Genetic Stock Structure of the Sailfish, Istiophorus Platypterus, Based on Nuclear and Mitochondrial DNA
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

On the Biology of Florida East Coast Atlantic Sailfisht' Íe (Istiophorus Platypterus)1
SJf..... z e e w e t e k r " - Ka,; ,y, -, V öi'*DzBZöm 8429 =*==,_ r^ä r.-,-. On the Biology of Florida East Coast Atlantic SailfishT' íe (Istiophorus platypterus)1 JOHN W. JOLLEY, JR .2 149299 ABSTRACT The sailfish, Istiophorus platypterus, is one of the most important species in southeast Florida’s marine sport fishery. Recently, the concern of Palm Beach anglers about apparent declines in numbers of sailfish caught annually prompted the Florida Department of Natural Resources Marine Research Laboratory to investigate the biological status of Florida’s east coast sailfish populations. Fresh specimens from local sport catches were examined monthly during May 1970 through September 1971. Monthly plankton and “ night-light” collections of larval and juvenile stages were also obtained. Attempts are being made to estimate sailfish age using concentric rings in dorsal fin spines. If successful, growth rates will be determined for each sex and age of initial maturity described. Females were found to be consistently larger than males and more numerous during winter. A significant difference in length-weight relationship was also noted between sexes. Fecundity estimates varied from 0.8 to 1.6 million “ ripe” ova, indicating that previous estimates (2.5 to 4.7 million ova) were probably high. Larval istiophorids collected from April through October coincided with the prominence of “ ripe” females in the sport catch. Microscopic examination of ovarian tissue and inspection of “ ripe” ovaries suggest multiple spawning. Florida’s marine sport fishery has been valued as a in 1948 at the request of the Florida Board of Con $200 million business (de Sylva, 1969). -

The 2016 SWFSC Billfish Newsletter
The SouthwestSWFSC Fisheries 2016 Billfish Science Newsletter Center’s 2016 Billfish Newsletter Global Tagging Map El Niño fishing conditions Catch-Photo-Release mobile phone application IGFA Great Marlin Race and satellite tagging 1 Top Anglers and Captains of 2015 SWFSC 2016 Billfish Newsletter Table of Contents Special Foreword …………………………………………………………….. 3 An Inside Look ……………………………………………………………..… 4 Prologue …………………………………………………………………….… 5 Introduction ……………………………………………………………..….… 5 The International Billfish Angler Survey ………………………………....... 7 Pacific blue marlin 9 Striped marlin 10 Indo-Pacific sailfish 11 Black marlin 13 Shortbill spearfish 13 Broadbill swordfish 14 The Billfish Tagging Program ……………………………………………..... 14 The Hawaiian Islands 16 2015 Tagging-at-a-Glance Map 17 Baja California and Guerrero, Mexico 18 Southern California 18 Western Pacific 18 Top Anglers and Captains Acknowledgements ……………………………. 19 Top Tagging Anglers 19 Top Tagging Captains 21 Tag Recoveries ……………………………………………………………….. 21 Science in Action: “The IGFA Great Marlin Race and Marlin Tagging” 23 Acknowledgements ………………………………………………………....... 25 Angler Photos ……………………………………………………………..….. 26 Congratulations to Captain Teddy Hoogs of the Bwana for winning this year’s cover photo contest! Teddy photographed this spectacular marlin off the coast of Hawaii. Fish on! 2 Special Forward James Wraith, director of the SWFSC Cooperative Billfish Tagging Program since 2007, recently left the SWFSC to move back to Australia. James was an integral part of the Highly Migratory Species (HMS) program. In addition to day-to-day work, James planned and organized the research cruises for HMS at the SWFSC and was involved in tagging thresher, blue, and mako sharks in the Southern California Bight for many years. We are sad to see him go but are excited for his future opportunities and thankful for his many contributions to the program over the last 10 years. -

Striped Marlin (Kajikia Audax)
I & I NSW WILD FISHERIES RESEARCH PROGRAM Striped Marlin (Kajikia audax) EXPLOITATION STATUS UNDEFINED Status is yet to be determined but will be consistent with the assessment of the south-west Pacific stock by the Scientific Committee of the Central and Western Pacific Fisheries Commission. SCIENTIFIC NAME STANDARD NAME COMMENT Kajikia audax striped marlin Previously known as Tetrapturus audax. Kajikia audax Image © I & I NSW Background lengths greater than 250cm (lower jaw-to-fork Striped marlin (Kajikia audax) is a highly length) and can attain a maximum weight of migratory pelagic species distributed about 240 kg. Females mature between 1.5 and throughout warm-temperate to tropical 2.5 years of age whilst males mature between waters of the Indian and Pacific Oceans.T he 1 and 2 years of age. Striped marlin are stock structure of striped marlin is uncertain multiple batch spawners with females shedding although there are thought to be separate eggs every 1-2 days over 4-41 events per stocks in the south-west, north-west, east and spawning season. An average sized female of south-central regions of the Pacific Ocean, as about 100 kg is able to produce up to about indicated by genetic research, tagging studies 120 million eggs annually. and the locations of identified spawning Striped marlin spend most of their time in grounds. The south-west Pacific Ocean (SWPO) surface waters above the thermocline, making stock of striped marlin spawn predominately them vulnerable to surface fisheries.T hey are during November and December each year caught mostly by commercial longline and in waters warmer than 24°C between 15-30°S recreational fisheries throughout their range. -

Fao Species Catalogue
FAO Fisheries Synopsis No. 125, Volume 5 FIR/S125 Vol. 5 FAO SPECIES CATALOGUE VOL. 5. BILLFISHES OF THE WORLD AN ANNOTATED AND ILLUSTRATED CATALOGUE OF MARLINS, SAILFISHES, SPEARFISHES AND SWORDFISHES KNOWN TO DATE UNITED NATIONS DEVELOPMENT PROGRAMME FOOD AND AGRICULTURE ORGANIZATION OF THE UNITED NATIONS FAO Fisheries Synopsis No. 125, Volume 5 FIR/S125 Vol.5 FAO SPECIES CATALOGUE VOL. 5 BILLFISHES OF THE WORLD An Annotated and Illustrated Catalogue of Marlins, Sailfishes, Spearfishes and Swordfishes Known to date MarIins, prepared by Izumi Nakamura Fisheries Research Station Kyoto University Maizuru Kyoto 625, Japan Prepared with the support from the United Nations Development Programme (UNDP) UNITED NATIONS DEVELOPMENT PROGRAMME FOOD AND AGRICULTURE ORGANIZATION OF THE UNITED NATIONS Rome 1985 The designations employed and the presentation of material in this publication do not imply the expression of any opinion whatsoever on the part of the Food and Agriculture Organization of the United Nations concerning the legal status of any country, territory. city or area or of its authorities, or concerning the delimitation of its frontiers or boundaries. M-42 ISBN 92-5-102232-1 All rights reserved . No part of this publicatlon may be reproduced. stored in a retriewal system, or transmitted in any form or by any means, electronic, mechanical, photocopying or otherwase, wthout the prior permission of the copyright owner. Applications for such permission, with a statement of the purpose and extent of the reproduction should be addressed to the Director, Publications Division, Food and Agriculture Organization of the United Nations Via delle Terme di Caracalla, 00100 Rome, Italy. -

Black Marlin (Makaira Indica) Blue Marlin (Makaira Nigricans) Blue
Black marlin (Makaira indica) Blue marlin (Makaira nigricans) Blue shark (Prionace glauca) Opah (Lampris guatus) Shorin mako shark (Isurus oxyrinchus) Striped marlin (Kaijkia audax) Western Central Pacific, North Pacific, South Pacific Pelagic longline July 12, 2016 Alexia Morgan, Consulng Researcher Disclaimer Seafood Watch® strives to have all Seafood Reports reviewed for accuracy and completeness by external sciensts with experse in ecology, fisheries science and aquaculture. Scienfic review, however, does not constute an endorsement of the Seafood Watch® program or its recommendaons on the part of the reviewing sciensts. Seafood Watch® is solely responsible for the conclusions reached in this report. Table of Contents Table of Contents 2 About Seafood Watch 3 Guiding Principles 4 Summary 5 Final Seafood Recommendations 5 Introduction 8 Assessment 10 Criterion 1: Impacts on the species under assessment 10 Criterion 2: Impacts on other species 22 Criterion 3: Management Effectiveness 45 Criterion 4: Impacts on the habitat and ecosystem 55 Acknowledgements 58 References 59 Appendix A: Extra By Catch Species 69 2 About Seafood Watch Monterey Bay Aquarium’s Seafood Watch® program evaluates the ecological sustainability of wild-caught and farmed seafood commonly found in the United States marketplace. Seafood Watch® defines sustainable seafood as originang from sources, whether wild-caught or farmed, which can maintain or increase producon in the long-term without jeopardizing the structure or funcon of affected ecosystems. Seafood Watch® makes its science-based recommendaons available to the public in the form of regional pocket guides that can be downloaded from www.seafoodwatch.org. The program’s goals are to raise awareness of important ocean conservaon issues and empower seafood consumers and businesses to make choices for healthy oceans. -

A Black Marlin, Makaira Indica, from the Early Pleistocene of the Philippines and the Zoogeography of Istiophorid B1llfishes
BULLETIN OF MARINE SCIENCE, 33(3): 718-728,1983 A BLACK MARLIN, MAKAIRA INDICA, FROM THE EARLY PLEISTOCENE OF THE PHILIPPINES AND THE ZOOGEOGRAPHY OF ISTIOPHORID B1LLFISHES Harry L Fierstine and Bruce 1. Welton ABSTRACT A nearly complete articulated head (including pectoral and pelvic girdles and fins) was collected from an Early Pleistocene, upper bathyal, volcanic ash deposit on Tambac Island, Northwest Central Luzon, Philippines. The specimen was positively identified because of its general resemblance to other large marlins and by its rigid pectoral fin, a characteristic feature of the black marlin. This is the first fossil billfish described from Asia and the first living species of billfi sh positively identified in the fossil record. The geographic distribution of the two living species of Makaira is discussed, Except for fossil localities bordering the Mediterranean Sea, the distribution offossil post-Oligocene istiophorids roughly corresponds to the distribution of living adult forms. During June 1980, we (Fierstine and Welton, in press) collected a large fossil billfish discovered on the property of Pacific Farms, Inc., Tambac Island, near the barrio of Zaragoza, Bolinao Peninsula, Pangasinan Province, Northwest Cen tral Luzon, Philippines (Fig. 1). In addition, associated fossils were collected and local geological outcrops were mapped. The fieldwork continued throughout the month and all specimens were brought to the Natural History Museum of the Los Angeles County for curation, preparation, and distribution to specialists for study. The fieldwork was important because no fossil bony fish had ever been reported from the Philippines (Hashimoto, 1969), fossil billfishes have never been described from Asia (Fierstine and Applegate, 1974), a living species of billfish has never been positively identified in the fossil record (Fierstine, 1978), and no collection of marine macrofossils has ever been made under strict stratigraphic control in the Bolinao area, if not in the entire Philippines. -
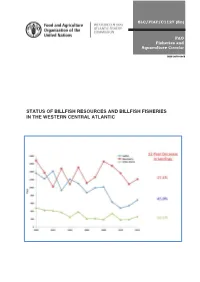
Status of Billfish Resources and the Billfish Fisheries in the Western
SLC/FIAF/C1127 (En) FAO Fisheries and Aquaculture Circular ISSN 2070-6065 STATUS OF BILLFISH RESOURCES AND BILLFISH FISHERIES IN THE WESTERN CENTRAL ATLANTIC Source: ICCAT (2015) FAO Fisheries and Aquaculture Circular No. 1127 SLC/FIAF/C1127 (En) STATUS OF BILLFISH RESOURCES AND BILLFISH FISHERIES IN THE WESTERN CENTRAL ATLANTIC by Nelson Ehrhardt and Mark Fitchett School of Marine and Atmospheric Science, University of Miami Miami, United States of America FOOD AND AGRICULTURE ORGANIZATION OF THE UNITED NATIONS Bridgetown, Barbados, 2016 The designations employed and the presentation of material in this information product do not imply the expression of any opinion whatsoever on the part of the Food and Agriculture Organization of the United Nations (FAO) concerning the legal or development status of any country, territory, city or area or of its authorities, or concerning the delimitation of its frontiers or boundaries. The mention of specific companies or products of manufacturers, whether or not these have been patented, does not imply that these have been endorsed or recommended by FAO in preference to others of a similar nature that are not mentioned. The views expressed in this information product are those of the author(s) and do not necessarily reflect the views or policies of FAO. ISBN 978-92-5-109436-5 © FAO, 2016 FAO encourages the use, reproduction and dissemination of material in this information product. Except where otherwise indicated, material may be copied, downloaded and printed for private study, research and teaching purposes, or for use in non-commercial products or services, provided that appropriate DFNQRZOHGJHPHQWRI)$2DVWKHVRXUFHDQGFRS\ULJKWKROGHULVJLYHQDQGWKDW)$2¶VHQGRUVHPHQWRI XVHUV¶YLHZVSURGXFWVRUVHUYLFHVLVQRWLPSOLHGLQDQ\ZD\ All requests for translation and adaptation rights, and for resale and other commercial use rights should be made via www.fao.org/contact-us/licence-request or addressed to [email protected]. -

(Tetrapturus Albidus) Released from Commercial Pelagic Longline Gear in the Western North
ART & EQUATIONS ARE LINKED 434 Abstract—To estimate postrelease Survival of white marlin (Tetrapturus albidus) survival of white marlin (Tetraptu- rus albidus) caught incidentally in released from commercial pelagic longline gear regular commercial pelagic longline fishing operations targeting sword- in the western North Atlantic* fish and tunas, short-duration pop- up satellite archival tags (PSATs) David W. Kerstetter were deployed on captured animals for periods of 5−43 days. Twenty John E. Graves (71.4%) of 28 tags transmitted data Virginia Institute of Marine Science at the preprogrammed time, includ- College of William and Mary ing one tag that separated from the Route 1208 Greate Road fish shortly after release and was Gloucester Point, Virginia 23062 omitted from subsequent analyses. Present address (for D. W. Kerstetter): Cooperative Institute for Marine and Atmospheric Studies Transmitted data from 17 of 19 Rosenstiel School for Marine and Atmospheric Science tags were consistent with survival University of Miami of those animals for the duration of 4600 Rickenbacker Causeway the tag deployment. Postrelease sur- Miami, Florida 33149 vival estimates ranged from 63.0% E-mail address (for D. W. Kerstetter): [email protected] (assuming all nontransmitting tags were evidence of mortality) to 89.5% (excluding nontransmitting tags from the analysis). These results indi- cate that white marlin can survive the trauma resulting from interac- White marlin (Tetrapturus albidus incidental catch of the international tion with pelagic longline gear, and Poey 1860) is an istiophorid billfish pelagic longline fishery, which targets indicate that current domestic and species widely distributed in tropi- tunas (Thunnus spp.) and swordfish international management measures cal and temperate waters through- (Xiphias gladius). -

Validity, Identification, and Distribution of the Roundscale Spearfish
Nova Southeastern University NSUWorks Marine & Environmental Sciences Faculty Articles Department of Marine and Environmental Sciences NSUWorks Citation Mahmood S. Shivji, Jennifer E. Magnussen, Lawrence R. Beerkircher, George Hinteregger, Dennis W. Lee, Joseph E. Serafy, and Eric D. Prince. 2006. Validity, Identification, and Distribution of the Roundscale Spearfish, Tetrapturus georgii (Teleostei: Istiophoridae): Morphological and Molecular Evidence .Bulletin of Marine Science , (3) : 483 -491. https://nsuworks.nova.edu/occ_facarticles/374. This Article is brought to you for free and open access by the Department of Marine and Environmental Sciences at NSUWorks. It has been accepted for inclusion in Marine & Environmental Sciences Faculty Articles by an authorized administrator of NSUWorks. For more information, please contact [email protected]. 11-1-2006 Validity, Identification, and Distribution of the Roundscale Spearfish, Tetrapturus georgii (Teleostei: Istiophoridae): Morphological and Molecular Evidence Mahmood S. Shivji Nova Southeastern University, <<span class="elink">[email protected] Jennifer E. Magnussen Nova Southeastern University Lawrence R. Beerkircher National Oceanic and Atmospheric Administration George Hinteregger National Oceanic and Atmospheric Administration Dennis W. Lee National Oceanic and Atmospheric Administration See next page for additional authors Find out more information about Nova Southeastern University and the Halmos College of Natural Sciences and Oceanography. Follow this and additional works at: https://nsuworks.nova.edu/occ_facarticles Part of the Genetics and Genomics Commons, Marine Biology Commons, and the Oceanography and Atmospheric Sciences and Meteorology Commons This Article has supplementary content. View the full record on NSUWorks here: https://nsuworks.nova.edu/occ_facarticles/374 Authors Joseph E. Serafy National Oceanic and Atmospheric Administration, [email protected] Eric D. -

63Rd Annual Labor Day White Marlin Tournament
Information THE OCEAN CITY MARLIN CLUB would like to extend a special thanks to our great sponsors. 63rd Annual Labor Day REGISTRATION White Marlin Level Sponsors: White Marlin Tournament Thursday, September 2 @ 6:30-8:00 p.m. Decatur Diner Ocean City Fishing Center Delaware Elevator Sunset Marina In-Person Captain’s Meeting 7:30 p.m. or Virtual Captain’s D.W. Burt Concrete Construction, Inc. SEPTEMBER 3, 4 & 5 2021 Meeting available on our website and Facebook. Over the You do not need to be phone registration available M-F 10:30-5:30 p.m. Blue Marlin Level Sponsors: a member of The Marlin Club to fish this tournament. 410-213-1613. A representative from each boat must Atlantic General Hospital Henley Construction Co. Inc. watch/attend the Captain’s Meeting. Bahia Marina Intrinsic Yacht & Ship Carey Distributors Ocean City Golf Club Open Bar - Beer & Wine Only & donated Appetizers by Consolidated Commercial Services PYY Marine Decatur Diner until 8:00 p.m. (while supplies last) DBS, Inc. Red Sun Custom Apparel Delmarvalous Photos Rt113 Boat Sales Fish in OC Ruth’s Chris Steak House AWARDS BANQUET September 5, 6:30-9:00 p.m. at the Club Billfish Level Sponsors: Awards Presentation: 8:00 p.m. Billfish Gear Ocean Downs Casino Four Tickets included with entry fee. Bluewater Yacht Sales PKS & Company, P.A. C & R Electrics, Inc. Premier Flooring Installation 1, Inc. C.J. Miller LLC Racetrack Auto & Marine FISHING DAYS Casual Designs Furniture SeaBoard Media Inc. (2 of 3) September 3, 4 & 5 Creative Concepts Steen Homes Lines In 8:00 a.m. -

Diet of the White Marlin (Tetrapturus Albidus) from the Southwestern Equatorial Atlantic Ocean
SCRS/2009/173 Collect. Vol. Sci. Pap. ICCAT, 65(5): 1843-1850 (2010) DIET OF THE WHITE MARLIN (TETRAPTURUS ALBIDUS) FROM THE SOUTHWESTERN EQUATORIAL ATLANTIC OCEAN P.B. Pinheiro1, T. Vaske Júnior2, F.H.V. Hazin2, P. Travassos3, M.T. Tolotti3 and T.M. Barbosa2 SUMMARY The aim of this study was to evaluate the diet of white marlin, regarding the number, weight, and frequency of occurrence of the prey items, prey-predator relationships, and feeding strategies in the southwestern equatorial Atlantic Ocean. A total of 257 white marlins were examined, of which 60 (23.3%) were male and 197 (76.7%) were female. Males ranged from 105 to 220 cm low jaw fork length (LJFL) and females ranged from 110 to 236 cm LJFL. Most prey (fish and cephalopods) ranged between 1.0 and 65.0 cm in body length, with a mean length around 10.1 cm. According to the IRI (Index of Relative Importance) ranking, the flying gurnard, Dactylopterus volitans, was the most important prey item, with 27.9% of occurrence, followed by the squid, Ornithoteuthis antillarum (Atlantic bird squid), with 21.2% occurrence. RÉSUMÉ Cette étude vise à évaluer le régime alimentaire des makaires blancs en ce qui concerne le nombre, poids et fréquence de présence des proies, la relation proie-prédateur et les stratégies trophiques dans l’Océan Atlantique équatorial sud-occidental. Un total de 257 makaires blancs ont été examinés, dont 60 (23,3%) étaient des mâles et 197 (76,7%) des femelles. La taille des mâles oscillait entre 105 et 220 cm (longueur maxillaire inférieur-fourche, LJFL) et celle des femelles entre 110 et 236 cm (LJFL). -

Updated Us Conventional Tagging Database For
SCRS/2009/047 Collect. Vol. Sci. Pap. ICCAT, 65(5): 1692-1700 (2010) UPDATED U.S. CONVENTIONAL TAGGING DATABASE FOR ATLANTIC SAILFISH (1955-2008), WITH COMMENTS ON POTENTIAL STOCK STRUCTURE Eric S. Orbesen, Derke Snodgrass, John P. Hoolihan, and Eric D. Prince1 SUMMARY The U.S. conventional tagging data base for Atlantic sailfish (1955-2008), Istiophorus platypterus, consists of data from the NOAA Southeast Fishery Science Center’s Cooperative Tagging Center (CTC) and The Billfish Foundation (TBF). We examined the patterns of sailfish tag release and recapture results in the Atlantic Ocean using a composite analysis from both agencies. In addition, we discuss tagging results and other data that might provide insight into Atlantic sailfish stock structure. RÉSUMÉ La base de données de marquage conventionnel des Etats-Unis pour les voiliers de l’Atlantique (Istiophorus platypterus) (1955-2008) est constituée de données provenant du Cooperative Tagging Center (CTC) du Southeast Fishery Science Center de la NOAA et du The Billfish Foundation (TBF). Nous avons examiné les modes des résultats d'apposition des marques sur les voiliers et de leur récupération dans l’océan Atlantique à l’aide d’une analyse composite émanant des deux agences. En outre, nous discutons des résultats de marquage et d’autres données susceptibles de nous éclairer sur la structure du stock de voiliers de l’Atlantique. RESUMEN La base de datos de marcado convencional estadounidense para el pez vela del Atlántico (1955-2008), Istiophorus platypterus, contiene datos del Cooperative Tagging Center (CTC) del Southeast Fishery Science Center de la NOAA y de The Billfish Foundation (TBF).