Frontotemporal Dementia: Insights Into the Biological Underpinnings of Disease Through Gene Co-Expression Network Analysis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Multiple Activities of Arl1 Gtpase in the Trans-Golgi Network Chia-Jung Yu1,2 and Fang-Jen S
© 2017. Published by The Company of Biologists Ltd | Journal of Cell Science (2017) 130, 1691-1699 doi:10.1242/jcs.201319 COMMENTARY Multiple activities of Arl1 GTPase in the trans-Golgi network Chia-Jung Yu1,2 and Fang-Jen S. Lee3,4,* ABSTRACT typical features of an Arf-family GTPase, including an amphipathic ADP-ribosylation factors (Arfs) and ADP-ribosylation factor-like N-terminal helix and a consensus site for N-myristoylation (Lu et al., proteins (Arls) are highly conserved small GTPases that function 2001; Price et al., 2005). In yeast, recruitment of Arl1 to the Golgi as main regulators of vesicular trafficking and cytoskeletal complex requires a second Arf-like GTPase, Arl3 (Behnia et al., reorganization. Arl1, the first identified member of the large Arl family, 2004; Setty et al., 2003). Yeast Arl3 lacks a myristoylation site and is an important regulator of Golgi complex structure and function in is, instead, N-terminally acetylated; this modification is required for organisms ranging from yeast to mammals. Together with its effectors, its recruitment to the Golgi complex by Sys1. In mammalian cells, Arl1 has been shown to be involved in several cellular processes, ADP-ribosylation-factor-related protein 1 (Arfrp1), a mammalian including endosomal trans-Golgi network and secretory trafficking, lipid ortholog of yeast Arl3, plays a pivotal role in the recruitment of Arl1 droplet and salivary granule formation, innate immunity and neuronal to the trans-Golgi network (TGN) (Behnia et al., 2004; Panic et al., development, stress tolerance, as well as the response of the unfolded 2003b; Setty et al., 2003; Zahn et al., 2006). -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

TMEM106B in Humans and Vac7 and Tag1 in Yeast Are Predicted to Be Lipid Transfer Proteins
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.12.435176; this version posted March 12, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. TMEM106B in humans and Vac7 and Tag1 in yeast are predicted to be lipid transfer proteins Tim P. Levine* UCL Institute of Ophthalmology, 11-43 Bath Street, London EC1V 9EL, United Kingdom. ORCID 0000-0002-7231-0775 *Corresponding author and lead contact: [email protected] Data availability statement: The data that support this study are freely available in Harvard Dataverse at https://dataverse.harvard.edu/dataverse/LEA_2. Acknowledgements: work was funded by the Higher Education Funding Council for England and the NIHR Moorfields Biomedical Research Centre Conflict of interest disclosure: the author declares that there is no conflict of interest Keywords: Structural bioinformatics, Lipid transfer protein, LEA_2, TMEM106B, Vac7, YLR173W, Endosome, Lysosome Running Title: TMEM106B & Vac7: lipid transfer proteins 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.03.12.435176; this version posted March 12, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract TMEM106B is an integral membrane protein of late endosomes and lysosomes involved in neuronal function, its over-expression being associated with familial frontotemporal lobar degeneration, and under-expression linked to hypomyelination. It has also been identified in multiple screens for host proteins required for productive SARS-CoV2 infection. Because standard approaches to understand TMEM106B at the sequence level find no homology to other proteins, it has remained a protein of unknown function. -

Eif2b Is a Decameric Guanine Nucleotide Exchange Factor with a G2e2 Tetrameric Core
ARTICLE Received 19 Feb 2014 | Accepted 15 Apr 2014 | Published 23 May 2014 DOI: 10.1038/ncomms4902 OPEN eIF2B is a decameric guanine nucleotide exchange factor with a g2e2 tetrameric core Yuliya Gordiyenko1,2,*, Carla Schmidt1,*, Martin D. Jennings3, Dijana Matak-Vinkovic4, Graham D. Pavitt3 & Carol V. Robinson1 eIF2B facilitates and controls protein synthesis in eukaryotes by mediating guanine nucleotide exchange on its partner eIF2. We combined mass spectrometry (MS) with chemical cross- linking, surface accessibility measurements and homology modelling to define subunit stoichiometry and interactions within eIF2B and eIF2. Although it is generally accepted that eIF2B is a pentamer of five non-identical subunits (a–e), here we show that eIF2B is a decamer. MS and cross-linking of eIF2B complexes allows us to propose a model for the subunit arrangements within eIF2B where the subunit assembly occurs through catalytic g- and e-subunits, with regulatory subunits arranged in asymmetric trimers associated with the core. Cross-links between eIF2 and eIF2B allow modelling of interactions that contribute to nucleotide exchange and its control by eIF2 phosphorylation. Finally, we identify that GTP binds to eIF2Bg, prompting us to propose a multi-step mechanism for nucleotide exchange. 1 Department of Chemistry, Physical and Theoretical Chemistry Laboratory, University of Oxford, South Parks Road, Oxford OX1 3QZ, UK. 2 MRC Laboratory of Molecular Biology, University of Cambridge, Francis Crick Avenue, Cambridge CB2 0QH, UK. 3 Faculty of Life Sciences, Michael Smith Building, University of Manchester, Oxford Road, Manchester M13 9PT, UK. 4 Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge CB2 1EW, UK. -

Cofactor Dependent Conformational Switching of Gtpases
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector 1704 Biophysical Journal Volume 95 August 2008 1704–1715 Cofactor Dependent Conformational Switching of GTPases Vasili Hauryliuk,*y Sebastian Hansson,z and Ma˚ns Ehrenberg* *Department of Cell and Molecular Biology, Molecular Biology Program, Uppsala University, Uppsala, Sweden; yEngelhardt Institute of Molecular Biology, Russian Academy of Sciences, Moscow, Russia; and zLaboratoire d’Enzymologie et Biochimie Structurales, CNRS, Avenue de la Terrasse, 91198 Gif-sur-Yvette, France ABSTRACT This theoretical work covers structural and biochemical aspects of nucleotide binding and GDP/GTP exchange of GTP hydrolases belonging to the family of small GTPases. Current models of GDP/GTP exchange regulation are often based on two specific assumptions. The first is that the conformation of a GTPase is switched by the exchange of the bound nucleotide from GDP to GTP or vice versa. The second is that GDP/GTP exchange is regulated by a guanine nucleotide exchange factor, which stabilizes a GTPase conformation with low nucleotide affinity. Since, however, recent biochemical and structural data seem to contradict this view, we present a generalized scheme for GTPase action. This novel ansatz accounts for those important cases when conformational switching in addition to guanine nucleotide exchange requires the presence of cofactors, and gives a more nuanced picture of how the nucleotide exchange is regulated. The scheme is also used to discuss some problems of interpretation that may arise when guanine nucleotide exchange mechanisms are inferred from experiments with analogs of GTP, like GDPNP, GDPCP, and GDP g S. -

A Dissertation Entitled Characterization of the CXCR4
A Dissertation entitled Characterization of the CXCR4-LASP1-eIF4F Axis in Triple-Negative Breast Cancer by Cory M Howard Submitted to the Graduate Faculty as partial fulfillment of the requirements for the Doctor of Philosophy Degree in Biomedical Sciences ___________________________________________ Dayanidhi Raman, B.V.Sc., Ph.D., Committee Chair ___________________________________________ Amit Tiwari, Ph.D., Committee Member ___________________________________________ Ritu Chakravarti, Ph.D., Committee Member ___________________________________________ Nagalakshmi Nadiminty, Ph.D., Committee Member ___________________________________________ Saori Furuta, Ph.D., Committee Member ___________________________________________ Shi-He Liu, M.D., Committee Member ___________________________________________ Amanda C. Bryant-Friedrich, Ph.D., Dean College of Graduate Studies The University of Toledo August 2020 © 2020 Cory M. Howard This document is copyrighted material. Under copyright law, no parts of this document may be reproduced without the expressed permission of the author. An Abstract of Characterization of the CXCR4-LASP1-eIF4F Axis in Triple-Negative Breast Cancer by Cory M. Howard Submitted to the Graduate Faculty as partial fulfillment of the requirements for the Doctor of Philosophy Degree in Biomedical Sciences The University of Toledo August 2020 Triple-negative breast cancer (TNBC) remains clinically challenging as effective targeted therapies are still lacking. In addition, patient mortality mainly results from the metastasized -

Exploring the Relationship Between Gut Microbiota and Major Depressive Disorders
E3S Web of Conferences 271, 03055 (2021) https://doi.org/10.1051/e3sconf/202127103055 ICEPE 2021 Exploring the Relationship between Gut Microbiota and Major Depressive Disorders Catherine Tian1 1Shanghai American School, Shanghai, China Abstract. Major Depressive Disorder (MDD) is a psychiatric disorder accompanied with a high rate of suicide, morbidity and mortality. With the symptom of an increasing or decreasing appetite, there is a possibility that MDD may have certain connections with gut microbiota, the colonies of microbes which reside in the human digestive system. In recent years, more and more studies started to demonstrate the links between MDD and gut microbiota from animal disease models and human metabolism studies. However, this relationship is still largely understudied, but it is very innovative since functional dissection of this relationship would furnish a new train of thought for more effective treatment of MDD. In this study, by using multiple genetic analytic tools including Allen Brain Atlas, genetic function analytical tools, and MicrobiomeAnalyst, I explored the genes that shows both expression in the brain and the digestive system to affirm that there is a connection between gut microbiota and the MDD. My approach finally identified 7 MDD genes likely to be associated with gut microbiota, implicating 3 molecular pathways: (1) Wnt Signaling, (2) citric acid cycle in the aerobic respiration, and (3) extracellular exosome signaling. These findings may shed light on new directions to understand the mechanism of MDD, potentially facilitating the development of probiotics for better psychiatric disorder treatment. 1 Introduction 1.1 Major Depressive Disorder Major Depressive Disorder (MDD) is a mood disorder that will affect the mood, behavior and other physical parts. -
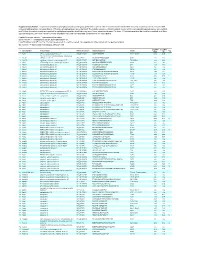
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

Anti-TMEM106A (RABBIT) Antibody - 600-401-FH1
Anti-TMEM106A (RABBIT) Antibody - 600-401-FH1 Code: 600-401-FH1 Size: 100 µg Product Description: Anti-TMEM106A (RABBIT) Antibody - 600-401-FH1 Concentration: 1 mg/mL by UV absorbance at 280 nm PhysicalState: Liquid (sterile filtered) Label Unconjugated Host Rabbit Gene Name TMEM106A Species Reactivity Human, mouse Buffer 0.01 M Sodium Phosphate, 0.25 M Sodium Chloride, pH 7.2 Stabilizer None Preservative 0.02% (w/v) Sodium Azide Storage Condition Store vial at -20° C prior to opening. Aliquot contents and freeze at -20° C or below for extended storage. Avoid cycles of freezing and thawing. Centrifuge product if not completely clear after standing at room temperature. This product is stable for several weeks at 4° C as an undiluted liquid. Dilute only prior to immediate use. Synonyms TMEM106A Antibody, Transmembrane protein 106A Application Note Anti-TMEM106A Antibody has been tested for use in ELISA, Western Blotting, Immunohistochemistry and Immunofluorescence. Specific conditions for reactivity should be optimized by the end user. Expect a band at approximately 29 kDa in Western Blots of specific cell lysates and tissues. Background Transmembrane protein 106A (TMEM106A) is a single-pass transmembrane protein that is closely related to TMEM106B, a protein that is thought to be a novel risk factor for frontotemporal lobar degeneration (FTLD), a group of clinically, pathologically and genetically heterogeneous disorders associated with atrophy in the frontal lobe and temporal lobe of the brain. The actual roles of TMEM106A and TMEM106B are still undetermined; however, as TMEM106B is involved in FTLD, it is possible that TMEM106A may also be a risk factor for FTLD. -

Genetic Regulation of Tmem106b in the Pathogenesis of Frontotemporal Lobar Degeneration
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2017 Genetic Regulation Of Tmem106b In The Pathogenesis Of Frontotemporal Lobar Degeneration Michael Gallagher University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Genetics Commons, Molecular Biology Commons, and the Neuroscience and Neurobiology Commons Recommended Citation Gallagher, Michael, "Genetic Regulation Of Tmem106b In The Pathogenesis Of Frontotemporal Lobar Degeneration" (2017). Publicly Accessible Penn Dissertations. 2294. https://repository.upenn.edu/edissertations/2294 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/2294 For more information, please contact [email protected]. Genetic Regulation Of Tmem106b In The Pathogenesis Of Frontotemporal Lobar Degeneration Abstract Neurodegenerative diseases are an emerging global health crisis, with the projected global cost of dementia alone expected to exceed $1 trillion, or >1% of world GDP, by 2018. However, there are no disease-modifying treatments for the major neurodegenerative diseases, such as Alzheimer’s disease, Parkinson’s disease, frontotemporal lobar degeneration (FTLD), and amyotrophic lateral sclerosis. Therefore, there is an urgent need for a better understanding of the pathophysiology underlying these diseases. While genome-wide association studies (GWAS) have identified ~200 genetic ariantsv that are associated with risk of developing neurodegenerative disease, the biological mechanisms underlying these associations are largely unknown. This dissertation investigates the mechanisms by which common genetic variation at TMEM106B, a GWAS-identified risk locus for FTLD, influences disease risk. First, using genetic and clinical data from thirty American and European medical centers, I demonstrate that the TMEM106B locus acts as a genetic modifier of a common Mendelian form of FTLD. -

A Systematic Genetic Assessment of ARFGEF2 Mutations in Periventricular Heterotopia
International Journal of Genetics and Genomics 2018; 6(1): 11-17 http://www.sciencepublishinggroup.com/j/ijgg doi: 10.11648/j.ijgg.20180601.13 ISSN: 2376-7340 (Print); ISSN: 2376-7359 (Online) A Systematic Genetic Assessment of ARFGEF2 Mutations in Periventricular Heterotopia Lina Al Neghery 1, 2, Rosan Kenana 1, Albandary Al Bakheet 1, Rawan Al Mass 1, Faten Al Mutairi 1, Maysoon Al Sagob 1, Aliya Qari 3, Rozeena Huma 3, Dilek Colak 4, Maha Daghestani 2, Namik Kaya 1, Moeenaldeen Al Sayed 3, * 1Department of Genetics, King Faisal Specialist Hospital and Research Center, Riyadh, Kingdom of Saudi Arabia 2Department of Zoology, College of Science, King Saud University, Riyadh, Kingdom of Saudi Arabia 3Department of Medical Genetics, King Faisal Specialist Hospital and Research Center, Riyadh, Kingdom of Saudi Arabia 4Department of Biostatistics, Epidemiology and Scientific Computing, King Faisal Specialist Hospital and Research Center, Riyadh, Kingdom of Saudi Arabia Email address: *Corresponding author To cite this article: Lina Al Neghery, Rosan Kenana, Albandary Al Bakheet, Rawan Al Mass, Faten Al Mutairi, Maysoon Al Sagob, Aliya Qari, Rozeena Huma, Dilek Colak, Maha Daghestani, Namik Kaya, Moeenaldeen Al Sayed. A Systematic Genetic Assessment of ARFGEF2 Mutations in Periventricular Heterotopia. International Journal of Genetics and Genomics . Vol. 6, No. 1, 2018, pp. 11-17. doi: 10.11648/j.ijgg.20180601.13 Received : January 12, 2018; Accepted : January 29, 2018; Published : May 5, 2018 Abstract: Mutations in ADP-ribosylation factor guanine nucleotide-exchange factor 2 ( ARFGEF2 ) lead to autosomal recessive periventricular heterotopia (PH). To date, 11 mutations, (six missense mutations, one splicing mutation, one small deletion, two small insertions, and one small deletion/insertion) have been reported. -

Nck Adapter Proteins: Functional Versatility in T Cells Marcus Lettau, Jennifer Pieper and Ottmar Janssen
Cell Communication and Signaling BioMed Central Review Open Access Nck adapter proteins: functional versatility in T cells Marcus Lettau, Jennifer Pieper and Ottmar Janssen Address: 1University Hospital Schleswig-Holstein Campus Kiel, Institute of Immunology, Molecular Immunology, Arnold-Heller-Str 3, Bldg 17, D-24105 Kiel, Germany E-mail: Marcus Lettau* - [email protected]; Jennifer Pieper - [email protected]; Ottmar Janssen - [email protected] *Corresponding author Published: 02 February 2009 Received: 2 December 2008 Cell Communication and Signaling 2009, 7:1 doi: 10.1186/1478-811X-7-1 Accepted: 2 February 2009 This article is available from: http://www.biosignaling.com/content/7/1/1 © 2009 Lettau et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Nck is a ubiquitously expressed adapter protein that is almost exclusively built of one SH2 domain and three SH3 domains. The two isoproteins of Nck are functionally redundant in many aspects and differ in only few amino acids that are mostly located in the linker regions between the interaction modules. Nck proteins connect receptor and non-receptor tyrosine kinases to the machinery of actin reorganisation. Thereby, Nck regulates activation-dependent processes during cell polarisation and migration and plays a crucial role in the signal transduction of a variety of receptors including for instance PDGF-, HGF-, VEGF- and Ephrin receptors. In most cases, the SH2 domain mediates binding to the phosphorylated receptor or associated phosphoproteins, while SH3domaininteractionsleadtotheformationoflargerproteincomplexes.InTlymphocytes,Nck plays a pivotal role in the T cell receptor (TCR)-induced reorganisation of the actin cytoskeleton and the formation of the immunological synapse.