Interdependence of EGFR with Ptps on Juxtaposed Membranes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Clinical and Biological Features of PTPN2-Deleted Adult and Pediatric T-Cell Acute Lymphoblastic Leukemia
REGULAR ARTICLE Clinical and biological features of PTPN2-deleted adult and pediatric T-cell acute lymphoblastic leukemia Marion Alcantara,1 Mathieu Simonin,1-3 Ludovic Lhermitte,1 Aurore Touzart,1 Marie Emilie Dourthe,1,4 Mehdi Latiri,1 Nathalie Grardel,5 Jean Michel Cayuela,6 Yves Chalandon,7 Carlos Graux,8 Herve´ Dombret,9 Norbert Ifrah,10 Arnaud Petit,2,3 Elizabeth Macintyre,1 Andre´ Baruchel,4 Nicolas Boissel,9 and Vahid Asnafi1 1Universite´ Paris Descartes Sorbonne Cite,´ Institut Necker-Enfants Malades, Institut National de Recherche Medicale´ U1151, Laboratory of Onco-Hematology, Assistance Publique–Hopitauxˆ de Paris, Hopitalˆ Necker Enfants-Malades, Paris, France; 2Department of Pediatric Hematology and Oncology, Assistance Publique–Hopitauxˆ de Paris, Groupe Hospitalier des Hopitauxˆ Universitaires Est Parisien, Armand Trousseau Hospital, Paris, France; 3Sorbonne Universites,´ Universite´ Pierre et Marie Curie Universite´ Paris 06, Unite´ Mixte de Recherche S 938, Centre de Recherche Saint-Antoine, Groupe de Recherche (GRC) Clinique n°07, GRC Myeloprolif´ erations´ aigues¨ et chroniques, Paris, France; 4Department of Pediatric Hematology and Immunology, Assistance Publique–Hopitauxˆ de Paris, Hopitalˆ Universitaire Robert-Debre,´ Paris, France; 5Laboratory of Hematology, Centre Hospitalier Regional´ Universitaire de Lille, Lille, France; 6Laboratory of Hematology, Saint-Louis Hospital, Assistance Publique–Hopitauxˆ de Paris, Paris, France; 7Swiss Group for Clinical Cancer Research, Bern, Hopitalˆ Universitaire, Geneva, Switzerland; 8Universite´ Catholique de Louvain, Centre Hospitalier Universitaire, Namur, Yvoir, Belgium; 9Adolescent and Young Adult Hematology Unit, Saint-Louis Hospital, Assistance Publique–Hopitauxˆ de Paris, Paris, France; and 10Centre Hospitalier Universitaire d’Angers, Angers, France Protein tyrosine phosphatase nonreceptor type 2 (PTPN2) is a phosphatase known to be Key Points a tumor suppressor gene in T-cell acute lymphoblastic leukemia (T-ALL). -

Inflammatory Cytokine Signalling by Protein Tyrosine Phosphatases in Pancreatic Β-Cells
59 4 W J STANLEY and others PTPN1 and PTPN6 modulate 59: 4 325–337 Research cytokine signalling in β-cells Differential regulation of pro- inflammatory cytokine signalling by protein tyrosine phosphatases in pancreatic β-cells William J Stanley1,2, Prerak M Trivedi1,2, Andrew P Sutherland1, Helen E Thomas1,2 and Esteban N Gurzov1,2,3 Correspondence should be addressed 1 St. Vincent’s Institute of Medical Research, Melbourne, Australia to E N Gurzov 2 Department of Medicine, St. Vincent’s Hospital, The University of Melbourne, Melbourne, Australia Email 3 ULB Center for Diabetes Research, Universite Libre de Bruxelles (ULB), Brussels, Belgium esteban.gurzov@unimelb. edu.au Abstract Type 1 diabetes (T1D) is characterized by the destruction of insulin-producing β-cells Key Words by immune cells in the pancreas. Pro-inflammatory including TNF-α, IFN-γ and IL-1β f pancreatic β-cells are released in the islet during the autoimmune assault and signal in β-cells through f protein tyrosine phosphorylation cascades, resulting in pro-apoptotic gene expression and eventually phosphatases β-cell death. Protein tyrosine phosphatases (PTPs) are a family of enzymes that regulate f PTPN1 phosphorylative signalling and are associated with the development of T1D. Here, we f PTPN6 observed expression of PTPN6 and PTPN1 in human islets and islets from non-obese f cytokines diabetic (NOD) mice. To clarify the role of these PTPs in β-cells/islets, we took advantage f inflammation Journal of Molecular Endocrinology of CRISPR/Cas9 technology and pharmacological approaches to inactivate both proteins. We identify PTPN6 as a negative regulator of TNF-α-induced β-cell death, through JNK- dependent BCL-2 protein degradation. -

The Proteintyrosine Phosphatase Receptor Type J Is Regulated by The
Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2013 The Protein-Tyrosine Phosphatase Receptor Type J is regulated by the pVHL-HIF axis in Clear Cell Renal Cell Carcinoma Casagrande, Silvia ; Ruf, Melanie ; Rechsteiner, Markus ; Morra, Laura ; Brun-Schmid, Sonja ; von Teichman, Adriana ; Krek, Wilhelm ; Schraml, Peter ; Moch, Holger Abstract: Mass spectrometry analysis of renal cancer cell lines recently suggested that the Protein- Tyrosine Phosphatase Receptor Type J (PTPRJ), an important regulator of tyrosine kinase receptors, is tightly linked to the von-Hippel Lindau protein (pVHL). Therefore, we aimed to characterize the biological relevance of PTPRJ for clear cell renal cell carcinoma (ccRCC). In pVHL negative ccRCC cell lines both RNA and protein expression levels of PTPRJ were lower than those in the corresponding pVHL reconstituted cells. Quantitative RT-PCR and Western blot analysis of ccRCC with known VHL mutation status and normal matched tissues as well as RNA in situ hybridization on a Tissue Microarray (TMA) confirmed a decrease of PTPRJ expression in more than 80 % of ccRCCs, but inonly12%of papillary RCCs. ccRCC patients with no or reduced PTPRJ mRNA expression had a less favourable outcome than those with a normal expression status (p = 0.05). Sequence analysis of 32 PTPRJ mRNA negative ccRCC samples showed five known polymorphisms, but no mutations implying other mechanisms leading to PTPRJ’s down-regulation. Selective silencing of HIF- by siRNA and reporter gene assays demonstrated that pVHL inactivation reduces PTPRJ expression through a HIF-dependent mechanism, which is mainly driven by HIF-2 stabilization. -

The Expression Patterns and the Prognostic Roles of PTPN Family Members in Digestive Tract Cancers
Preprint: Please note that this article has not completed peer review. The expression patterns and the prognostic roles of PTPN family members in digestive tract cancers CURRENT STATUS: UNDER REVIEW Jing Chen The First Affiliated Hospital of China Medical University Xu Zhao Liaoning Vocational College of Medicine Yuan Yuan The First Affiliated Hospital of China Medical University Jing-jing Jing The First Affiliated Hospital of China Medical University [email protected] Author ORCiD: https://orcid.org/0000-0002-9807-8089 DOI: 10.21203/rs.3.rs-19689/v1 SUBJECT AREAS Cancer Biology KEYWORDS PTPN family members, digestive tract cancers, expression, prognosis, clinical features 1 Abstract Background Non-receptor protein tyrosine phosphatases (PTPNs) are a set of enzymes involved in the tyrosyl phosphorylation. The present study intended to clarify the associations between the expression patterns of PTPN family members and the prognosis of digestive tract cancers. Method Expression profiling of PTPN family genes in digestive tract cancers were analyzed through ONCOMINE and UALCAN. Gene ontology enrichment analysis was conducted using the DAVID database. The gene–gene interaction network was performed by GeneMANIA and the protein–protein interaction (PPI) network was built using STRING portal couple with Cytoscape. Data from The Cancer Genome Atlas (TCGA) were downloaded for validation and to explore the relationship of the PTPN expression with clinicopathological parameters and survival of digestive tract cancers. Results Most PTPN family members were associated with digestive tract cancers according to Oncomine, Ualcan and TCGA data. For esophageal carcinoma (ESCA), expression of PTPN1, PTPN4 and PTPN12 were upregulated; expression of PTPN20 was associated with poor prognosis. -

The Regulatory Roles of Phosphatases in Cancer
Oncogene (2014) 33, 939–953 & 2014 Macmillan Publishers Limited All rights reserved 0950-9232/14 www.nature.com/onc REVIEW The regulatory roles of phosphatases in cancer J Stebbing1, LC Lit1, H Zhang, RS Darrington, O Melaiu, B Rudraraju and G Giamas The relevance of potentially reversible post-translational modifications required for controlling cellular processes in cancer is one of the most thriving arenas of cellular and molecular biology. Any alteration in the balanced equilibrium between kinases and phosphatases may result in development and progression of various diseases, including different types of cancer, though phosphatases are relatively under-studied. Loss of phosphatases such as PTEN (phosphatase and tensin homologue deleted on chromosome 10), a known tumour suppressor, across tumour types lends credence to the development of phosphatidylinositol 3--kinase inhibitors alongside the use of phosphatase expression as a biomarker, though phase 3 trial data are lacking. In this review, we give an updated report on phosphatase dysregulation linked to organ-specific malignancies. Oncogene (2014) 33, 939–953; doi:10.1038/onc.2013.80; published online 18 March 2013 Keywords: cancer; phosphatases; solid tumours GASTROINTESTINAL MALIGNANCIES abs in sera were significantly associated with poor survival in Oesophageal cancer advanced ESCC, suggesting that they may have a clinical utility in Loss of PTEN (phosphatase and tensin homologue deleted on ESCC screening and diagnosis.5 chromosome 10) expression in oesophageal cancer is frequent, Cao et al.6 investigated the role of protein tyrosine phosphatase, among other gene alterations characterizing this disease. Zhou non-receptor type 12 (PTPN12) in ESCC and showed that PTPN12 et al.1 found that overexpression of PTEN suppresses growth and protein expression is higher in normal para-cancerous tissues than induces apoptosis in oesophageal cancer cell lines, through in 20 ESCC tissues. -

(12) Patent Application Publication (10) Pub. No.: US 2003/0082511 A1 Brown Et Al
US 20030082511A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2003/0082511 A1 Brown et al. (43) Pub. Date: May 1, 2003 (54) IDENTIFICATION OF MODULATORY Publication Classification MOLECULES USING INDUCIBLE PROMOTERS (51) Int. Cl." ............................... C12O 1/00; C12O 1/68 (52) U.S. Cl. ..................................................... 435/4; 435/6 (76) Inventors: Steven J. Brown, San Diego, CA (US); Damien J. Dunnington, San Diego, CA (US); Imran Clark, San Diego, CA (57) ABSTRACT (US) Correspondence Address: Methods for identifying an ion channel modulator, a target David B. Waller & Associates membrane receptor modulator molecule, and other modula 5677 Oberlin Drive tory molecules are disclosed, as well as cells and vectors for Suit 214 use in those methods. A polynucleotide encoding target is San Diego, CA 92121 (US) provided in a cell under control of an inducible promoter, and candidate modulatory molecules are contacted with the (21) Appl. No.: 09/965,201 cell after induction of the promoter to ascertain whether a change in a measurable physiological parameter occurs as a (22) Filed: Sep. 25, 2001 result of the candidate modulatory molecule. Patent Application Publication May 1, 2003 Sheet 1 of 8 US 2003/0082511 A1 KCNC1 cDNA F.G. 1 Patent Application Publication May 1, 2003 Sheet 2 of 8 US 2003/0082511 A1 49 - -9 G C EH H EH N t R M h so as se W M M MP N FIG.2 Patent Application Publication May 1, 2003 Sheet 3 of 8 US 2003/0082511 A1 FG. 3 Patent Application Publication May 1, 2003 Sheet 4 of 8 US 2003/0082511 A1 KCNC1 ITREXCHO KC 150 mM KC 2000000 so 100 mM induced Uninduced Steady state O 100 200 300 400 500 600 700 Time (seconds) FIG. -

Protein Tyrosine Phosphatase Non-Receptor Type 2 Function in Dendritic Cells Is Crucial to Maintain Tissue Tolerance
ORIGINAL RESEARCH published: 18 August 2020 doi: 10.3389/fimmu.2020.01856 Protein Tyrosine Phosphatase Non-Receptor Type 2 Function in Dendritic Cells Is Crucial to Maintain Tissue Tolerance Larissa Hering 1, Egle Katkeviciute 1, Marlene Schwarzfischer 1, Philipp Busenhart 1, Claudia Gottier 1, Dunja Mrdjen 2, Juliana Komuczki 2†, Marcin Wawrzyniak 1, Silvia Lang 1, Kirstin Atrott 1, Burkhard Becher 2, Gerhard Rogler 1,3, Michael Scharl 1,3* and Marianne R. Spalinger 1 1 Department of Gastroenterology and Hepatology, University Hospital Zurich, University of Zurich, Zurich, Switzerland, 2 Institute of Experimental Immunology, University of Zurich, Zurich, Switzerland, 3 Zurich Center for Integrative Human Physiology, University of Zurich, Zurich, Switzerland Edited by: Michele Marie Kosiewicz, University of Louisville, United States Protein tyrosine phosphatase non-receptor type 2 (PTPN2) plays a pivotal role in Reviewed by: immune homeostasis and has been associated with human autoimmune and chronic Cong-Yi Wang, inflammatory diseases. Though PTPN2 is well-characterized in lymphocytes, little is Tongji Medical College, China Andrew L. Mellor, known about its function in innate immune cells. Our findings demonstrate that dendritic Newcastle University, United Kingdom cell (DC)-intrinsic PTPN2 might be the key to explain the central role for PTPN2 in *Correspondence: the immune system to maintain immune tolerance. Partial genetic PTPN2 ablation in Michael Scharl DCs resulted in spontaneous inflammation, particularly in skin, liver, lung and kidney 22 [email protected] weeks post-birth. DC-specific PTPN2 controls steady-state immune cell composition † Present address: and even incomplete PTPN2 deficiency in DCs resulted in enhanced organ infiltration of Juliana Komuczki, Roche Innovation Center Basel, conventional type 2 DCs, accompanied by expansion of IFNγ-producing effector T-cells. -

Pdf/Infopackage Kinex.Pdf for a Com- Domain Inhibition (15)
Protein Tyrosine Phosphatase Receptor Type γ Is a JAK Phosphatase and Negatively Regulates Leukocyte Integrin Activation This information is current as Michela Mirenda, Lara Toffali, Alessio Montresor, Giovanni of October 1, 2021. Scardoni, Claudio Sorio and Carlo Laudanna J Immunol 2015; 194:2168-2179; Prepublished online 26 January 2015; doi: 10.4049/jimmunol.1401841 http://www.jimmunol.org/content/194/5/2168 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2015/01/23/jimmunol.140184 Material 1.DCSupplemental http://www.jimmunol.org/ References This article cites 46 articles, 16 of which you can access for free at: http://www.jimmunol.org/content/194/5/2168.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on October 1, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2015 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Protein Tyrosine Phosphatase Receptor Type g Is a JAK Phosphatase and Negatively Regulates Leukocyte Integrin Activation Michela Mirenda,* Lara Toffali,*,† Alessio Montresor,*,† Giovanni Scardoni,† Claudio Sorio,* and Carlo Laudanna*,† Regulation of signal transduction networks depends on protein kinase and phosphatase activities. -

A Homozygous Deletion Within the Carbonic Anhydrase-Like Domain of the Ptprg Gene in Murine L-Cells1
(CANCER RESEARCH 53. 14«)»-1502.April1. 1993| Advances in Brief A Homozygous Deletion within the Carbonic Anhydrase-like Domain of the Ptprg Gene in Murine L-Cells1 Kishore K. Wary, Zhuangwei Lou, Arthur M. Buchberg, Linda D. Siracusa, Teresa Druck, Sal LaForgia, and Kay Huebner2 Jefferson Cancer Instilare. Thomas Jefferson Medical College, Philadelphia. Pennsylvania ¡9ÃŒ07 Abstract and murine predicted amino acid sequences for this isoform are greater than 90% identical: the Ptprg gene contains a single trans- Protein tyrosine phosphatases, on purely theoretical grounds, were sug membrane domain and two tandem tyrosine phosphatase domains. gested as possible tumor suppressor genes, and receptor protein tyrosine The extracellular region contains one fibronectin type repeat and. like phosphatase 7 (PTPRG ) has been proposed, on the basis of its location at the PTPRZ gene (7).' an NH2-terminal region of 266 amino acids with human chromosome region 3pl4.2, specifically as a tumor suppressor gene —¿30%sequence similarity to members of the carbonic anhydrase for renal cell carcinoma. We have isolated murine genomic and complementary DNA clones for enzyme family (8). analysis and mapping of the murine Ptprg locus; interspecific backcross During an investigation of the genomic organization of human and analysis showed that the Ptprg locus maps to the centramene region of mouse Ptprg loci, we noted extensive polymorphism of the murine mouse chromosome 14. We also observed a homozygous, intragenic dele PÃprglocus5 and a homozygous deletion of a portion of the CA-like tion in the Ptprg gene in all donai derivatives of the original I.-cell strain, domain in murine L-cell lines, which we undertook to describe in a methylcholanthrene-treated mouse connective tissue cell line which pro detail, since homozygous deletion of a gene is one of the hallmarks of duces sarcomas in syngeneic mice. -

Genetic Alterations of Protein Tyrosine Phosphatases in Human Cancers
Oncogene (2015) 34, 3885–3894 © 2015 Macmillan Publishers Limited All rights reserved 0950-9232/15 www.nature.com/onc REVIEW Genetic alterations of protein tyrosine phosphatases in human cancers S Zhao1,2,3, D Sedwick3,4 and Z Wang2,3 Protein tyrosine phosphatases (PTPs) are enzymes that remove phosphate from tyrosine residues in proteins. Recent whole-exome sequencing of human cancer genomes reveals that many PTPs are frequently mutated in a variety of cancers. Among these mutated PTPs, PTP receptor T (PTPRT) appears to be the most frequently mutated PTP in human cancers. Beside PTPN11, which functions as an oncogene in leukemia, genetic and functional studies indicate that most of mutant PTPs are tumor suppressor genes. Identification of the substrates and corresponding kinases of the mutant PTPs may provide novel therapeutic targets for cancers harboring these mutant PTPs. Oncogene (2015) 34, 3885–3894; doi:10.1038/onc.2014.326; published online 29 September 2014 INTRODUCTION tyrosine/threonine-specific phosphatases. (4) Class IV PTPs include Protein tyrosine phosphorylation has a critical role in virtually all four Drosophila Eya homologs (Eya1, Eya2, Eya3 and Eya4), which human cellular processes that are involved in oncogenesis.1 can dephosphorylate both tyrosine and serine residues. Protein tyrosine phosphorylation is coordinately regulated by protein tyrosine kinases (PTKs) and protein tyrosine phosphatases 1 THE THREE-DIMENSIONAL STRUCTURE AND CATALYTIC (PTPs). Although PTKs add phosphate to tyrosine residues in MECHANISM OF PTPS proteins, PTPs remove it. Many PTKs are well-documented oncogenes.1 Recent cancer genomic studies provided compelling The three-dimensional structures of the catalytic domains of evidence that many PTPs function as tumor suppressor genes, classical PTPs (RPTPs and non-RPTPs) are extremely well because a majority of PTP mutations that have been identified in conserved.5 Even the catalytic domain structures of the dual- human cancers are loss-of-function mutations. -
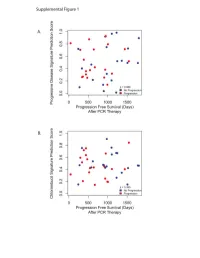
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

Supplementary Table 2
Supplementary Table 2. Differentially Expressed Genes following Sham treatment relative to Untreated Controls Fold Change Accession Name Symbol 3 h 12 h NM_013121 CD28 antigen Cd28 12.82 BG665360 FMS-like tyrosine kinase 1 Flt1 9.63 NM_012701 Adrenergic receptor, beta 1 Adrb1 8.24 0.46 U20796 Nuclear receptor subfamily 1, group D, member 2 Nr1d2 7.22 NM_017116 Calpain 2 Capn2 6.41 BE097282 Guanine nucleotide binding protein, alpha 12 Gna12 6.21 NM_053328 Basic helix-loop-helix domain containing, class B2 Bhlhb2 5.79 NM_053831 Guanylate cyclase 2f Gucy2f 5.71 AW251703 Tumor necrosis factor receptor superfamily, member 12a Tnfrsf12a 5.57 NM_021691 Twist homolog 2 (Drosophila) Twist2 5.42 NM_133550 Fc receptor, IgE, low affinity II, alpha polypeptide Fcer2a 4.93 NM_031120 Signal sequence receptor, gamma Ssr3 4.84 NM_053544 Secreted frizzled-related protein 4 Sfrp4 4.73 NM_053910 Pleckstrin homology, Sec7 and coiled/coil domains 1 Pscd1 4.69 BE113233 Suppressor of cytokine signaling 2 Socs2 4.68 NM_053949 Potassium voltage-gated channel, subfamily H (eag- Kcnh2 4.60 related), member 2 NM_017305 Glutamate cysteine ligase, modifier subunit Gclm 4.59 NM_017309 Protein phospatase 3, regulatory subunit B, alpha Ppp3r1 4.54 isoform,type 1 NM_012765 5-hydroxytryptamine (serotonin) receptor 2C Htr2c 4.46 NM_017218 V-erb-b2 erythroblastic leukemia viral oncogene homolog Erbb3 4.42 3 (avian) AW918369 Zinc finger protein 191 Zfp191 4.38 NM_031034 Guanine nucleotide binding protein, alpha 12 Gna12 4.38 NM_017020 Interleukin 6 receptor Il6r 4.37 AJ002942