BMP Signaling Orchestrates a Transcriptional Network to Control the Fate of Mesenchymal Stem Cells (Mscs)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Genome-Wide DNA Methylation Analysis of KRAS Mutant Cell Lines Ben Yi Tew1,5, Joel K
www.nature.com/scientificreports OPEN Genome-wide DNA methylation analysis of KRAS mutant cell lines Ben Yi Tew1,5, Joel K. Durand2,5, Kirsten L. Bryant2, Tikvah K. Hayes2, Sen Peng3, Nhan L. Tran4, Gerald C. Gooden1, David N. Buckley1, Channing J. Der2, Albert S. Baldwin2 ✉ & Bodour Salhia1 ✉ Oncogenic RAS mutations are associated with DNA methylation changes that alter gene expression to drive cancer. Recent studies suggest that DNA methylation changes may be stochastic in nature, while other groups propose distinct signaling pathways responsible for aberrant methylation. Better understanding of DNA methylation events associated with oncogenic KRAS expression could enhance therapeutic approaches. Here we analyzed the basal CpG methylation of 11 KRAS-mutant and dependent pancreatic cancer cell lines and observed strikingly similar methylation patterns. KRAS knockdown resulted in unique methylation changes with limited overlap between each cell line. In KRAS-mutant Pa16C pancreatic cancer cells, while KRAS knockdown resulted in over 8,000 diferentially methylated (DM) CpGs, treatment with the ERK1/2-selective inhibitor SCH772984 showed less than 40 DM CpGs, suggesting that ERK is not a broadly active driver of KRAS-associated DNA methylation. KRAS G12V overexpression in an isogenic lung model reveals >50,600 DM CpGs compared to non-transformed controls. In lung and pancreatic cells, gene ontology analyses of DM promoters show an enrichment for genes involved in diferentiation and development. Taken all together, KRAS-mediated DNA methylation are stochastic and independent of canonical downstream efector signaling. These epigenetically altered genes associated with KRAS expression could represent potential therapeutic targets in KRAS-driven cancer. Activating KRAS mutations can be found in nearly 25 percent of all cancers1. -

The Regulation of Lunatic Fringe During Somitogenesis
THE REGULATION OF LUNATIC FRINGE DURING SOMITOGENESIS DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Emily T. Shifley ***** The Ohio State University 2009 Dissertation Committee: Approved by Professor Susan Cole, Advisor Professor Christine Beattie _________________________________ Professor Mark Seeger Advisor Graduate Program in Molecular Genetics Professor Michael Weinstein ABSTRACT Somitogenesis is the morphological hallmark of vertebrate segmentation. Somites bud from the presomitic mesoderm (PSM) in a sequential, periodic fashion and give rise to the rib cage, vertebrae, and dermis and muscles of the back. The regulation of somitogenesis is complex. In the posterior region of the PSM, a segmentation clock operates to organize cohorts of cells into presomites, while in the anterior region of the PSM the presomites are patterned into rostral and caudal compartments (R/C patterning). Both of these stages of somitogenesis are controlled, at least in part, by the Notch pathway and Lunatic fringe (Lfng), a glycosyltransferase that modifies the Notch receptor. To dissect the roles played by Lfng during somitogenesis, we created a novel allele that lacks cyclic Lfng expression within the segmentation clock, but that maintains expression during R/C somite patterning (Lfng∆FCE1). Lfng∆FCE1/∆FCE1 mice have severe defects in their anterior vertebrae and rib cages, but relatively normal sacral and tail vertebrae, unlike Lfng knockouts. Segmentation clock function is differentially affected by the ∆FCE1 deletion; during anterior somitogenesis the expression patterns of many clock genes are disrupted, while during posterior somitogenesis, certain clock components have recovered. R/C patterning occurs relatively normally in Lfng∆FCE1/∆FCE1 embryos, likely contributing to the partial phenotype rescue, and confirming that Lfng ii plays separate roles in the two regions of the PSM. -

NKX2-8 Antibody (Pab)
21.10.2014NKX2-8 antibody (pAb) Rabbit Anti -Human/Mouse/Rat NK2 homeobox 8 (NKXH, NKX2 -9) Instruction Manual Catalog Number PK-AB718-6753 Synonyms NKX2-8 Antibody: NK2 homeobox 8, NKXH, NKX2-9 Description NKX2-8 (NK2 homeobox 8) is a member of a family of transcription factors that are involved in embryonic development and cell fate. It is expressed in the ventral foregut, the developing heart, the epithelial layers of the branchial arches and in the dorsal mesoderm. In conjunction with related protein, NKX2-5, NKX2-8 may play a role in cardiac embryonic development. NKX2-8 is also thought to be involved in lung development and is suspected of being an oncogene in lung cancer that is activated by way of gene amplification at chromosome 14q13. Quantity 100 µg Source / Host Rabbit Immunogen NKX2-8 antibody was raised against a 19 amino acid synthetic peptide near the center of human NKX2-8. Purification Method Affinity chromatography purified via peptide column. Clone / IgG Subtype Polyclonal antibody Species Reactivity Human, Mouse, Rat Specificity NKX2-8 antibody is predicted to not cross-react with other NK2 homeobox family members. Formulation Antibody is supplied in PBS containing 0.02% sodium azide. Reconstitution During shipment, small volumes of antibody will occasionally become entrapped in the seal of the product vial. For products with volumes of 200 μl or less, we recommend gently tapping the vial on a hard surface or briefly centrifuging the vial in a tabletop centrifuge to dislodge any liquid in the container’s cap. Storage & Stability Antibody can be stored at 4ºC for three months and at -20°C for up to one year. -

Single Cell Regulatory Landscape of the Mouse Kidney Highlights Cellular Differentiation Programs and Disease Targets
ARTICLE https://doi.org/10.1038/s41467-021-22266-1 OPEN Single cell regulatory landscape of the mouse kidney highlights cellular differentiation programs and disease targets Zhen Miao 1,2,3,8, Michael S. Balzer 1,2,8, Ziyuan Ma 1,2,8, Hongbo Liu1,2, Junnan Wu 1,2, Rojesh Shrestha 1,2, Tamas Aranyi1,2, Amy Kwan4, Ayano Kondo 4, Marco Pontoglio 5, Junhyong Kim6, ✉ Mingyao Li 7, Klaus H. Kaestner2,4 & Katalin Susztak 1,2,4 1234567890():,; Determining the epigenetic program that generates unique cell types in the kidney is critical for understanding cell-type heterogeneity during tissue homeostasis and injury response. Here, we profile open chromatin and gene expression in developing and adult mouse kidneys at single cell resolution. We show critical reliance of gene expression on distal regulatory elements (enhancers). We reveal key cell type-specific transcription factors and major gene- regulatory circuits for kidney cells. Dynamic chromatin and expression changes during nephron progenitor differentiation demonstrates that podocyte commitment occurs early and is associated with sustained Foxl1 expression. Renal tubule cells follow a more complex differentiation, where Hfn4a is associated with proximal and Tfap2b with distal fate. Mapping single nucleotide variants associated with human kidney disease implicates critical cell types, developmental stages, genes, and regulatory mechanisms. The single cell multi-omics atlas reveals key chromatin remodeling events and gene expression dynamics associated with kidney development. 1 Renal, Electrolyte, and Hypertension Division, Department of Medicine, University of Pennsylvania, Perelman School of Medicine, Philadelphia, PA, USA. 2 Institute for Diabetes, Obesity, and Metabolism, University of Pennsylvania, Perelman School of Medicine, Philadelphia, PA, USA. -

Genome-Wide DNA Methylation Analysis Reveals Molecular Subtypes of Pancreatic Cancer
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 17), pp: 28990-29012 Research Paper Genome-wide DNA methylation analysis reveals molecular subtypes of pancreatic cancer Nitish Kumar Mishra1 and Chittibabu Guda1,2,3,4 1Department of Genetics, Cell Biology and Anatomy, University of Nebraska Medical Center, Omaha, NE, 68198, USA 2Bioinformatics and Systems Biology Core, University of Nebraska Medical Center, Omaha, NE, 68198, USA 3Department of Biochemistry and Molecular Biology, University of Nebraska Medical Center, Omaha, NE, 68198, USA 4Fred and Pamela Buffet Cancer Center, University of Nebraska Medical Center, Omaha, NE, 68198, USA Correspondence to: Chittibabu Guda, email: [email protected] Keywords: TCGA, pancreatic cancer, differential methylation, integrative analysis, molecular subtypes Received: October 20, 2016 Accepted: February 12, 2017 Published: March 07, 2017 Copyright: Mishra et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC-BY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Pancreatic cancer (PC) is the fourth leading cause of cancer deaths in the United States with a five-year patient survival rate of only 6%. Early detection and treatment of this disease is hampered due to lack of reliable diagnostic and prognostic markers. Recent studies have shown that dynamic changes in the global DNA methylation and gene expression patterns play key roles in the PC development; hence, provide valuable insights for better understanding the initiation and progression of PC. In the current study, we used DNA methylation, gene expression, copy number, mutational and clinical data from pancreatic patients. -

Automatic Detection of the Circulating Cell-Free Methylated DNA Pattern
Supplementary Files Automatic Detection of the Circulating Cell‐Free Methylated DNA Pattern of GCM2, ITPRIPL1 and CCDC181 for Detection of Early Breast Cancer and Surgical Treatment Response Sheng‐Chao Wang, Li‐Min Liao, Muhamad Ansar, Shih‐Yun Lin, Wei‐Wen Hsu and Chih‐Ming Su, Yu‐Mei Chung, Cai‐Cing Liu, Chin‐Sheng Hung and Ruo‐Kai Lin Figure S1. Representative standard sequencing diagram for bisulfite direct sequencing of the CCDC181, GCM2, ITPRIPL1, ENPP2, LOC643719, ZNF177, ADCY4 and RASSF1 genes. Figure S2. Representative figures showing the DNA methylation levels of the candidate genes CCDC181, GCM2, ITPRIPL1, ENPP2, LOC643719, ZNF177, ADCY4 and RASSF1 using qMSP in breast cancer patients. Table S1. The clinical parameters of breast cancer patients for plasma ccfDNA analysis. Automatic Automatic Manual Manual Labturbo Duo Prime Characteristics 0.2 mL Plasma 0.5 mL Plasma 1.6 mL Plasma 1.6 mL Plasma N (%) N (%) N (%) N (%) Overall 45 34 63 57 Type 45 19(42.2) 13(38.2) 9(14.3) 10(17.5) 45 26(57.8) 21(61.8) 54(85.7) 47(82.5) Type DCIS 3(6.7) 10(29.4) 7(11.1) 1(1.8) IDC 40(88.9) 22(64.7) 47(74.6) 48(84.2) ILC 1(2.2) 2(5.9) 1(1.6) 2(3.5) Others 1(2.2) 0 8(12.7) 6(10.5) Tumor Stage 0, I and II 32(71.1) 29(85.3) 52(82.5) 44(77.2) III and IV 13(28.9) 5(14.7) 11(17.5) 13(22.8) Tumor Size T0–T2 38(84.4) 31(91.2) 56(88.9) 49(86.0) T3–T4 7(15.6) 3(8.8) 7(11.1) 8(14.0) Lymph node N = 0 16(35.6) 22(64.7) 31(49.2) 30(52.6) N > 0 29(64.4) 1235.3) 32(50.8) 27(47.4) ER Negative 21(46.7) 7(20.6) 19(30.2) 19(33.3) Positive 24(53.3) 27(79.4) 44(69.8) 38(66.7) PR Negative 20(44.4) 11(32.4) 21(33.3) 22(38.6) Positive 25(55.6) 23(67.6) 42(66.7) 35(61.4) HER2 Negative 35(77.8) 26(76.5) 46(73.0) 35(61.4) Positive 10(22.2) 8(23.5) 17(27.0) 22(38.6) Ki‐67 High 24(53.3) 24(70.6) 39(61.9) 39(68.4) Low 21(46.7) 10(29.4) 24(38.1) 12(21.1) n.d. -

An OTX2-PAX3 Signaling Axis Regulates Group 3 Medulloblastoma Cell Fate
ARTICLE https://doi.org/10.1038/s41467-020-17357-4 OPEN An OTX2-PAX3 signaling axis regulates Group 3 medulloblastoma cell fate Jamie Zagozewski1, Ghazaleh M. Shahriary 1, Ludivine Coudière Morrison1, Olivier Saulnier 2,3, Margaret Stromecki1, Agnes Fresnoza4, Gareth Palidwor5, Christopher J. Porter 5, Antoine Forget6,7, Olivier Ayrault 6,7, Cynthia Hawkins2,8,9, Jennifer A. Chan10, Maria C. Vladoiu2,3,9, Lakshmikirupa Sundaresan2,3, Janilyn Arsenio11,12, Michael D. Taylor 2,3,9,13, Vijay Ramaswamy 2,3,14,15 & ✉ Tamra E. Werbowetski-Ogilvie 1 1234567890():,; OTX2 is a potent oncogene that promotes tumor growth in Group 3 medulloblastoma. However, the mechanisms by which OTX2 represses neural differentiation are not well characterized. Here, we perform extensive multiomic analyses to identify an OTX2 regulatory network that controls Group 3 medulloblastoma cell fate. OTX2 silencing modulates the repressive chromatin landscape, decreases levels of PRC2 complex genes and increases the expression of neurodevelopmental transcription factors including PAX3 and PAX6. Expression of PAX3 and PAX6 is significantly lower in Group 3 medulloblastoma patients and is cor- related with reduced survival, yet only PAX3 inhibits self-renewal in vitro and increases survival in vivo. Single cell RNA sequencing of Group 3 medulloblastoma tumorspheres demonstrates expression of an undifferentiated progenitor program observed in primary tumors and characterized by translation/elongation factor genes. Identification of mTORC1 signaling as a downstream effector of OTX2-PAX3 reveals roles for protein synthesis pathways in regulating Group 3 medulloblastoma pathogenesis. 1 Regenerative Medicine Program, Department of Biochemistry and Medical Genetics, University of Manitoba, Winnipeg, MB, Canada. 2 The Arthur and Sonia Labatt Brain Tumour Research Center, The Hospital for Sick Children, Toronto, ON, Canada. -

Mutational Analysis of AXIN2, MSX1, and PAX9 in Two Mexican Oligodontia Families
Mutational analysis of AXIN2, MSX1, and PAX9 in two Mexican oligodontia families Y.D. Mu1,2, Z. Xu1, C.I. Contreras1, J.S. McDaniel1, K.J. Donly1 and S. Chen1 1Department of Developmental Dentistry, Dental School, University of Texas Health Science Center, San Antonio, Texas, USA 2Stomotology Department, Sichuan Provincial People’s Hospital, Chengdu, Sichuan, China Corresponding author: S. Chen E-mail: [email protected] Genet. Mol. Res. 12 (4): 4446-4458 (2013) Received July 4, 2012 Accepted May 15, 2013 Published October 10, 2013 DOI http://dx.doi.org/10.4238/2013.October.10.10 ABSTRACT. The genes for axin inhibition protein 2 (AXIN2), msh homeobox 1 (MSX1), and paired box gene 9 (PAX9) are involved in tooth root formation and tooth development. Mutations of the AXIN2, MSX1, and PAX9 genes are associated with non-syndromic oligodontia. In this study, we investigated phenotype and AXIN2, MSX1, and PAX9 gene variations in two Mexican families with non-syndromic oligodontia. Individuals from two families underwent clinical examinations, including an intra-oral examination and panoramic radiograph. Retrospective data were reviewed, and peripheral blood samples were collected. The exons and exon-intronic boundaries of the AXIN2, MSX1, and PAX9 genes were sequenced and analyzed. Protein and messenger RNA structures were predicted using bioinformative software programs. Clinical and oral examinations revealed isolated non-syndromic oligodontia in the two Mexican families. The average number of missing teeth was 12. The sequence analysis of exons and exon-intronic regions of AXIN2, MSX1, and PAX9 revealed 11 single- nucleotide polymorphisms (SNPs), including seven in AXIN2, two Genetics and Molecular Research 12 (4): 4446-4458 (2013) ©FUNPEC-RP www.funpecrp.com.br AXIN2, MSX1, and PAX9 SNPs and oligodontia 4447 in MSX1, and three in PAX9. -
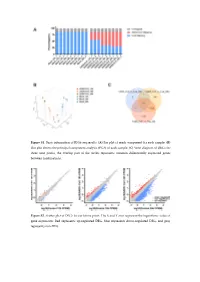
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

Pancreatic Α- and Β-Cellular Clocks Have Distinct Molecular Properties and Impact on Islet Hormone Secretion and Gene Expression
Downloaded from genesdev.cshlp.org on October 4, 2021 - Published by Cold Spring Harbor Laboratory Press Pancreatic α- and β-cellular clocks have distinct molecular properties and impact on islet hormone secretion and gene expression Volodymyr Petrenko,1,2,3 Camille Saini,1,2,3,9 Laurianne Giovannoni,1,2,3,9 Cedric Gobet,4,5,9 Daniel Sage,6 Michael Unser,6 Mounia Heddad Masson,1 Guoqiang Gu,7 Domenico Bosco,8 Frédéric Gachon,4,5,10 Jacques Philippe,1,10 and Charna Dibner1,2,3 1Endocrinology, Diabetes, Hypertension, and Nutrition, University Hospital of Geneva, CH-1211 Geneva, Switzerland; 2Department of Cellular Physiology and Metabolism, Diabetes Center, Faculty of Medicine, University of Geneva, CH-1211 Geneva, Switzerland; 3Institute of Genetics and Genomics in Geneva (iGE3), University of Geneva, CH-1211 Geneva, Switzerland; 4Department of Diabetes and Circadian Rhythms, Nestlé Institute of Health Sciences, CH-1015 Lausanne, Switzerland; 5School of Life Sciences, Ecole Polytechnique Fédérale de Lausanne (EPFL), CH-1015 Lausanne, Switzerland; 6Biomedical Imaging Group, EPFL, CH-1015 Lausanne, Switzerland; 7Department of Cell and Developmental Biology, Vanderbilt University Medical Center, Nashville, Tennessee 37240, USA; 8Department of Surgery, Cell Isolation and Transplantation Centre, University Hospital of Geneva, CH-1211 Geneva, Switzerland A critical role of circadian oscillators in orchestrating insulin secretion and islet gene transcription has been dem- onstrated recently. However, these studies focused on whole islets and did not explore the interplay between α-cell and β-cell clocks. We performed a parallel analysis of the molecular properties of α-cell and β-cell oscillators using a mouse model expressing three reporter genes: one labeling α cells, one specific for β cells, and a third monitoring circadian gene expression. -

Evaluation of the Role of Egfr, Sox2, and Pax9 in Oral
EVALUATION OF THE ROLE OF EGFR, SOX2, AND PAX9 IN ORAL CARCINOGENESIS Timothy John Bates A thesis submitted in partial fulfilment of the requirements for the degree of Doctor of Philosophy Northern Institute for Cancer Research Newcastle University August 2015 Acknowledgements I would like to acknowledge my three supervisors, Dr M. Robinson, Dr R. Kist, and Professor P. Sloan, without whom this project would not have been possible. Dr Kist and Dr Robinson shared the initial vision for this project and, in collaboration, designed and developed its overall structure. The project therefore arose from a unique collaboration between principle investigators with backgrounds in developmental oral biology and oral pathology. Professor Sloan’s experience and insight have proved invaluable to the analysis, interpretation, and presentation of our data. I would like to thank Miss A. Long and all members of the Research and Development Team, Department of Cellular Pathology, for their essential contribution to the processing, sectioning, and automated immunostaining of both human and mouse tissue during this project. Dr R. Mahsoub and Miss V. Rook contributed to the manual immunohistochemical staining of sections during this project, work which formed part of their Masters by Research projects. Mr M. Kennedy played a vital role in collecting clinical data for the group of early-stage oral squamous cell carcinoma patients, during a Master of Science project that he undertook at the University of Edinburgh. I would like to thank Miss M. Elias and all the team in the Dermatology Laboratory, Institute of Cellular Medicine. The generation of the tetracycline- inducible cell lines would not have been possible without the support of Martina and others in the Dermatology team: Carol, Keith, and Emma, to name just a few.