Genome-Based Analysis of the Nonhuman Primate Macaca Fascicularis As a Model for Drug Safety Assessment
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

CYP26C1 Is a Hydroxylase of Multiple Active Retinoids and Interacts with Cellular Retinoic Acid Binding Proteins S
Supplemental material to this article can be found at: http://molpharm.aspetjournals.org/content/suppl/2018/02/23/mol.117.111039.DC1 1521-0111/93/5/489–503$35.00 https://doi.org/10.1124/mol.117.111039 MOLECULAR PHARMACOLOGY Mol Pharmacol 93:489–503, May 2018 Copyright ª 2018 by The American Society for Pharmacology and Experimental Therapeutics CYP26C1 Is a Hydroxylase of Multiple Active Retinoids and Interacts with Cellular Retinoic Acid Binding Proteins s Guo Zhong, David Ortiz, Alex Zelter, Abhinav Nath, and Nina Isoherranen Departments of Pharmaceutics (G.Z., N.I.) and Medicinal Chemistry (D.O., A.N.), School of Pharmacy, and Department of Biochemistry, School of Medicine (A.Z.), University of Washington, Seattle, Washington Received October 31, 2017; accepted February 22, 2018 ABSTRACT Downloaded from The clearance of retinoic acid (RA) and its metabolites is believed orientation of retinoids within the CYP26C1 active site. In compar- to be regulated by the CYP26 enzymes, but the specific roles of ison with other CYP26 family members, CYP26C1 was up to CYP26A1, CYP26B1, and CYP26C1 in clearing active vitamin A 10-fold more efficient in clearing 4-oxo-atRA (intrinsic clearance metabolites have not been defined. The goal of this study was to 153 ml/min/pmol) than CYP26A1 and CYP26B1, suggesting that establish the substrate specificity of CYP26C1, and determine CYP26C1 may be important in clearing this active retinoid. In whether CYP26C1 interacts with cellular retinoic acid binding support of this, CRABPs delivered 4-oxo-atRA and atRA for proteins (CRABPs). CYP26C1 was found to effectively metabo- metabolism by CYP26C1. -

Impaired Hepatic Drug and Steroid Metabolism in Congenital Adrenal
European Journal of Endocrinology (2010) 163 919–924 ISSN 0804-4643 CLINICAL STUDY Impaired hepatic drug and steroid metabolism in congenital adrenal hyperplasia due to P450 oxidoreductase deficiency Dorota Tomalik-Scharte1, Dominique Maiter2, Julia Kirchheiner3, Hannah E Ivison, Uwe Fuhr1 and Wiebke Arlt School of Clinical and Experimental Medicine, Centre for Endocrinology, Diabetes and Metabolism (CEDAM), University of Birmingham, Birmingham B15 2TT, UK, 1Department of Pharmacology, University Hospital, University of Cologne, 50931 Cologne, Germany, 2Department of Endocrinology, University Hospital Saint Luc, 1200 Brussels, Belgium and 3Department of Pharmacology of Natural Products and Clinical Pharmacology, University of Ulm, 89019 Ulm, Germany (Correspondence should be addressed to W Arlt; Email: [email protected]) Abstract Objective: Patients with congenital adrenal hyperplasia due to P450 oxidoreductase (POR) deficiency (ORD) present with disordered sex development and glucocorticoid deficiency. This is due to disruption of electron transfer from mutant POR to microsomal cytochrome P450 (CYP) enzymes that play a key role in glucocorticoid and sex steroid synthesis. POR also transfers electrons to all major drug- metabolizing CYP enzymes, including CYP3A4 that inactivates glucocorticoid and oestrogens. However, whether ORD results in impairment of in vivo drug metabolism has never been studied. Design: We studied an adult patient with ORD due to homozygous POR A287P, the most frequent POR mutation in Caucasians, and her clinically unaffected, heterozygous mother. The patient had received standard dose oestrogen replacement from 17 until 37 years of age when it was stopped after she developed breast cancer. Methods: Both subjects underwent in vivo cocktail phenotyping comprising the oral administration of caffeine, tolbutamide, omeprazole, dextromethorphan hydrobromide and midazolam to assess the five major drug-metabolizing CYP enzymes. -

NIH Public Access Author Manuscript Pharmacogenet Genomics
NIH Public Access Author Manuscript Pharmacogenet Genomics. Author manuscript; available in PMC 2013 February 01. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Pharmacogenet Genomics. 2012 February ; 22(2): 159–165. doi:10.1097/FPC.0b013e32834d4962. PharmGKB summary: very important pharmacogene information for cytochrome P450, family 2, subfamily C, polypeptide 19 Stuart A. Scotta, Katrin Sangkuhlc, Alan R. Shuldinere,f, Jean-Sébastien Hulotb,g, Caroline F. Thornc, Russ B. Altmanc,d, and Teri E. Kleinc aDepartment of Genetics and Genomic Sciences bCardiovascular Research Center, Mount Sinai School of Medicine, New York, New York cDepartments of Genetics dBioengineering, Stanford University, Stanford, California eDivision of Endocrinology, Diabetes and Nutrition, University of Maryland School of Medicine fGeriatric Research and Education Clinical Center, Veterans Administration Medical Center, Baltimore, Maryland, USA gDepartment of Pharmacology, Université Pierre et Marie Curie-Paris 6, INSERM UMR S 956, Pitié-Salpêtrière University Hospital, Paris, France Abstract This PharmGKB summary briefly discusses the CYP2C19 gene and current understanding of its function, regulation, and pharmacogenomic relevance. Keywords antidepressants; clopidogrel; CYP2C19*17; CYP2C19*2; CYP2C19; proton pump inhibitors; rs4244285 Introduction The cytochrome P450, family 2, subfamily C, polypeptide 19 (CYP2C19) gene is located within a cluster of cytochrome P450 genes (centromere-CYP2C18-CYP2C19-CYP2C9- CYP2C8-telomere) on chromosome 10q23.33. The CYP2C19 enzyme contributes to the metabolism of a large number of clinically relevant drugs and drug classes such as antidepressants [1], benzodiazepines [2], mephenytoin [3], proton pump inhibitors (PPIs) [4], and the antiplatelet prodrug clopidogrel [5]. Similar to other CYP450 genes, inherited genetic variation in CYP2C19 and its variable hepatic expression contributes to the interindividual phenotypic variability in CYP2C19 substrate metabolism. -

Identification and Developmental Expression of the Full Complement Of
Goldstone et al. BMC Genomics 2010, 11:643 http://www.biomedcentral.com/1471-2164/11/643 RESEARCH ARTICLE Open Access Identification and developmental expression of the full complement of Cytochrome P450 genes in Zebrafish Jared V Goldstone1, Andrew G McArthur2, Akira Kubota1, Juliano Zanette1,3, Thiago Parente1,4, Maria E Jönsson1,5, David R Nelson6, John J Stegeman1* Abstract Background: Increasing use of zebrafish in drug discovery and mechanistic toxicology demands knowledge of cytochrome P450 (CYP) gene regulation and function. CYP enzymes catalyze oxidative transformation leading to activation or inactivation of many endogenous and exogenous chemicals, with consequences for normal physiology and disease processes. Many CYPs potentially have roles in developmental specification, and many chemicals that cause developmental abnormalities are substrates for CYPs. Here we identify and annotate the full suite of CYP genes in zebrafish, compare these to the human CYP gene complement, and determine the expression of CYP genes during normal development. Results: Zebrafish have a total of 94 CYP genes, distributed among 18 gene families found also in mammals. There are 32 genes in CYP families 5 to 51, most of which are direct orthologs of human CYPs that are involved in endogenous functions including synthesis or inactivation of regulatory molecules. The high degree of sequence similarity suggests conservation of enzyme activities for these CYPs, confirmed in reports for some steroidogenic enzymes (e.g. CYP19, aromatase; CYP11A, P450scc; CYP17, steroid 17a-hydroxylase), and the CYP26 retinoic acid hydroxylases. Complexity is much greater in gene families 1, 2, and 3, which include CYPs prominent in metabolism of drugs and pollutants, as well as of endogenous substrates. -

(12) Patent Application Publication (10) Pub. No.: US 2011/0190389 A1 Arterburn Et Al
US 2011 0190389A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2011/0190389 A1 Arterburn et al. (43) Pub. Date: Aug. 4, 2011 (54) OXYLIPINS FROM LONG CHAIN C07C 57/03 (2006.01) POLYUNSATURATED FATTY ACDS AND A6IP 29/00 (2006.01) METHODS OF MAKING AND USING THE CI2P 7/64 (2006.01) SAME CI2P I 7/02 (2006.01) C07D 303/38 (2006.01) (76) Inventors: Linda Arterburn, Ellicott City, A6IP 25/28 (2006.01) MD (US); William Barclay, Boulder, CO (US); Bindi Dangi, (52) U.S. Cl. ......... 514/475: 514/560; 554/124; 435/134; Elkridge, MD (US); James Flatt, 435/123:549/561 Colorado Springs, CO (US); Jung Lee, McLean, VA (US); Dutt (57) ABSTRACT Vinjamoori, Chesterfield, MO Disclosed are novel oxylipins, referred to herein as (US); Mary Van Elswyk, docosanoids and eicosanoids, that are derived from C22 poly Longmont, CO (US) unsaturated fatty acids and from C20 polyunsaturated fatty acids, respectively, and methods of making and using Such (21) Appl. No.: 12/531,344 oxylipins. Also disclosed is the use of docosapentaenoic acid 1-1. (C22:5n-6) (DPAn-6), docosapentaenoic acid (C22:5n-3) (22) PCT Filed: Feb. 20, 2008 (DPAn-3), and docosatetraenoic acid (DTAn-6: C22:4n-6), (86). PCT No.: PCT/USO8/54.456 docosatrienoic acid (C22:3n-3) (DTrAn-3), docosadienoic acid (C22:2n-6) (DDAn-6), eicosatrienoic acid (C20:3n-3) S371 (c)(1), (ETrAn-3) eicosapentaenoic acid and arachidonic acid as (2), (4) Date: Feb. 15, 2011 substrates for the production of novel oxylipins, and to the oxylipins produced thereby. -

Metallothionein Monoclonal Antibody, Clone N11-G
Metallothionein monoclonal antibody, clone N11-G Catalog # : MAB9787 規格 : [ 50 uL ] List All Specification Application Image Product Rabbit monoclonal antibody raised against synthetic peptide of MT1A, Western Blot (Recombinant protein) Description: MT1B, MT1E, MT1F, MT1G, MT1H, MT1IP, MT1L, MT1M, MT2A. Immunogen: A synthetic peptide corresponding to N-terminus of human MT1A, MT1B, MT1E, MT1F, MT1G, MT1H, MT1IP, MT1L, MT1M, MT2A. Host: Rabbit enlarge Reactivity: Human, Mouse Immunoprecipitation Form: Liquid Enzyme-linked Immunoabsorbent Assay Recommend Western Blot (1:1000) Usage: ELISA (1:5000-1:10000) The optimal working dilution should be determined by the end user. Storage Buffer: In 20 mM Tris-HCl, pH 8.0 (10 mg/mL BSA, 0.05% sodium azide) Storage Store at -20°C. Instruction: Note: This product contains sodium azide: a POISONOUS AND HAZARDOUS SUBSTANCE which should be handled by trained staff only. Datasheet: Download Applications Western Blot (Recombinant protein) Western blot analysis of recombinant Metallothionein protein with Metallothionein monoclonal antibody, clone N11-G (Cat # MAB9787). Lane 1: 1 ug. Lane 2: 3 ug. Lane 3: 5 ug. Immunoprecipitation Enzyme-linked Immunoabsorbent Assay ASSP5 MT1A MT1B MT1E MT1F MT1G MT1H MT1M MT1L MT1IP Page 1 of 5 2021/6/2 Gene Information Entrez GeneID: 4489 Protein P04731 (Gene ID : 4489);P07438 (Gene ID : 4490);P04732 (Gene ID : Accession#: 4493);P04733 (Gene ID : 4494);P13640 (Gene ID : 4495);P80294 (Gene ID : 4496);P80295 (Gene ID : 4496);Q8N339 (Gene ID : 4499);Q86YX0 (Gene ID : 4490);Q86YX5 -

Identification and Characterization of CYP2C18 in the Cynomolgus Macaque (Macaca Fascicularis)
NOTE Toxicology Identification and Characterization of CYP2C18 in the Cynomolgus Macaque (Macaca fascicularis) Yasuhiro UNO1)*, Kiyomi MATSUNO1), Chika NAKAMURA1), Masahiro UTOH1) and Hiroshi YAMAZAKI2) 1)Pharmacokinetics and Bioanalysis Center (PBC), Shin Nippon Biomedical Laboratories (SNBL), Kainan, Wakayama 642–0017 and 2)Laboratory of Drug Metabolism and Pharmacokinetics, Showa Pharmaceutical University, Machida, Tokyo 194–8543, Japan (Received 3 August 2009/Accepted 15 September 2009/Published online in J-STAGE 25 November 2009) ABSTRACT. The macaque is widely used for investigation of drug metabolism due to its evolutionary closeness to the human. However, the genetic backgrounds of drug-metabolizing enzymes have not been fully investigated; therefore, identification and characterization of drug-metabolizing enzyme genes are important for understanding drug metabolism in this species. In this study, we isolated and char- acterized a novel cytochrome P450 2C18 (CYP2C18) cDNA in cynomolgus macaques. This cDNA was highly homologous (96%) to human CYP2C18 cDNA. Cynomolgus CYP2C18 was preferentially expressed in the liver and kidney. Moreover, a metabolic assay using cynomolgus CYP2C18 protein heterologously expressed in Escherichia coli revealed its activity toward S-mephenytoin 4’-hydrox- ylation. These results suggest that cynomolgus CYP2C18 could function as a drug-metabolizing enzyme in the liver. KEY WORDS: cloning, CYP2C18, cytochrome P450, liver, monkey. J. Vet. Med. Sci. 72(2): 225–228, 2010 Cytochrome P450 (CYP) is a superfamily of some of the Such species differences also could be explained by func- most important drug-metabolizing enzymes and consists of tional disparity of the orthologous CYPs between the two a large number of subfamilies [14]. In humans, the CYP2C species, such as, if an ortholog to human CYP with low (or subfamilies contain important enzymes that metabolize no) expression and drug-metabolizing activity, for example approximately 20% of all prescribed drugs [3]. -

CD56+ T-Cells in Relation to Cytomegalovirus in Healthy Subjects and Kidney Transplant Patients
CD56+ T-cells in Relation to Cytomegalovirus in Healthy Subjects and Kidney Transplant Patients Institute of Infection and Global Health Department of Clinical Infection, Microbiology and Immunology Thesis submitted in accordance with the requirements of the University of Liverpool for the degree of Doctor in Philosophy by Mazen Mohammed Almehmadi December 2014 - 1 - Abstract Human T cells expressing CD56 are capable of tumour cell lysis following activation with interleukin-2 but their role in viral immunity has been less well studied. The work described in this thesis aimed to investigate CD56+ T-cells in relation to cytomegalovirus infection in healthy subjects and kidney transplant patients (KTPs). Proportions of CD56+ T cells were found to be highly significantly increased in healthy cytomegalovirus-seropositive (CMV+) compared to cytomegalovirus-seronegative (CMV-) subjects (8.38% ± 0.33 versus 3.29%± 0.33; P < 0.0001). In donor CMV-/recipient CMV- (D-/R-)- KTPs levels of CD56+ T cells were 1.9% ±0.35 versus 5.42% ±1.01 in D+/R- patients and 5.11% ±0.69 in R+ patients (P 0.0247 and < 0.0001 respectively). CD56+ T cells in both healthy CMV+ subjects and KTPs expressed markers of effector memory- RA T-cells (TEMRA) while in healthy CMV- subjects and D-/R- KTPs the phenotype was predominantly that of naïve T-cells. Other surface markers, CD8, CD4, CD58, CD57, CD94 and NKG2C were expressed by a significantly higher proportion of CD56+ T-cells in healthy CMV+ than CMV- subjects. Functional studies showed levels of pro-inflammatory cytokines IFN-γ and TNF-α, as well as granzyme B and CD107a were significantly higher in CD56+ T-cells from CMV+ than CMV- subjects following stimulation with CMV antigens. -

Investigation of Structural Properties of Methylated Human Promoter Regions in Terms of Dna Helical Rise
INVESTIGATION OF STRUCTURAL PROPERTIES OF METHYLATED HUMAN PROMOTER REGIONS IN TERMS OF DNA HELICAL RISE A THESIS SUBMITTED TO THE GRADUATE SCHOOL OF INFORMATICS OF MIDDLE EAST TECHNICAL UNIVERSITY BY BURCU YALDIZ IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF MASTER OF SCIENCE IN BIOINFORMATICS AUGUST 2014 INVESTIGATION OF STRUCTURAL PROPERTIES OF METHYLATED HUMAN PROMOTER REGIONS IN TERMS OF DNA HELICAL RISE submitted by Burcu YALDIZ in partial fulfillment of the requirements for the degree of Master of Science, Bioinformatics Program, Middle East Technical University by, Prof. Dr. Nazife Baykal _____________________ Director, Informatics Institute Assist. Prof. Dr. Yeşim Aydın Son _____________________ Head of Department, Health Informatics, METU Assist. Prof. Dr. Yeşim Aydın Son Supervisor, Health Informatics, METU _____________________ Examining Committee Members: Assoc. Prof. Dr. Tolga Can _____________________ METU, CENG Assist. Prof. Dr. Yeşim Aydın Son _____________________ METU, Health Informatics Assist. Prof. Dr. Aybar Can Acar _____________________ METU, Health Informatics Assist. Prof. Dr. Özlen Konu _____________________ Bilkent University, Molecular Biology and Genetics Assoc. Prof. Dr. Çağdaş D. Son _____________________ METU, Biology Date: 27.08.2014 I hereby declare that all information in this document has been obtained and presented in accordance with academic rules and ethical conduct. I also declare that, as required by these rules and conduct, I have fully cited and referenced all material and results that are not original to this work. Name, Last name : Burcu Yaldız Signature : iii ABSTRACT INVESTIGATION OF STRUCTURAL PROPERTIES OF METHYLATED HUMAN PROMOTER REGIONS IN TERMS OF DNA HELICAL RISE Yaldız, Burcu M.Sc. Bioinformatics Program Advisor: Assist. Prof. Dr. Yeşim Aydın Son August 2014, 60 pages The infamous double helix structure of DNA was assumed to be a rigid, uniformly observed structure throughout the genomic DNA. -

Synonymous Single Nucleotide Polymorphisms in Human Cytochrome
DMD Fast Forward. Published on February 9, 2009 as doi:10.1124/dmd.108.026047 DMD #26047 TITLE PAGE: A BIOINFORMATICS APPROACH FOR THE PHENOTYPE PREDICTION OF NON- SYNONYMOUS SINGLE NUCLEOTIDE POLYMORPHISMS IN HUMAN CYTOCHROME P450S LIN-LIN WANG, YONG LI, SHU-FENG ZHOU Department of Nutrition and Food Hygiene, School of Public Health, Peking University, Beijing 100191, P. R. China (LL Wang & Y Li) Discipline of Chinese Medicine, School of Health Sciences, RMIT University, Bundoora, Victoria 3083, Australia (LL Wang & SF Zhou). 1 Copyright 2009 by the American Society for Pharmacology and Experimental Therapeutics. DMD #26047 RUNNING TITLE PAGE: a) Running title: Prediction of phenotype of human CYPs. b) Author for correspondence: A/Prof. Shu-Feng Zhou, MD, PhD Discipline of Chinese Medicine, School of Health Sciences, RMIT University, WHO Collaborating Center for Traditional Medicine, Bundoora, Victoria 3083, Australia. Tel: + 61 3 9925 7794; fax: +61 3 9925 7178. Email: [email protected] c) Number of text pages: 21 Number of tables: 10 Number of figures: 2 Number of references: 40 Number of words in Abstract: 249 Number of words in Introduction: 749 Number of words in Discussion: 1459 d) Non-standard abbreviations: CYP, cytochrome P450; nsSNP, non-synonymous single nucleotide polymorphism. 2 DMD #26047 ABSTRACT Non-synonymous single nucleotide polymorphisms (nsSNPs) in coding regions that can lead to amino acid changes may cause alteration of protein function and account for susceptivity to disease. Identification of deleterious nsSNPs from tolerant nsSNPs is important for characterizing the genetic basis of human disease, assessing individual susceptibility to disease, understanding the pathogenesis of disease, identifying molecular targets for drug treatment and conducting individualized pharmacotherapy. -

A Bayesian Approach to Mediation Analysis Predicts 206 Causal Target Genes in Alzheimer’S Disease
bioRxiv preprint first posted online Nov. 14, 2017; doi: http://dx.doi.org/10.1101/219428. The copyright holder for this preprint (which was not peer-reviewed) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Title A Bayesian approach to mediation analysis predicts 206 causal target genes in Alzheimer’s disease Authors Yongjin Park+;1;2, Abhishek K Sarkar+;3, Liang He+;1;2, Jose Davila-Velderrain1;2, Philip L De Jager2;4, Manolis Kellis1;2 +: equal contribution. 1: Computer Science and Artificial Intelligence Laboratory, Massachusetts Institute of Technology, Cambridge, MA, USA 2: Broad Institute of MIT and Harvard, Cambridge, MA, USA 3: Department of Human Genetics, University of Chicago, Chicago, IL, USA 4: Department of Neurology, Columbia University Medical Center, New York, NY, USA MK: [email protected] Abstract Characterizing the intermediate phenotypes, such as gene expression, that mediate genetic effects on complex dis- eases is a fundamental problem in human genetics. Existing methods utilize genotypic data and summary statistics to identify putative disease genes, but cannot distinguish pleiotropy from causal mediation and are limited by overly strong assumptions about the data. To overcome these limitations, we develop Causal Multivariate Mediation within Extended Linkage disequilibrium (CaMMEL), a novel Bayesian inference framework to jointly model multiple medi- ated and unmediated effects relying only on summary statistics. We show in simulation that CaMMEL accurately distinguishes between mediating and pleiotropic genes unlike existing methods. We applied CaMMEL to Alzheimer’s disease (AD) and found 206 causal genes in sub-threshold loci (p < 10−4). -
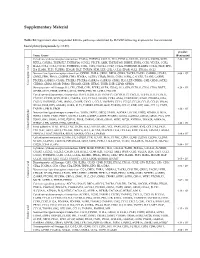
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R,