The First Affiliated Hospital of Sun Yat-Sen University, Guangzhou, China, 2 Department of Ultrasonic Medicine, The First Affiliated Hospital of Sun Yat-Sen University, Guangzhou, China, 3 Jiangmen Central Hospital, Affiliated Jiangmen Hospital of Sun Yat-Sen University, Jiangmen, China, 4 EFS, Univ Brest, Inserm, UMR 1078, GGB, Brest, France, 5 CHU Brest, Service de Génétique, Brest, France, 6Institute of Medical Genetics, School of Medicine, Cardiff University, Cardiff, United Kingdom Polyhydramnios is sometimes associated with genetic defects. However, establishing an Edited by: Fan Jin, accurate diagnosis and pinpointing the precise genetic cause of polyhydramnios in any Zhejiang University, China given case represents a major challenge because it is known to occur in association with Reviewed by: over 200 different conditions. Whole exome sequencing (WES) is now a routine part of Liang-Liang Fan, Central South University, China the clinical workup, particularly with diseases characterized by atypical manifestations and Wenbin Zou, significant genetic heterogeneity. Here we describe the identification, by means of WES, Changhai Hospital, China of novel compound heterozygous truncating variants in the LMOD3 gene [i.e., c.1412delA *Correspondence: (p.Lys471Serfs*18) and c.1283dupC (p.Gly429Trpfs*35)] in a Chinese family with two Yi Zhou [email protected] successive fetuses affected with polyhydramnios, thereby potentiating the prenatal diagnosis of nemaline myopathy (NM) in the proband. -

Loss of LMOD1 Impairs Smooth Muscle Cytocontractility And
Loss of LMOD1 impairs smooth muscle cytocontractility PNAS PLUS and causes megacystis microcolon intestinal hypoperistalsis syndrome in humans and mice Danny Halima,1, Michael P. Wilsonb,1, Daniel Oliverc, Erwin Brosensa, Joke B. G. M. Verheijd, Yu Hanb, Vivek Nandab, Qing Lyub, Michael Doukase, Hans Stoope, Rutger W. W. Brouwerf, Wilfred F. J. van IJckenf, Orazio J. Slivanob, Alan J. Burnsa,g, Christine K. Christieb, Karen L. de Mesy Bentleyh, Alice S. Brooksa, Dick Tibboeli, Suowen Xub, Zheng Gen Jinb, Tono Djuwantonoj, Wei Yanc, Maria M. Alvesa, Robert M. W. Hofstraa,g,2, and Joseph M. Mianob,2 aDepartment of Clinical Genetics, Erasmus University Medical Center, 3015 CN Rotterdam, The Netherlands; bAab Cardiovascular Research Institute, University of Rochester School of Medicine and Dentistry, Rochester, NY 14642; cDepartment of Physiology and Cell Biology, University of Nevada School of Medicine, Reno, NV 89557; dDepartment of Genetics, University Medical Center, University of Groningen, 9700 RB Groningen, The Netherlands; eDepartment of Pathology, Erasmus University Medical Center, 3015 CN Rotterdam, The Netherlands; fCenter for Biomics, Erasmus University Medical Center, 3015 CN Rotterdam, The Netherlands; gStem Cells and Regenerative Medicine, Birth Defects Research Centre, University College London Institute of Child Health, London WC1N 1EH, United Kingdom; hDepartment of Pathology and Laboratory Medicine, University of Rochester School of Medicine and Dentistry, Rochester, NY 14642; iDepartment of Pediatric Surgery, Erasmus University Medical Center, 3015 CN Rotterdam, The Netherlands; and jDepartment of Obstetrics and Gynecology, Faculty of Medicine, Universitas Padjadjaran, Bandung, Indonesia Edited by Eric N. Olson, University of Texas Southwestern Medical Center, Dallas, TX, and approved February 21, 2017 (received for review December 13, 2016) Megacystis microcolon intestinal hypoperistalsis syndrome (MMIHS) is lacking (6, 7). -

A Long-Read RNA-Seq Approach to Identify Novel Transcripts of Very Large Genes
Downloaded from genome.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Method A long-read RNA-seq approach to identify novel transcripts of very large genes Prech Uapinyoying,1,2,3,6 Jeremy Goecks,4 Susan M. Knoblach,1,2 Karuna Panchapakesan,1 Carsten G. Bonnemann,1,3 Terence A. Partridge,1,2 Jyoti K. Jaiswal,1,2 and Eric P. Hoffman1,5 1Center for Genetic Medicine Research, Children’s Research Institute, Children’s National Health System, Washington, D.C. 20010, USA; 2Department of Genomics and Precision Medicine, The George Washington University School of Medicine and Health Sciences, Washington, D.C. 20052, USA; 3Neuromuscular and Neurogenetic Disorders of Childhood Section, National Institute of Neurological Disorders and Stroke, National Institutes of Health, Bethesda, Maryland 20892, USA; 4Computational Biology Program, Oregon Health and Science University, Portland, Oregon 97239, USA; 5Department of Pharmaceutical Sciences, School of Pharmacy and Pharmaceutical Sciences, Binghamton University, Binghamton, New York 13902, USA RNA-seq is widely used for studying gene expression, but commonly used sequencing platforms produce short reads that only span up to two exon junctions per read. This makes it difficult to accurately determine the composition and phasing of exons within transcripts. Although long-read sequencing improves this issue, it is not amenable to precise quantitation, which limits its utility for differential expression studies. We used long-read isoform sequencing combined with a novel anal- ysis approach to compare alternative splicing of large, repetitive structural genes in muscles. Analysis of muscle structural genes that produce medium (Nrap: 5 kb), large (Neb: 22 kb), and very large (Ttn: 106 kb) transcripts in cardiac muscle, and fast and slow skeletal muscles identified unannotated exons for each of these ubiquitous muscle genes. -

Global MEF2 Target Gene Analysis In
Published online 12 September 2014 Nucleic Acids Research, 2014, Vol. 42, No. 18 11349–11362 doi: 10.1093/nar/gku813 Global MEF2 target gene analysis in cardiac and skeletal muscle reveals novel regulation of DUSP6 by p38MAPK-MEF2 signaling Stephanie Wales1,2,3, Sara Hashemi1,2,3, Alexandre Blais4 and John C. McDermott1,2,3,5,* 1Department of Biology, York University, 4700 Keele Street Toronto, Ontario, M3J 1P3 Canada, 2Muscle Health Research Centre (MHRC), York University, 4700 Keele Street, Toronto, Ontario, M3J 1P3 Canada, 3Centre for Research on Biomolecular Interactions (CRBI), 4700 Keele Street, Toronto, Ontario, M3J 1P3 Canada, 4Ottawa Institute of Systems Biology, University of Ottawa, Health Sciences Campus, 451 Smyth Road, Ottawa, Ontario, K1H 8M5 Canada and 5Centre for Research in Mass Spectrometry (CRMS), York University, 4700 Keele Street, Toronto, Ontario, M3J 1P3 Canada Downloaded from Received June 12, 2014; Revised August 23, 2014; Accepted August 28, 2014 ABSTRACT INTRODUCTION http://nar.oxfordjournals.org/ MEF2 plays a profound role in the regulation of tran- Myocyte enhancer factor-2 (MEF2) is a member of the scription in cardiac and skeletal muscle lineages. MADS-box super family of transcriptional regulatory pro- To define the overlapping and unique MEF2A ge- teins originally identified in skeletal muscle but are now nomic targets, we utilized ChIP-exo analysis of car- an established component in the regulation of a diverse diomyocytes and skeletal myoblasts. Of the 2783 and number of tissues, including smooth, cardiac and skele- tal muscle, neurons and T cells (1–4). In vertebrates 1648 MEF2A binding peaks in skeletal myoblasts and there are four MEF2 isoforms (A–D) which bind to the cardiomyocytes, respectively, 294 common binding consensus sequence (C/T TA(A/T)4TA G/A) within the sites were identified. -

Human Social Genomics in the Multi-Ethnic Study of Atherosclerosis
Getting “Under the Skin”: Human Social Genomics in the Multi-Ethnic Study of Atherosclerosis by Kristen Monét Brown A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Epidemiological Science) in the University of Michigan 2017 Doctoral Committee: Professor Ana V. Diez-Roux, Co-Chair, Drexel University Professor Sharon R. Kardia, Co-Chair Professor Bhramar Mukherjee Assistant Professor Belinda Needham Assistant Professor Jennifer A. Smith © Kristen Monét Brown, 2017 [email protected] ORCID iD: 0000-0002-9955-0568 Dedication I dedicate this dissertation to my grandmother, Gertrude Delores Hampton. Nanny, no one wanted to see me become “Dr. Brown” more than you. I know that you are standing over the bannister of heaven smiling and beaming with pride. I love you more than my words could ever fully express. ii Acknowledgements First, I give honor to God, who is the head of my life. Truly, without Him, none of this would be possible. Countless times throughout this doctoral journey I have relied my favorite scripture, “And we know that all things work together for good, to them that love God, to them who are called according to His purpose (Romans 8:28).” Secondly, I acknowledge my parents, James and Marilyn Brown. From an early age, you two instilled in me the value of education and have been my biggest cheerleaders throughout my entire life. I thank you for your unconditional love, encouragement, sacrifices, and support. I would not be here today without you. I truly thank God that out of the all of the people in the world that He could have chosen to be my parents, that He chose the two of you. -

Investigating the Effect of Chronic Activation of AMP-Activated Protein
Investigating the effect of chronic activation of AMP-activated protein kinase in vivo Alice Pollard CASE Studentship Award A thesis submitted to Imperial College London for the degree of Doctor of Philosophy September 2017 Cellular Stress Group Medical Research Council London Institute of Medical Sciences Imperial College London 1 Declaration I declare that the work presented in this thesis is my own, and that where information has been derived from the published or unpublished work of others it has been acknowledged in the text and in the list of references. This work has not been submitted to any other university or institute of tertiary education in any form. Alice Pollard The copyright of this thesis rests with the author and is made available under a Creative Commons Attribution Non-Commercial No Derivatives license. Researchers are free to copy, distribute or transmit the thesis on the condition that they attribute it, that they do not use it for commercial purposes and that they do not alter, transform or build upon it. For any reuse or redistribution, researchers must make clear to others the license terms of this work. 2 Abstract The prevalence of obesity and associated diseases has increased significantly in the last decade, and is now a major public health concern. It is a significant risk factor for many diseases, including cardiovascular disease (CVD) and type 2 diabetes. Characterised by excess lipid accumulation in the white adipose tissue, which drives many associated pathologies, obesity is caused by chronic, whole-organism energy imbalance; when caloric intake exceeds energy expenditure. Whilst lifestyle changes remain the most effective treatment for obesity and the associated metabolic syndrome, incidence continues to rise, particularly amongst children, placing significant strain on healthcare systems, as well as financial burden. -

Agricultural University of Athens
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΤΩΝ ΖΩΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΖΩΙΚΗΣ ΠΑΡΑΓΩΓΗΣ ΕΡΓΑΣΤΗΡΙΟ ΓΕΝΙΚΗΣ ΚΑΙ ΕΙΔΙΚΗΣ ΖΩΟΤΕΧΝΙΑΣ ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ ΑΘΗΝΑ 2020 ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Genome-wide association analysis and gene network analysis for (re)production traits in commercial broilers ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ Τριμελής Επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Επταμελής εξεταστική επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Πηνελόπη Μπεμπέλη (Καθ. ΓΠΑ) Δημήτριος Βλαχάκης (Επ. Καθ. ΓΠΑ) Ευάγγελος Ζωίδης (Επ.Καθ. ΓΠΑ) Γεώργιος Θεοδώρου (Επ.Καθ. ΓΠΑ) 2 Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Περίληψη Σκοπός της παρούσας διδακτορικής διατριβής ήταν ο εντοπισμός γενετικών δεικτών και υποψηφίων γονιδίων που εμπλέκονται στο γενετικό έλεγχο δύο τυπικών πολυγονιδιακών ιδιοτήτων σε κρεοπαραγωγικά ορνίθια. Μία ιδιότητα σχετίζεται με την ανάπτυξη (σωματικό βάρος στις 35 ημέρες, ΣΒ) και η άλλη με την αναπαραγωγική -

UC San Diego Electronic Theses and Dissertations
UC San Diego UC San Diego Electronic Theses and Dissertations Title Cardiac Stretch-Induced Transcriptomic Changes are Axis-Dependent Permalink https://escholarship.org/uc/item/7m04f0b0 Author Buchholz, Kyle Stephen Publication Date 2016 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California UNIVERSITY OF CALIFORNIA, SAN DIEGO Cardiac Stretch-Induced Transcriptomic Changes are Axis-Dependent A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Bioengineering by Kyle Stephen Buchholz Committee in Charge: Professor Jeffrey Omens, Chair Professor Andrew McCulloch, Co-Chair Professor Ju Chen Professor Karen Christman Professor Robert Ross Professor Alexander Zambon 2016 Copyright Kyle Stephen Buchholz, 2016 All rights reserved Signature Page The Dissertation of Kyle Stephen Buchholz is approved and it is acceptable in quality and form for publication on microfilm and electronically: Co-Chair Chair University of California, San Diego 2016 iii Dedication To my beautiful wife, Rhia. iv Table of Contents Signature Page ................................................................................................................... iii Dedication .......................................................................................................................... iv Table of Contents ................................................................................................................ v List of Figures ................................................................................................................... -

Autosomal Dominant Familial Dyskinesia and Facial Myokymia Single Exome Sequencing Identifies a Mutation in Adenylyl Cyclase 5
ORIGINAL CONTRIBUTION Autosomal Dominant Familial Dyskinesia and Facial Myokymia Single Exome Sequencing Identifies a Mutation in Adenylyl Cyclase 5 Ying-Zhang Chen, MD, PhD; Mark M. Matsushita, BS; Peggy Robertson, PhD; Mark Rieder, PhD; Santhosh Girirajan, MBBS, PhD; Francesca Antonacci, PhD; Hillary Lipe, MN, ARNP; Evan E. Eichler, PhD; Deborah A. Nickerson, PhD; Thomas D. Bird, MD; Wendy H. Raskind, MD, PhD Background: Familial dyskinesia with facial myoky- Results: The exome contained 23 428 single-nucleotide mia (FDFM) is an autosomal dominant disorder that is ex- variants, of which 9391 were missense, nonsense, or splice acerbated by anxiety. In a 5-generation family of German site alterations. The critical region contained 323 variants, ancestry, we previously mapped FDFM to chromosome 5 of which were not present in 1 of the sequence databases. band 3p21-3q21. The 72.5-Mb linkage region was too large Adenylyl cyclase 5 (ADCY5) was the only gene in which for traditional positional mutation identification. the variant (c.2176GϾA) was co-transmitted perfectly with disease status and was not present in 3510 control white Objective: To identify the gene responsible for FDFM exomes. This residue is highly conserved, and the change by exome resequencing of a single affected individual. is nonconservative and predicted to be damaging. Participants: We performed whole exome sequencing Conclusions: ADCY5 is highly expressed in striatum. in 1 affected individual and used a series of bioinformatic Mice deficient in Adcy5 develop a movement disorder that filters, including functional significance and presence in is worsened by stress. We conclude that FDFM likely re- dbSNP or the 1000 Genomes Project, to reduce the num- sults from a missense mutation in ADCY5. -

Identification of Acquired Copy Number Alterations and Uniparental
Leukemia (2010) 24, 438–449 & 2010 Macmillan Publishers Limited All rights reserved 0887-6924/10 $32.00 www.nature.com/leu ORIGINAL ARTICLE Identification of acquired copy number alterations and uniparental disomies in cytogenetically normal acute myeloid leukemia using high-resolution single-nucleotide polymorphism analysis L Bullinger1,4, J Kro¨nke1,4, C Scho¨n1,4, I Radtke2, K Urlbauer1, U Botzenhardt1, V Gaidzik1, A Cario´ 1, C Senger1, RF Schlenk1, JR Downing2, K Holzmann3,KDo¨hner1 and H Do¨hner1 1Department of Internal Medicine III, University Hospital of Ulm, Ulm, Germany; 2Department of Pathology, St Jude Children’s Research Hospital, Memphis, TN, USA and 3Microarray Core Facility, University of Ulm, Ulm, Germany Recent advances in genome-wide single-nucleotide polymorph- tyrosine kinase 3 (FLT3) gene the CCAAT/enhancer binding ism (SNP) analyses have revealed previously unrecognized protein-a (CEBPA) gene, the myeloid-lymphoid or mixed-line- microdeletions and uniparental disomy (UPD) in a broad age leukemia (MLL) gene, the neuroblastoma RAS viral spectrum of human cancers. As acute myeloid leukemia (AML) represents a genetically heterogeneous disease, this oncogene homolog (NRAS) gene, the Wilms’ tumor 1 (WT1) technology might prove helpful, especially for cytogenetically gene, and the runt-related transcription factor 1 (RUNX1) 4,5 normal AML (CN-AML) cases. Thus, we performed high- gene. These aberrations underscore the genetic complexity resolution SNP analyses in 157 adult cases of CN-AML. Regions of CN-AML and many additional genetic lesions are expected to of acquired UPDs were identified in 12% of cases and in the be present in this subset of AML. -
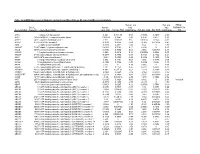
Mrna Expression in Human Leiomyoma and Eker Rats As Measured by Microarray Analysis
Table 3S: mRNA Expression in Human Leiomyoma and Eker Rats as Measured by Microarray Analysis Human_avg Rat_avg_ PENG_ Entrez. Human_ log2_ log2_ RAPAMYCIN Gene.Symbol Gene.ID Gene Description avg_tstat Human_FDR foldChange Rat_avg_tstat Rat_FDR foldChange _DN A1BG 1 alpha-1-B glycoprotein 4.982 9.52E-05 0.68 -0.8346 0.4639 -0.38 A1CF 29974 APOBEC1 complementation factor -0.08024 0.9541 -0.02 0.9141 0.421 0.10 A2BP1 54715 ataxin 2-binding protein 1 2.811 0.01093 0.65 0.07114 0.954 -0.01 A2LD1 87769 AIG2-like domain 1 -0.3033 0.8056 -0.09 -3.365 0.005704 -0.42 A2M 2 alpha-2-macroglobulin -0.8113 0.4691 -0.03 6.02 0 1.75 A4GALT 53947 alpha 1,4-galactosyltransferase 0.4383 0.7128 0.11 6.304 0 2.30 AACS 65985 acetoacetyl-CoA synthetase 0.3595 0.7664 0.03 3.534 0.00388 0.38 AADAC 13 arylacetamide deacetylase (esterase) 0.569 0.6216 0.16 0.005588 0.9968 0.00 AADAT 51166 aminoadipate aminotransferase -0.9577 0.3876 -0.11 0.8123 0.4752 0.24 AAK1 22848 AP2 associated kinase 1 -1.261 0.2505 -0.25 0.8232 0.4689 0.12 AAMP 14 angio-associated, migratory cell protein 0.873 0.4351 0.07 1.656 0.1476 0.06 AANAT 15 arylalkylamine N-acetyltransferase -0.3998 0.7394 -0.08 0.8486 0.456 0.18 AARS 16 alanyl-tRNA synthetase 5.517 0 0.34 8.616 0 0.69 AARS2 57505 alanyl-tRNA synthetase 2, mitochondrial (putative) 1.701 0.1158 0.35 0.5011 0.6622 0.07 AARSD1 80755 alanyl-tRNA synthetase domain containing 1 4.403 9.52E-05 0.52 1.279 0.2609 0.13 AASDH 132949 aminoadipate-semialdehyde dehydrogenase -0.8921 0.4247 -0.12 -2.564 0.02993 -0.32 AASDHPPT 60496 aminoadipate-semialdehyde -
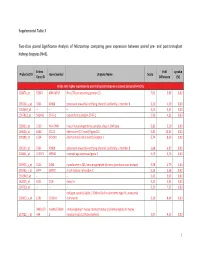
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at