Gli3 Utilizes Hand2 to Synergistically Regulate Tissue-Specific Transcriptional Networks 5 6 Kelsey H
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Uterine Double-Conditional Inactivation of Smad2 and Smad3 in Mice Causes Endometrial Dysregulation, Infertility, and Uterine Cancer
Uterine double-conditional inactivation of Smad2 and Smad3 in mice causes endometrial dysregulation, infertility, and uterine cancer Maya Krisemana,b, Diana Monsivaisa,c, Julio Agnoa, Ramya P. Masanda, Chad J. Creightond,e, and Martin M. Matzuka,c,f,g,h,1 aDepartment of Pathology and Immunology, Baylor College of Medicine, Houston, TX 77030; bReproductive Endocrinology and Infertility, Baylor College of Medicine/Texas Children’s Hospital Women’s Pavilion, Houston, TX 77030; cCenter for Drug Discovery, Baylor College of Medicine, Houston, TX 77030; dDepartment of Medicine, Baylor College of Medicine, Houston, TX 77030; eDan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, TX 77030; fDepartment of Molecular and Cellular Biology, Baylor College of Medicine, Houston, TX 77030; gDepartment of Molecular and Human Genetics, Baylor College of Medicine, Houston, TX 77030; and hDepartment of Pharmacology and Chemical Biology, Baylor College of Medicine, Houston, TX 77030 Contributed by Martin M. Matzuk, December 6, 2018 (sent for review April 30, 2018; reviewed by Milan K. Bagchi and Thomas E. Spencer) SMAD2 and SMAD3 are downstream proteins in the transforming in endometrial function. Notably, members of the transforming growth factor-β (TGF β) signaling pathway that translocate signals growth factor β (TGF β) family are involved in many cellular from the cell membrane to the nucleus, bind DNA, and control the processes and serve as principal regulators of numerous biological expression of target genes. While SMAD2/3 have important roles functions, including female reproduction. Previous studies have in the ovary, we do not fully understand the roles of SMAD2/3 in shown the TGF β family to have key roles in ovarian folliculo- the uterus and their implications in the reproductive system. -

Ncomms2254.Pdf
ARTICLE Received 3 May 2012 | Accepted 5 Nov 2012 | Published 4 Dec 2012 DOI: 10.1038/ncomms2254 The RB family is required for the self-renewal and survival of human embryonic stem cells Jamie F. Conklin1,2,3, Julie Baker2,3 & Julien Sage1,2,3 The mechanisms ensuring the long-term self-renewal of human embryonic stem cells are still only partly understood, limiting their use in cellular therapies. Here we found that increased activity of the RB cell cycle inhibitor in human embryonic stem cells induces cell cycle arrest, differentiation and cell death. Conversely, inactivation of the entire RB family (RB, p107 and p130) in human embryonic stem cells triggers G2/M arrest and cell death through functional activation of the p53 pathway and the cell cycle inhibitor p21. Differences in E2F target gene activation upon loss of RB family function between human embryonic stem cells, mouse embryonic stem cells and human fibroblasts underscore key differences in the cell cycle regulatory networks of human embryonic stem cells. Finally, loss of RB family function pro- motes genomic instability in both human and mouse embryonic stem cells, uncoupling cell cycle defects from chromosomal instability. These experiments indicate that a homeostatic level of RB activity is essential for the self-renewal and the survival of human embryonic stem cells. 1 Department of Pediatrics, Stanford Medical School, Stanford, California 94305, USA. 2 Department of Genetics, Stanford Medical School, Stanford, California 94305, USA. 3 Institute for Stem Cell Biology and Regenerative Medicine, Stanford Medical School, Stanford, California 94305, USA. Correspondence and requests for materials should be addressed to J.S. -

Prospective Isolation of NKX2-1–Expressing Human Lung Progenitors Derived from Pluripotent Stem Cells
The Journal of Clinical Investigation RESEARCH ARTICLE Prospective isolation of NKX2-1–expressing human lung progenitors derived from pluripotent stem cells Finn Hawkins,1,2 Philipp Kramer,3 Anjali Jacob,1,2 Ian Driver,4 Dylan C. Thomas,1 Katherine B. McCauley,1,2 Nicholas Skvir,1 Ana M. Crane,3 Anita A. Kurmann,1,5 Anthony N. Hollenberg,5 Sinead Nguyen,1 Brandon G. Wong,6 Ahmad S. Khalil,6,7 Sarah X.L. Huang,3,8 Susan Guttentag,9 Jason R. Rock,4 John M. Shannon,10 Brian R. Davis,3 and Darrell N. Kotton1,2 2 1Center for Regenerative Medicine, and The Pulmonary Center and Department of Medicine, Boston University School of Medicine, Boston, Massachusetts, USA. 3Center for Stem Cell and Regenerative Medicine, Brown Foundation Institute of Molecular Medicine, University of Texas Health Science Center, Houston, Texas, USA. 4Department of Anatomy, UCSF, San Francisco, California, USA. 5Division of Endocrinology, Diabetes and Metabolism, Beth Israel Deaconess Medical Center and Harvard Medical School, Boston, Massachusetts, USA. 6Department of Biomedical Engineering and Biological Design Center, Boston University, Boston, Massachusetts, USA. 7Wyss Institute for Biologically Inspired Engineering, Harvard University, Boston, Massachusetts, USA. 8Columbia Center for Translational Immunology & Columbia Center for Human Development, Columbia University Medical Center, New York, New York, USA. 9Department of Pediatrics, Monroe Carell Jr. Children’s Hospital, Vanderbilt University, Nashville, Tennessee, USA. 10Division of Pulmonary Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA. It has been postulated that during human fetal development, all cells of the lung epithelium derive from embryonic, endodermal, NK2 homeobox 1–expressing (NKX2-1+) precursor cells. -

Table S1 the Four Gene Sets Derived from Gene Expression Profiles of Escs and Differentiated Cells
Table S1 The four gene sets derived from gene expression profiles of ESCs and differentiated cells Uniform High Uniform Low ES Up ES Down EntrezID GeneSymbol EntrezID GeneSymbol EntrezID GeneSymbol EntrezID GeneSymbol 269261 Rpl12 11354 Abpa 68239 Krt42 15132 Hbb-bh1 67891 Rpl4 11537 Cfd 26380 Esrrb 15126 Hba-x 55949 Eef1b2 11698 Ambn 73703 Dppa2 15111 Hand2 18148 Npm1 11730 Ang3 67374 Jam2 65255 Asb4 67427 Rps20 11731 Ang2 22702 Zfp42 17292 Mesp1 15481 Hspa8 11807 Apoa2 58865 Tdh 19737 Rgs5 100041686 LOC100041686 11814 Apoc3 26388 Ifi202b 225518 Prdm6 11983 Atpif1 11945 Atp4b 11614 Nr0b1 20378 Frzb 19241 Tmsb4x 12007 Azgp1 76815 Calcoco2 12767 Cxcr4 20116 Rps8 12044 Bcl2a1a 219132 D14Ertd668e 103889 Hoxb2 20103 Rps5 12047 Bcl2a1d 381411 Gm1967 17701 Msx1 14694 Gnb2l1 12049 Bcl2l10 20899 Stra8 23796 Aplnr 19941 Rpl26 12096 Bglap1 78625 1700061G19Rik 12627 Cfc1 12070 Ngfrap1 12097 Bglap2 21816 Tgm1 12622 Cer1 19989 Rpl7 12267 C3ar1 67405 Nts 21385 Tbx2 19896 Rpl10a 12279 C9 435337 EG435337 56720 Tdo2 20044 Rps14 12391 Cav3 545913 Zscan4d 16869 Lhx1 19175 Psmb6 12409 Cbr2 244448 Triml1 22253 Unc5c 22627 Ywhae 12477 Ctla4 69134 2200001I15Rik 14174 Fgf3 19951 Rpl32 12523 Cd84 66065 Hsd17b14 16542 Kdr 66152 1110020P15Rik 12524 Cd86 81879 Tcfcp2l1 15122 Hba-a1 66489 Rpl35 12640 Cga 17907 Mylpf 15414 Hoxb6 15519 Hsp90aa1 12642 Ch25h 26424 Nr5a2 210530 Leprel1 66483 Rpl36al 12655 Chi3l3 83560 Tex14 12338 Capn6 27370 Rps26 12796 Camp 17450 Morc1 20671 Sox17 66576 Uqcrh 12869 Cox8b 79455 Pdcl2 20613 Snai1 22154 Tubb5 12959 Cryba4 231821 Centa1 17897 -

Expression of the Human PAC1 Receptor Leads to Dose- Dependent Hydrocephalus-Related Abnormalities in Mice
Expression of the human PAC1 receptor leads to dose- dependent hydrocephalus-related abnormalities in mice Bing Lang, … , Anthony J. Harmar, Sanbing Shen J Clin Invest. 2006;116(7):1924-1934. https://doi.org/10.1172/JCI27597. Research Article Neuroscience Hydrocephalus is a common and potentially devastating birth defect affecting the CNS, and its relationship with G protein–coupled receptors (GPCRs) is unknown. We have expressed 2, 4, or 6 copies of a GPCR — the human PAC1 receptor with a 130-kb transgene in the mouse nervous system in a pattern closely resembling that of the endogenous gene. Consistent with PAC1 actions, PKA and PKC activity were elevated in the brains of Tg mice. Remarkably, Tg mice developed dose-dependent hydrocephalus-like characteristics, including enlarged third and lateral ventricles and reduced cerebral cortex, corpus callosum, and subcommissural organ (SCO). Neuronal proliferation and apoptosis were implicated in hydrocephalus, and we observed significantly reduced neuronal proliferation and massively increased neuronal apoptosis in the developing cortex and SCO of Tg embryos, while neurite outgrowth and neuronal migration in vitro remain uncompromised. Ventricular ependymal cilia are crucial for directing cerebrospinal fluid flow, and ependyma of Tg mice exhibited disrupted cilia with increased phospho-CREB immunoreactivity. These data demonstrate that altered neuronal proliferation/apoptosis and disrupted ependymal cilia are the main factors contributing to hydrocephalus in PAC1-overexpressing mice. This is the first report to our knowledge demonstrating that misregulation of GPCRs can be involved in hydrocephalus-related neurodevelopmental disorders. Find the latest version: https://jci.me/27597/pdf Research article Related Commentary, page 1828 Expression of the human PAC1 receptor leads to dose-dependent hydrocephalus- related abnormalities in mice Bing Lang,1 Bing Song,1 Wendy Davidson,1 Alastair MacKenzie,1 Norman Smith,2 Colin D. -

Co-Occupancy by Multiple Cardiac Transcription Factors Identifies
Co-occupancy by multiple cardiac transcription factors identifies transcriptional enhancers active in heart Aibin Hea,b,1, Sek Won Konga,b,c,1, Qing Maa,b, and William T. Pua,b,2 aDepartment of Cardiology and cChildren’s Hospital Informatics Program, Children’s Hospital Boston, Boston, MA 02115; and bHarvard Stem Cell Institute, Harvard University, Cambridge, MA 02138 Edited by Eric N. Olson, University of Texas Southwestern, Dallas, TX, and approved February 23, 2011 (received for review November 12, 2010) Identification of genomic regions that control tissue-specific gene study of a handful of model genes (e.g., refs. 7–10), it has not been expression is currently problematic. ChIP and high-throughput se- evaluated using unbiased, genome-wide approaches. quencing (ChIP-seq) of enhancer-associated proteins such as p300 In this study, we used a modified ChIP-seq approach to define identifies some but not all enhancers active in a tissue. Here we genome wide the binding sites of these cardiac TFs (1). We show that co-occupancy of a chromatin region by multiple tran- provide unbiased support for collaborative TF interactions in scription factors (TFs) identifies a distinct set of enhancers. GATA- driving cardiac gene expression and use this principle to show that chromatin co-occupancy by multiple TFs identifies enhancers binding protein 4 (GATA4), NK2 transcription factor-related, lo- with cardiac activity in vivo. The majority of these multiple TF- cus 5 (NKX2-5), T-box 5 (TBX5), serum response factor (SRF), and “ binding loci (MTL) enhancers were distinct from p300-bound myocyte-enhancer factor 2A (MEF2A), here referred to as cardiac enhancers in location and functional properties. -

To Proliferate Or to Die: Role of Id3 in Cell Cycle Progression and Survival of Neural Crest Progenitors
Downloaded from genesdev.cshlp.org on October 4, 2021 - Published by Cold Spring Harbor Laboratory Press To proliferate or to die: role of Id3 in cell cycle progression and survival of neural crest progenitors Yun Kee and Marianne Bronner-Fraser1 Division of Biology, California Institute of Technology, Pasadena, California 91125, USA The neural crest is a unique population of mitotically active, multipotent progenitors that arise at the vertebrate neural plate border. Here, we show that the helix–loop–helix transcriptional regulator Id3 has a novel role in cell cycle progression and survival of neural crest progenitors in Xenopus. Id3 is localized at the neural plate border during gastrulation and neurulation, overlapping the domain of neural crest induction. Morpholino oligonucleotide-mediated depletion of Id3 results in the absence of neural crest precursors and a resultant loss of neural crest derivatives. This appears to be mediated by cell cycle inhibition followed by cell death of the neural crest progenitor pool, rather than a cell fate switch. Conversely, overexpression of Id3 increases cell proliferation and results in expansion of the neural crest domain. Our data suggest that Id3 functions by a novel mechanism, independent of cell fate determination, to mediate the decision of neural crest precursors to proliferate or die. [Keywords: Xenopus; Id3; neural crest; cell cycle; survival] Received September 1, 2004; revised version accepted January 19, 2005. The neural crest is an embryonic cell population that origi- et al. 2003), c-Myc (Bellmeyer et al. 2003), and Msx1 nates from the lateral edges of the neural plate during (Tribulo et al. 2003) have been identified in Xenopus nervous system formation. -

Lncmyod Promotes Skeletal Myogenesis and Regulates Skeletal Muscle Fiber-Type Composition by Sponging Mir-370-3P
G C A T T A C G G C A T genes Article LncMyoD Promotes Skeletal Myogenesis and Regulates Skeletal Muscle Fiber-Type Composition by Sponging miR-370-3p Peiwen Zhang 1,2,† , Jingjing Du 1,2,†, Xinyu Guo 1,2, Shuang Wu 1,2, Jin He 1,2 , Xinrong Li 1,2, Linyuan Shen 1,2 , Lei Chen 1,2, Bohong Li 1, Jingjun Zhang 1, Yuhao Xie 1, Lili Niu 1,2, Dongmei Jiang 1,2, Xuewei Li 1,2, Shunhua Zhang 1,2 and Li Zhu 1,2,* 1 College of Animal Science and Technology, Sichuan Agricultural University, Chengdu 611130, China; [email protected] (P.Z.); [email protected] (J.D.); [email protected] (X.G.); [email protected] (S.W.); [email protected] (J.H.); [email protected] (X.L.); [email protected] (L.S.); [email protected] (L.C.); [email protected] (B.L.); [email protected] (J.Z.); [email protected] (Y.X.); [email protected] (L.N.); [email protected] (D.J.); [email protected] (X.L.); [email protected] (S.Z.) 2 Farm Animal Genetic Resources Exploration and Innovation Key Laboratory of Sichuan Province, Sichuan Agricultural University, Chengdu 611130, China * Correspondence: [email protected] † These authors contributed equally to this work. Abstract: The development of skeletal muscle is a highly ordered and complex biological process. Increasing evidence has shown that noncoding RNAs, especially long-noncoding RNAs (lncRNAs) and microRNAs, play a vital role in the development of myogenic processes. -

HAND2‑Mediated Proteolysis Negatively Regulates the Function of Estrogen Receptor Α
5538 MOLECULAR MEDICINE REPORTS 12: 5538-5544, 2015 HAND2‑mediated proteolysis negatively regulates the function of estrogen receptor α TOMOHIKO FUKUDA*, AKIRA SHIRANE*, OSAMU WADA-HIRAIKE, KATSUTOSHI ODA, MICHIHIRO TANIKAWA, AYAKO SAKUABASHI, MANA HIRANO, HOUJU FU, YOSHIHIRO MORITA, YUICHIRO MIYAMOTO, KANAKO INABA, KEI KAWANA, YUTAKA OSUGA and TOMOYUKI FUJII Department of Obstetrics and Gynecology, Graduate School of Medicine, The University of Tokyo, Tokyo 113-8655, Japan Received October 2, 2014; Accepted June 11, 2015 DOI: 10.3892/mmr.2015.4070 Abstract. A previous study demonstrated that the transcriptional activation function of ERα was possibly attrib- progesterone-inducible HAND2 gene product is a basic uted to the proteasomic degradation of ERα by HAND2. These helix-loop-helix transcription factor and prevents mitogenic results indicate a novel anti-tumorigenic function of HAND2 effects of estrogen receptor α (ERα) by inhibiting fibroblast in regulating ERα-dependent gene expression. Considering growth factor signalling in mouse uteri. However, whether that HAND2 is commonly hypermethylated and silenced in HAND2 directly affects the transcriptional activation func- endometrial cancer, it is hypothesized that HAND2 may serve tion of ERα remains to be elucidated. In the present study, as a possible tumor suppressor, particularly in uterine tissue. the physical interaction between HAND2 and ERα was investigating by performing an immunoprecipitation assay Introduction and an in vitro pull-down assay. The results demonstrated that HAND2 and ERα interacted in a ligand-independent manner. Endometrial cancer is one of the most common types of The in vitro pull-down assays revealed a direct interaction gynecologic malignancy, increasing each year (1). Based on between HAND2 and the amino-terminus of ERα, termed a pathological view, endometrial cancer can be divided into the activation function-1 domain. -
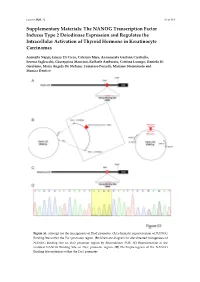
The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas
Cancers 2020, 12 S1 of S18 Supplementary Materials: The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas Annarita Nappi, Emery Di Cicco, Caterina Miro, Annunziata Gaetana Cicatiello, Serena Sagliocchi, Giuseppina Mancino, Raffaele Ambrosio, Cristina Luongo, Daniela Di Girolamo, Maria Angela De Stefano, Tommaso Porcelli, Mariano Stornaiuolo and Monica Dentice Figure S1. Strategy for the mutagenesis of Dio2 promoter. (A) Schematic representation of NANOG Binding Site within the Dio2 promoter region. (B) Schematic diagram for site‐directed mutagenesis of NANOG Binding Site on Dio2 promoter region by Recombinant PCR. (C) Representation of the mutated NANOG Binding Site on Dio2 promoter region. (D) Electropherogram of the NANOG Binding Site mutation within the Dio2 promoter. Cancers 2020, 12 S2 of S18 Figure S2. Strategy for the silencing of NANOG expression. (A) Cloning strategies for the generation of NANOG shRNA expression vectors. (B) Electropherograms of the NANOG shRNA sequences cloned into pcDNA3.1 vector. (C) Validation of effective NANOG down-modulation by two different NANOG shRNA vectors was assessed by Western Blot analysis of NANOG expression in BCC cells. (D) Quantification of NANOG protein levels versus Tubulin levels in the same experiment as in C is represented by histograms. Cancers 2020, 12 S3 of S18 Figure S3. The CD34+ cells are characterized by the expression of typical epithelial stemness genes. The mRNA levels of a panel of indicated stemness markers of epidermis were measured by Real Time PCR in the same experiment indicated in figure 3F and G. Cancers 2020, 12 S4 of S18 Figure S4. -

Genetic Factors in Nonsyndromic Orofacial Clefts
Published online: 2021-02-12 THIEME Review Article 101 Genetic Factors in Nonsyndromic Orofacial Clefts Mahamad Irfanulla Khan1 Prashanth C.S.2 Narasimha Murthy Srinath3 1 Department of Orthodontics & Dentofacial Orthopedics, The Oxford Address for correspondence Mahamad Irfanulla Khan, BDS, MDS, Dental College, Bangalore, Karnataka, India Department of Orthodontics & Dentofacial Orthopedics, The Oxford 2 Department of Orthodontics & Dentofacial Orthopedics, DAPM R.V. Dental College, Bangalore, Karnataka 560068, India Dental College, Bangalore, Karnataka, India (e-mail: [email protected]). 3 Department of Oral & Maxillofacial Surgery, Krishnadevaraya College of Dental Sciences, Bangalore, Karnataka, India Global Med Genet 2020;7:101–108. Abstract Orofacial clefts (OFCs) are the most common congenital birth defects in humans and immediately recognized at birth. The etiology remains complex and poorly understood and seems to result from multiple genetic and environmental factors along with gene– environment interactions. It can be classified into syndromic (30%) and nonsyndromic (70%) clefts. Nonsyndromic OFCs include clefts without any additional physical or Keywords cognitive deficits. Recently, various genetic approaches, such as genome-wide associ- ► orofacial clefts ation studies (GWAS), candidate gene association studies, and linkage analysis, have ► nonsyndromic identified multiple genes involved in the etiology of OFCs. ► genetics This article provides an insight into the multiple genes involved in the etiology of OFCs. ► gene mutation Identification of specific genetic causes of clefts helps in a better understanding of the ► genome-wide molecular pathogenesis of OFC. In the near future, it helps to provide a more accurate association study diagnosis, genetic counseling, personalized medicine for better clinical care, and ► linkage analysis prevention of OFCs. -

Tumor and Stem Cell Biology Research
Published OnlineFirst September 17, 2012; DOI: 10.1158/0008-5472.CAN-12-0614 Cancer Tumor and Stem Cell Biology Research OTX2 Represses Myogenic and Neuronal Differentiation in Medulloblastoma Cells Ren-Yuan Bai1, Verena Staedtke1, Hart G. Lidov3, Charles G. Eberhart2, and Gregory J. Riggins1 Abstract The brain development transcription factor OTX2 is overexpressed and/or genomically amplified in most medulloblastomas, but the mechanistic basis for its contributions in this setting are not understood. In this study, we identified OTX2 as a transcriptional repressor and a gatekeeper of myogenic and neuronal differentiation in medulloblastoma cells. OTX2 binds to the MyoD1 core enhancer through its homeobox domain, and the remarkable repressor activity exhibited by the homeobox domain renders OTX2 transcrip- tionally repressive. RNA interference–mediated attenuation of OTX2 expression triggered myogenic and neuronal differentiation in vitro and prolonged the survival in an orthotopic medulloblastoma mouse model. Conversely, inducing myogenic conversion of medulloblastoma cells led to the loss of OTX2 expression. In medullomyoblastoma, a medulloblastoma subtype containing muscle elements, myogenic cells share cytogenetic signatures with the primitive tumor cells and OTX2 expression was lost in the differentiated myogenic cells. Thus, OTX2 functions via its homeobox domain as a suppressor of differentiation, and the loss of OTX2 expression is linked to the myogenesis in medullomyoblastoma. Together, our findings illustrate the origin of muscle cells in medullomyoblastomas and the oncogenic mechanism of OTX2 as a repressor of diverse differentiating potential. Cancer Res; 72(22); 1–14. Ó2012 AACR. Introduction is featured with rhabdomyoblastic elements immunoreactive Medulloblastoma, the most common type of pediatric brain to desmin, myoglobin, and fast myosin (3).