How to Stabilize Protein: Stability Screens for Thermal Shift Assays and Nano Differential Scanning Fluorimetry in the Virus-X Project
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Identifying Drug Targets in Tissues and Whole Blood with Thermal-Shift Profiling
LETTERS https://doi.org/10.1038/s41587-019-0388-4 Identifying drug targets in tissues and whole blood with thermal-shift profiling Jessica Perrin1,3, Thilo Werner1,3, Nils Kurzawa 2,3, Anna Rutkowska1, Dorothee D. Childs2, Mathias Kalxdorf1, Daniel Poeckel1, Eugenia Stonehouse1, Katrin Strohmer1, Bianca Heller1, Douglas W. Thomson1, Jana Krause1, Isabelle Becher 2, H. Christian Eberl1, Johanna Vappiani1, Daniel C. Sevin1, Christina E. Rau1, Holger Franken1, Wolfgang Huber2, Maria Faelth-Savitski1, Mikhail M. Savitski2*, Marcus Bantscheff 1* and Giovanna Bergamini 1* Monitoring drug–target interactions with methods such as and only after heating proteins are extracted with a mild deter- the cellular thermal-shift assay (CETSA) is well established gent (Supplementary Fig. 1), resulting in better solubilization of for simple cell systems but remains challenging in vivo. Here membrane-bound proteins. The percentage of membrane pro- we introduce tissue thermal proteome profiling (tissue-TPP), teins detected in crude cell extracts following this protocol was which measures binding of small-molecule drugs to proteins fourfold higher than in detergent-free extracts (PBS extracts) and in tissue samples from drug-treated animals by detecting almost comparable to that in intact cells (Supplementary Fig. 2a changes in protein thermal stability using quantitative mass and Supplementary Data 1). In line with previous reports, melt- spectrometry. We report organ-specific, proteome-wide ing temperatures (Tm) measured in HepG2 cells and HepG2 crude thermal stability maps and derive target profiles of the non- extracts showed a correlation of r = 0.58, an expected value owing covalent histone deacetylase inhibitor panobinostat in rat to differences in concentration of cellular co-factors and the altera- liver, lung, kidney and spleen and of the B-Raf inhibitor vemu- tion of protein–protein interactions (Supplementary Fig. -

Modulation of Alpha-Synuclein Protein Folding by a Marine-Sourced
MODULATION OF ALPHA-SYNUCLEIN PROTEIN FOLDING BY A MARINE-SOURCED EXTRACT by James Christopher Giffin Submitted in partial fulfilment of the requirements for the degree of Master of Science at Dalhousie University Halifax, Nova Scotia April 2016 © Copyright by James Christopher Giffin, 2016 TABLE OF CONTENTS List of Figures ................................................................................................................................. v Abstract .......................................................................................................................................... vi List of Abbreviations and Symbols Used ..................................................................................... vii Acknowledgements ...................................................................................................................... viii 1 CHAPTER 1 INTRODUCTION ............................................................................................. 1 1.1 Amyloid ............................................................................................................................ 1 1.1.1 Protein Folding and Misfolding ................................................................................ 1 1.1.2 Amyloid Formation and Human Disease .................................................................. 5 1.2 Parkinson’s Disease.......................................................................................................... 7 1.2.1 Alpha Synuclein ....................................................................................................... -

Protein-Ligand Interactions Investigated by Thermal Shift Assays
research papers Acta Crystallographica Section D Biological Protein–ligand interactions investigated by Crystallography thermal shift assays (TSA) and dual polarization ISSN 1399-0047 interferometry (DPI) Morten K. Grøftehauge,a* Over the last decades, a wide range of biophysical techniques Received 12 February 2014 Nelly R. Hajizadeh,a Marcus J. investigating protein–ligand interactions have become indis- Accepted 17 July 2014 Swannb and Ehmke Pohlc* pensable tools to complement high-resolution crystal structure determinations. Current approaches in solution range from high-throughput-capable methods such as thermal shift assays a Chemistry Department, Durham University, (TSA) to highly accurate techniques including microscale South Road, Durham DH1 3LE, England, bFarfield, Biolin Scientific, 62 Wellington Road thermophoresis (MST) and isothermal titration calorimetry South, Stockport, Cheshire SK1 3SU, England, (ITC) that can provide a full thermodynamic description and cChemistry Department and School of of binding events. Surface-based methods such as surface Biological and Biomedical Sciences, Durham plasmon resonance (SPR) and dual polarization interfero- University, South Road, Durham DH1 3LE, metry (DPI) allow real-time measurements and can provide England kinetic parameters as well as binding constants. DPI provides additional spatial information about the binding event. Here, Correspondence e-mail: an account is presented of new developments and recent [email protected], applications of TSA and DPI connected to crystallography. -

Integrated Biophysical Approach to Fragment Screening and Validation for Fragment-Based Lead Discovery
Integrated biophysical approach to fragment screening and validation for fragment-based lead discovery Hernani Leonardo Silvestrea,1, Thomas L. Blundella, Chris Abellb, and Alessio Ciullib,1,2 aDepartment of Biochemistry, University of Cambridge, Cambridge CB2 1GA, United Kingdom; and bUniversity Chemical Laboratory, Department of Chemistry, University of Cambridge, Cambridge CB2 1EW, United Kingdom Edited by Dagmar Ringe, Brandeis University, Waltham, MA, and accepted by the Editorial Board June 13, 2013 (received for review March 5, 2013) In fragment-based drug discovery, the weak affinities exhibited by detection, and sensitivity capabilities (21). Although several fragments pose significant challenges for screening. Biophysical methods—for example, NMR, MS, and ITC—are conducted in techniques are used to address this challenge, but there is no clear solution, others require protein crystals (X-ray crystallography) consensus on which cascade of methods is best suited to identify or immobilization to a surface (SPR). Some of these methods fragment hits that ultimately translate into bound X-ray structures can be applied with significant automation and throughput. Be- and provide bona fide starting points for synthesis. We have cause of the broad differences among the techniques that are benchmarked an integrated biophysical approach for fragment used for fragment screening and their complementary advan- screening and validation against Mycobacterium tuberculosis pan- tages and disadvantages, there is a lack of consensus as to which tothenate -

Supporting Information
Supporting Information Chemical validation of Mycobacterium tuberculosis phosphopantetheine adenylyltransferase using fragment linking and CRISPR interference Jamal El Bakali1,†, Michal Blaszczyk2,†, Joanna C. Evans3, Jennifer A. Boland1, William J. McCarthy1, Marcio V. B. Dias2, Anthony G. Coyne1, Valerie Mizrahi3, Tom L. Blundell2, Chris Abell1,* and Christina Spry1,* 1Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge CB2 1EW (UK) 2Department of Biochemistry, University of Cambridge, 80 Tennis Court Road, Cambridge CB2 1GA (UK) 3MRC/NHLS/UCT Molecular Mycobacteriology Research Unit, DST/NRF Centre of Excellence for Biomedical TB Research & Wellcome Centre for Infectious Diseases Research in Africa, Institute of Infectious Disease and Molecular Medicine and Department of Pathology, Faculty of Health Sciences, University of Cape Town, Anzio Road, Observatory 7925 (South Africa) †These authors contributed equally to this work *Correspondence: Chris Abell ([email protected]) or Christina Spry ([email protected]) Table of Contents Supplementary Figures……………………………………………………………………….2 Figure S1……………………………………………………………………………….2 Figure S2……………………………………………………………………………….3 Figure S3……………………………………………………………………………….4 Figure S4……………………………………………………………………………….5 Figure S5……………………………………………………………………………….6 Figure S6……………………………………………………………………………….7 Figure S7……………………………………………………………………………….8 Figure S8……………………………………………………………………………….9 Figure S9……………………………………………………………………………...10 Figure S10…………………………………………………………………………….11 Figure S11…………………………………………………………………………….12 -

Identification of Drug Targets and Their Mechanisms of Action
Direct and indirect approaches to identify drug modes of action Lindsay B. Tulloch1, Stefanie K. Menzies2, Ross P. Coron3, Matthew D. Roberts4, Gordon J. Florence1* & Terry K. Smith1* 1 EaStChem School of Chemistry and School of Biology, Biomedical Sciences Research Complex, University of St Andrews, St Andrews, Fife KY16 9ST, UK. 2 Current address: Department of Molecular Microbiology, Washington University School of Medicine, St. Louis, Missouri 63110 USA 3 Current address: Department of Parasites and Insect Vectors, Institut Pasteur, 25 Rue du Docteur Roux, 75015, Paris, France 4 Current address: Department of Nutritional Sciences, Faculty of Health and Medical Sciences, University of Surrey, Guildford GU2 7WG, UK * Corresponding authors Email: [email protected] (TKS) / [email protected] (GJF) Keywords: Drug mode of action, Target ID, Genomics, Transcriptomics, Proteomics, Metabolomics, Affinity chromatography, Photo-affinity labeling. Abbreviations cDNA complementary DNA CETSA cellular thermal shift assay CRISPR clustered regularly interspaced short palindromic repeats DARTS Drug affinity response target stability dsRNA double-stranded RNA ESI electrospray ionisation HIV human immunodeficiency virus ICAT isotope-coded affinity tagging IEX ion exchange iTRAQ isobaric tags for relative and absolute quantitation LC liquid chromatography LC-MS liquid chromatography – mass spectrometry LC-MS/MS liquid chromatography – tandem mass spectrometry MALDI matrix-assisted laser desorption ionisation MOA mode of action mRNA messenger RNA -

Label-Free Technologies for Target Identification and Validation†
MedChemComm Accepted Manuscript This is an Accepted Manuscript, which has been through the Royal Society of Chemistry peer review process and has been accepted for publication. Accepted Manuscripts are published online shortly after acceptance, before technical editing, formatting and proof reading. Using this free service, authors can make their results available to the community, in citable form, before we publish the edited article. We will replace this Accepted Manuscript with the edited and formatted Advance Article as soon as it is available. You can find more information about Accepted Manuscripts in the Information for Authors. Please note that technical editing may introduce minor changes to the text and/or graphics, which may alter content. The journal’s standard Terms & Conditions and the Ethical guidelines still apply. In no event shall the Royal Society of Chemistry be held responsible for any errors or omissions in this Accepted Manuscript or any consequences arising from the use of any information it contains. www.rsc.org/medchemcomm Page 1 of 15 PleaseMedChemComm do not adjust margins Journal Name ARTICLE Label-free technologies for target identification and validation† Jing Li, a* Hua Xu, a Graham M. West b and Lyn H. Jones a Received 00th January 20xx, Accepted 00th January 20xx Phenotypic screening is a powerful strategy for identifying active molecules with particular biological effects in cellular or DOI: 10.1039/x0xx00000x animal disease models. Functionalized chemical probes have been instrumental in revealing new targets and confirming target engagement. However, substantial effort and resources are required to design and synthesize these bioactive www.rsc.org/ probes. -

Abstract Characterization of Potential Anti-Infective
ABSTRACT CHARACTERIZATION OF POTENTIAL ANTI-INFECTIVE AGENTS OF Burkholderia pseudomallei TARGETING IspF Joy Michelle Blain, Ph.D. Department of Chemistry and Biochemistry Northern Illinois University, 2016 James R. Horn, Co-Director Timothy J. Hagen, Co-Director There is an urgent need for new anti-infective agents. Pathogenic organisms continue to develop resistance to modern-day antibiotics, while there has been a significant decline in the development of antibiotics over the last 30 years. One particular pathway of interest in the development of new antibiotics is the biosynthesis of isoprenoids. These compounds are essential chemical building blocks in pathogenic organisms such as Plasmodium falciparum, Mycobacterium tuberculosis, and Burkholderia pseudomallei. Notably, there are two distinct pathways in the biosynthesis of isoprenoids called the Mevalonate (MVA) pathway and the 2C-methyl-D-erythritol phosphate (MEP) pathway. The MEP pathway enzymes are attractive targets in developing novel anti-infective agents due to the fact that humans only use the MVA pathway to produce isoprenoids. Burkholderia pseudomallei is a Gram-negative bacterium found in untreated water and soil in Southeast Asia and northern Australia. The bacterium has been classified as a category B bioterrorism agent/disease by the category B bioterrorism agent/disease by the Centers for Disease Control and Prevention. Antibiotic resistance to traditional anti-microbial agents in such pathogenic organisms has increased due to the primary mechanisms of resistance, which are enzyme inactivation, drug efflux from the cell, and target deletion. Consequently, new anti- infective agents targeting B. pseudomallei are of great interest. The focus of this research project was to examine the inhibition of the fifth enzyme in the MEP pathway, 2C-methyl-D-erythritol 2,4-cyclodiphosphate synthase (IspF) using small molecules that were developed based on fragment hits and structure activity relationships (SAR). -
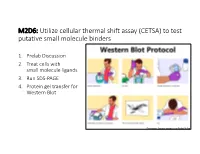
M2D6: Utilize Cellular Thermal Shift Assay (CETSA) to Test Putative Small Molecule Binders
M2D6: Utilize cellular thermal shift assay (CETSA) to test putative small molecule binders 1. Prelab Discussion 2. Treat cells with small molecule ligands 3. Run SDS-PAGE 4. Protein gel transfer for Western Blot Pedromics, Science cartoons by Pedro Veliça Mod2 Overview Cellular thermal shift assay (CETSA) ON THE MOD2 OVERVIEW PAGE: • The ΔTm indicates protein stabilization / destabilization compared to control • Assesses thermal stabilization of protein Native proteins denature at in presence / absence lower temperature than of drug (small molecule) native folded proteins associated with drug in the cell Source: Sygnature Discovery Cellular thermal shift assay (CETSA) • The ΔTm indicates protein stabilization / Why is testing fordestabilization ligand binding incompared a to control • Assesses thermal cell important?stabilization of protein Native proteins denature at in presence / absence lower temperature than native folded proteins of ligand in the cell associated with drug Source: Sygnature Discovery CETSA : Protein stability is measured by Western blot Identify presence of native folded proteins Source: Sygnature Discovery Major experimental steps of CETSA Heat causes protein denaturation • As proteins denature, ‘melted’ primary structures aggregate • Aggregates precipitate out of solution and can be removed via centrifugation soluble Aggregate Native protein insoluble pellet Purpose of CETSA steps • Heat: • Snap freeze: • Spin: spin Only soluble proteins are measured by Western blot Western Blot soluble insoluble pellet Imaging for -

Stabilization of Amyloidogenic Immunoglobulin Light Chains by Small Molecules
Stabilization of amyloidogenic immunoglobulin light chains by small molecules Gareth J. Morgana,b,1,2,3, Nicholas L. Yana,b,1, David E. Mortensona,b, Enrico Rennellac,d,e, Joshua M. Blundona,b, Ryan M. Gwina,b, Chung-Yon Lina,b, Robyn L. Stanfieldf, Steven J. Browna, Hugh Rosena, Timothy P. Spicerg, Virneliz Fernandez-Vegag, Giampaolo Merlinih,i, Lewis E. Kayc,d,e,j, Ian A. Wilsonf,k, and Jeffery W. Kellya,b,k,2 aDepartment of Molecular Medicine, The Scripps Research Institute, La Jolla, CA 92037; bDepartment of Chemistry, The Scripps Research Institute, La Jolla, CA 92037; cDepartment of Molecular Genetics, The University of Toronto, Toronto, ON M5S1A8, Canada; dDepartment of Biochemistry, The University of Toronto, Toronto, ON M5S1A8, Canada; eDepartment of Chemistry, The University of Toronto, Toronto, ON M5S1A8, Canada; fDepartment of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037; gDepartment of Molecular Medicine, The Scripps Research Institute, Jupiter, FL 33458; hDepartment of Molecular Medicine, University of Pavia, 27100 Pavia, Italy; iAmyloidosis Research and Treatment Center, Foundation Istituto di Ricovero e Cura a Carattere Scientifico Policlinico San Matteo, 27100 Pavia, Italy; jProgram in Molecular Medicine, The Hospital for Sick Children, Toronto, ON M5G1X8, Canada; and kThe Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, CA 92037 Edited by Susan Marqusee, University of California, Berkeley, CA, and approved March 13, 2019 (received for review October 11, 2018) In Ig light-chain (LC) amyloidosis (AL), the unique antibody LC that we and others have identified as potential targets for protein that is secreted by monoclonal plasma cells in each patient stabilization (21, 22). -

Measurement of Nanomolar Dissociation Constants by Titration Calorimetry and Thermal Shift Assay – Radicicol Binding to Hsp90 and Ethoxzolamide Binding to CAII
Int. J. Mol. Sci. 2009, 10, 2662-2680; doi:10.3390/ijms10062662 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Article Measurement of Nanomolar Dissociation Constants by Titration Calorimetry and Thermal Shift Assay – Radicicol Binding to Hsp90 and Ethoxzolamide Binding to CAII Asta Zubrienė, Jurgita Matulienė, Lina Baranauskienė, Jelena Jachno, Jolanta Torresan, Vilma Michailovienė, Piotras Cimmperman and Daumantas Matulis * Laboratory of Biothermodynamics and Drug Design / Institute of Biotechnology, Graičiūno 8, Vilnius, LT-02241, Lithuania; E-Mails: [email protected] (A.Z.); [email protected] (J.M.); [email protected] (L.B.); [email protected] (J.J.); [email protected] (J.T.); [email protected] (V.M.); [email protected] (P.C.) * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel. +370-5-269-1884; Fax: +370-5-260-2116 Received: 25 March 2009; in revised form: 30 May 2009 / Accepted: 3 June 2009 / Published: 10 June 2009 Abstract: The analysis of tight protein-ligand binding reactions by isothermal titration calorimetry (ITC) and thermal shift assay (TSA) is presented. The binding of radicicol to the N-terminal domain of human heat shock protein 90 (Hsp90αN) and the binding of ethoxzolamide to human carbonic anhydrase (hCAII) were too strong to be measured accurately by direct ITC titration and therefore were measured by displacement ITC and by observing the temperature-denaturation transitions of ligand-free and ligand-bound protein. Stabilization of both proteins by their ligands was profound, increasing the melting temperature by more than 10 ºC, depending on ligand concentration. Analysis of the melting temperature dependence on the protein and ligand concentrations yielded dissociation constants equal to 1 nM and 2 nM for Hsp90αN-radicicol and hCAII- ethoxzolamide, respectively. -

An Engineered Thermal-Shift Screen Reveals Specific Lipid Preferences Of
ARTICLE DOI: 10.1038/s41467-018-06702-3 OPEN An engineered thermal-shift screen reveals specific lipid preferences of eukaryotic and prokaryotic membrane proteins Emmanuel Nji 1, Yurie Chatzikyriakidou1, Michael Landreh2 & David Drew1 Membrane bilayers are made up of a myriad of different lipids that regulate the functional activity, stability, and oligomerization of many membrane proteins. Despite their importance, 1234567890():,; screening the structural and functional impact of lipid–protein interactions to identify specific lipid requirements remains a major challenge. Here, we use the FSEC-TS assay to show cardiolipin-dependent stabilization of the dimeric sodium/proton antiporter NhaA, demon- strating its ability to detect specific protein-lipid interactions. Based on the principle of FSEC- TS, we then engineer a simple thermal-shift assay (GFP-TS), which facilitates the high- throughput screening of lipid- and ligand- interactions with membrane proteins. By com- paring the thermostability of medically relevant eukaryotic membrane proteins and a selec- tion of bacterial counterparts, we reveal that eukaryotic proteins appear to have evolved to be more dependent to the presence of specific lipids. 1 Centre for Biomembrane Research, Department of Biochemistry and Biophysics, Stockholm University, SE-106 91 Stockholm, Sweden. 2 SciLifeLab and Department of Microbiology, Tumor and Cell Biology, Karolinska Institutet, SE-171 65 Stockholm, Sweden. These authors contributed equally: Emmanuel Nji, Yurie Chatzikyriakidou. Correspondence and requests for materials should be addressed to D.D. (email: [email protected]) NATURE COMMUNICATIONS | (2018) 9:4253 | DOI: 10.1038/s41467-018-06702-3 | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | DOI: 10.1038/s41467-018-06702-3 embrane bilayers are assembled from a plethora of lipid Consistent with the MS data15, FSEC-TS strongly indicated Mclasses with varying biophysical properties.