Loss of FOXC1 Contributes to the Corneal Epithelial Fate Switch and Pathogenesis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Id Proteins Promote a Cancer Stem Cell Phenotype in Triple Negative Breast
bioRxiv preprint doi: https://doi.org/10.1101/497313; this version posted March 11, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. 1 Id proteins promote a cancer stem cell phenotype in triple negative 2 breast cancer via negative regulation of Robo1 3 Wee S. Teo1,2, Holly Holliday1,2#, Nitheesh Karthikeyan3#, Aurélie S. Cazet1,2, Daniel L. 4 Roden1,2, Kate Harvey1, Christina Valbirk Konrad1, Reshma Murali3, Binitha Anu Varghese3, 5 Archana P. T.3,4, Chia-Ling Chan1,2, Andrea McFarland1, Simon Junankar1,2, Sunny Ye1, 6 Jessica Yang1, Iva Nikolic1,2, Jaynish S. Shah5, Laura A. Baker1,2, Ewan K.A. Millar1,6,7,8, 7 Mathew J. Naylor1,2,10, Christopher J. Ormandy1,2, Sunil R. Lakhani11, Warren Kaplan1,12, 8 Albert S. Mellick13,14, Sandra A. O’Toole1,9, Alexander Swarbrick1,2*, Radhika Nair1,2,3* 9 Affiliations 10 1Garvan Institute of Medical Research, Darlinghurst, New South Wales, Australia 11 2St Vincent’s Clinical School, Faculty of Medicine, UNSW Sydney, New South Wales, 12 Australia 13 3 Cancer Research Program, Rajiv Gandhi Centre for Biotechnology, Kerala, India 14 4Manipal Academy of Higher Education, Manipal, Karnataka, India 15 5Centenary Institute, The University of Sydney, New South Wales, Australia 16 6 Department of Anatomical Pathology, NSW Health Pathology, St George Hospital, 17 Kogarah, NSW, Australia 18 7 School of Medical Sciences, UNSW Sydney, Kensington NSW, Australia. -

A Molecular Switch from STAT2-IRF9 to ISGF3 Underlies Interferon-Induced Gene Transcription
ARTICLE https://doi.org/10.1038/s41467-019-10970-y OPEN A molecular switch from STAT2-IRF9 to ISGF3 underlies interferon-induced gene transcription Ekaterini Platanitis 1, Duygu Demiroz1,5, Anja Schneller1,5, Katrin Fischer1, Christophe Capelle1, Markus Hartl 1, Thomas Gossenreiter 1, Mathias Müller2, Maria Novatchkova3,4 & Thomas Decker 1 Cells maintain the balance between homeostasis and inflammation by adapting and inte- grating the activity of intracellular signaling cascades, including the JAK-STAT pathway. Our 1234567890():,; understanding of how a tailored switch from homeostasis to a strong receptor-dependent response is coordinated remains limited. Here, we use an integrated transcriptomic and proteomic approach to analyze transcription-factor binding, gene expression and in vivo proximity-dependent labelling of proteins in living cells under homeostatic and interferon (IFN)-induced conditions. We show that interferons (IFN) switch murine macrophages from resting-state to induced gene expression by alternating subunits of transcription factor ISGF3. Whereas preformed STAT2-IRF9 complexes control basal expression of IFN-induced genes (ISG), both type I IFN and IFN-γ cause promoter binding of a complete ISGF3 complex containing STAT1, STAT2 and IRF9. In contrast to the dogmatic view of ISGF3 formation in the cytoplasm, our results suggest a model wherein the assembly of the ISGF3 complex occurs on DNA. 1 Max Perutz Labs (MPL), University of Vienna, Vienna 1030, Austria. 2 Institute of Animal Breeding and Genetics, University of Veterinary Medicine Vienna, Vienna 1210, Austria. 3 Institute of Molecular Biotechnology of the Austrian Academy of Sciences (IMBA), Vienna 1030, Austria. 4 Research Institute of Molecular Pathology (IMP), Vienna Biocenter (VBC), Vienna 1030, Austria. -

Genome-Scale Identification of Transcription Factors That Mediate An
ARTICLE DOI: 10.1038/s41467-018-04406-2 OPEN Genome-scale identification of transcription factors that mediate an inflammatory network during breast cellular transformation Zhe Ji 1,2,4, Lizhi He1, Asaf Rotem1,2,5, Andreas Janzer1,6, Christine S. Cheng2,7, Aviv Regev2,3 & Kevin Struhl 1 Transient activation of Src oncoprotein in non-transformed, breast epithelial cells can initiate an epigenetic switch to the stably transformed state via a positive feedback loop that involves 1234567890():,; the inflammatory transcription factors STAT3 and NF-κB. Here, we develop an experimental and computational pipeline that includes 1) a Bayesian network model (AccessTF) that accurately predicts protein-bound DNA sequence motifs based on chromatin accessibility, and 2) a scoring system (TFScore) that rank-orders transcription factors as candidates for being important for a biological process. Genetic experiments validate TFScore and suggest that more than 40 transcription factors contribute to the oncogenic state in this model. Interestingly, individual depletion of several of these factors results in similar transcriptional profiles, indicating that a complex and interconnected transcriptional network promotes a stable oncogenic state. The combined experimental and computational pipeline represents a general approach to comprehensively identify transcriptional regulators important for a biological process. 1 Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115, USA. 2 Broad Institute of MIT and Harvard, Cambridge, MA 02142, USA. 3 Department of Biology, Howard Hughes Medical Institute and David H. Koch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, MA 20140, USA. 4Present address: Department of Pharmacology and Biomedical Engineering, Northwestern University, Evanston 60611 IL, USA. -

Analysis of the Indacaterol-Regulated Transcriptome in Human Airway
Supplemental material to this article can be found at: http://jpet.aspetjournals.org/content/suppl/2018/04/13/jpet.118.249292.DC1 1521-0103/366/1/220–236$35.00 https://doi.org/10.1124/jpet.118.249292 THE JOURNAL OF PHARMACOLOGY AND EXPERIMENTAL THERAPEUTICS J Pharmacol Exp Ther 366:220–236, July 2018 Copyright ª 2018 by The American Society for Pharmacology and Experimental Therapeutics Analysis of the Indacaterol-Regulated Transcriptome in Human Airway Epithelial Cells Implicates Gene Expression Changes in the s Adverse and Therapeutic Effects of b2-Adrenoceptor Agonists Dong Yan, Omar Hamed, Taruna Joshi,1 Mahmoud M. Mostafa, Kyla C. Jamieson, Radhika Joshi, Robert Newton, and Mark A. Giembycz Departments of Physiology and Pharmacology (D.Y., O.H., T.J., K.C.J., R.J., M.A.G.) and Cell Biology and Anatomy (M.M.M., R.N.), Snyder Institute for Chronic Diseases, Cumming School of Medicine, University of Calgary, Calgary, Alberta, Canada Received March 22, 2018; accepted April 11, 2018 Downloaded from ABSTRACT The contribution of gene expression changes to the adverse and activity, and positive regulation of neutrophil chemotaxis. The therapeutic effects of b2-adrenoceptor agonists in asthma was general enriched GO term extracellular space was also associ- investigated using human airway epithelial cells as a therapeu- ated with indacaterol-induced genes, and many of those, in- tically relevant target. Operational model-fitting established that cluding CRISPLD2, DMBT1, GAS1, and SOCS3, have putative jpet.aspetjournals.org the long-acting b2-adrenoceptor agonists (LABA) indacaterol, anti-inflammatory, antibacterial, and/or antiviral activity. Numer- salmeterol, formoterol, and picumeterol were full agonists on ous indacaterol-regulated genes were also induced or repressed BEAS-2B cells transfected with a cAMP-response element in BEAS-2B cells and human primary bronchial epithelial cells by reporter but differed in efficacy (indacaterol $ formoterol . -

Single Cell Regulatory Landscape of the Mouse Kidney Highlights Cellular Differentiation Programs and Disease Targets
ARTICLE https://doi.org/10.1038/s41467-021-22266-1 OPEN Single cell regulatory landscape of the mouse kidney highlights cellular differentiation programs and disease targets Zhen Miao 1,2,3,8, Michael S. Balzer 1,2,8, Ziyuan Ma 1,2,8, Hongbo Liu1,2, Junnan Wu 1,2, Rojesh Shrestha 1,2, Tamas Aranyi1,2, Amy Kwan4, Ayano Kondo 4, Marco Pontoglio 5, Junhyong Kim6, ✉ Mingyao Li 7, Klaus H. Kaestner2,4 & Katalin Susztak 1,2,4 1234567890():,; Determining the epigenetic program that generates unique cell types in the kidney is critical for understanding cell-type heterogeneity during tissue homeostasis and injury response. Here, we profile open chromatin and gene expression in developing and adult mouse kidneys at single cell resolution. We show critical reliance of gene expression on distal regulatory elements (enhancers). We reveal key cell type-specific transcription factors and major gene- regulatory circuits for kidney cells. Dynamic chromatin and expression changes during nephron progenitor differentiation demonstrates that podocyte commitment occurs early and is associated with sustained Foxl1 expression. Renal tubule cells follow a more complex differentiation, where Hfn4a is associated with proximal and Tfap2b with distal fate. Mapping single nucleotide variants associated with human kidney disease implicates critical cell types, developmental stages, genes, and regulatory mechanisms. The single cell multi-omics atlas reveals key chromatin remodeling events and gene expression dynamics associated with kidney development. 1 Renal, Electrolyte, and Hypertension Division, Department of Medicine, University of Pennsylvania, Perelman School of Medicine, Philadelphia, PA, USA. 2 Institute for Diabetes, Obesity, and Metabolism, University of Pennsylvania, Perelman School of Medicine, Philadelphia, PA, USA. -

Foxc1 Foxc2 Col2a Sox9
bioRxiv preprint doi: https://doi.org/10.1101/2021.01.27.428508; this version posted January 27, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Loss of Foxc1 and Foxc2 function in chondroprogenitor cells disrupts endochondral ossification. 1 2 2 1 1 3 Asra Almubarak , Rotem Lavy , Nikola Srnic , Yawen Hu , Devi P. Maripuri , Tsutomo Kume , Fred B Berry1,2 1 Department of Medical Genetics, University of Alberta, Edmonton AB Canada 2 Department of Surgery, University of Alberta, Edmonton AB, Canada 3 Feinberg Cardiovascular and Renal Research Institute, Feinberg School of Medicine,Department of Medicine, Northwestern University, Chicago, Illinois Foxc1 and Foxc2 function in chondrocytes to regulate endochondral ossification. Key words: Forkhead box, transcription factors, chondrocytes, bone, Gene regulation 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.01.27.428508; this version posted January 27, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Endochondral ossification forms and grows the majority of the mammalian skeleton and is tightly controlled through gene regulatory networks. The forkhead box transcription factors Foxc1 and Foxc2 have been demonstrated to regulate aspects of osteoblast function in the formation of the skeleton but their roles in chondrocytes to control endochondral ossification are less clear. We demonstrate that Foxc1 expression is directly regulated by SOX9 activity, one of the earliest transcription factors to specify the chondrocyte lineages. Moreover we demonstrate that elevelated expression of Foxc1 promotes chondrocyte differentiation in mouse embryonic stem cells and loss of Foxc1 function inhibits chondrogenesis in vitro. -

Expression Microarray Analysis of Renal Development and Human Renal
UNIVERSITY OF CINCINNATI Date:___________________ I, _________________________________________________________, hereby submit this work as part of the requirements for the degree of: in: It is entitled: This work and its defense approved by: Chair: _______________________________ _______________________________ _______________________________ _______________________________ _______________________________ Expression Microarray Analysis of Renal Development and Human Renal Disease A dissertation submitted to the Division of Graduate Studies and Research of the University of Cincinnati in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate Program in Molecular and Developmental Biology of the College of Medicine 2006 by Kristopher Robert Schwab B.A., Blackburn College, 2001 Committee Chair: S. Steven Potter, Ph.D. Tom Doetschman, Ph.D. Chia-Yi Kuan, M.D., Ph.D. Dan Wiginton, Ph.D. James Wells, Ph.D. Abstract Renal morphogenesis involves the reciprocal inductive interactions between the ureteric bud and metanephric mesenchyme forming the collecting ducts and nephrons within adult kidney. We applied microarray technology to the study of renal morphogenesis in order to better understand the molecular mechanisms underlying development. Additionally, the techniques employed in the expression analysis of the embryonic kidney were extended to the study of renal disease. Embryonic kidneys representing different stages of renal development were analyzed using expression microarrays. Renal developmental -

FOXC2 Augments Tumor Propagation and Metastasis in Osteosarcoma
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 7, No. 42 Research Paper FOXC2 augments tumor propagation and metastasis in osteosarcoma Maricel C. Gozo1,2,*, Dongyu Jia1,*, Paul-Joseph Aspuria1, Dong-Joo Cheon1, Naoyuki Miura3, Ann E. Walts4, Beth Y. Karlan1,5, Sandra Orsulic1,5 1Women’s Cancer Program at the Samuel Oschin Comprehensive Cancer Institute, Cedars-Sinai Medical Center, Los Angeles, CA, USA 2Graduate Program in Biomedical Science and Translational Medicine, Cedars-Sinai Medical Center, Los Angeles, CA, USA 3Department of Biochemistry, Hamamatsu University School of Medicine, Hamamatsu, Japan 4Department of Pathology and Laboratory Medicine, Cedars-Sinai Medical Center, Los Angeles, CA, USA 5Department of Obstetrics and Gynecology, David Geffen School of Medicine, University of California, Los Angeles, Los Angeles, CA, USA *These authors have contributed equally to this work Correspondence to: Sandra Orsulic, email: [email protected] Keywords: anoikis, forkhead box C2, osteosarcoma, metastasis, invasion Received: January 19, 2016 Accepted: September 02, 2016 Published: September 13, 2016 ABSTRACT Osteosarcoma is a highly malignant tumor that contains a small subpopulation of tumor-propagating cells (also known as tumor-initiating cells) characterized by drug resistance and high metastatic potential. The molecular mechanism by which tumor- propagating cells promote tumor growth is poorly understood. Here, we report that the transcription factor forkhead box C2 (FOXC2) is frequently expressed in human osteosarcomas and is important in maintaining osteosarcoma cells in a stem-like state. In osteosarcoma cell lines, we show that anoikis conditions stimulate FOXC2 expression. Downregulation of FOXC2 decreases anchorage-independent growth and invasion in vitro and lung metastasis in vivo, while overexpression of FOXC2 increases tumor propagation in vivo. -

The Impact of Transcription Factor Prospero Homeobox 1 on the Regulation of Thyroid Cancer Malignancy
International Journal of Molecular Sciences Review The Impact of Transcription Factor Prospero Homeobox 1 on the Regulation of Thyroid Cancer Malignancy Magdalena Rudzi ´nska 1,2 and Barbara Czarnocka 1,* 1 Department of Biochemistry and Molecular Biology, Centre of Postgraduate Medical Education, 01-813 Warsaw, Poland; [email protected] 2 Institute of Molecular Medicine, Sechenov First Moscow State Medical University, 119991 Moscow, Russia * Correspondence: [email protected]; Tel.: +48-225693812; Fax: +48-225693712 Received: 7 April 2020; Accepted: 30 April 2020; Published: 2 May 2020 Abstract: Transcription factor Prospero homeobox 1 (PROX1) is continuously expressed in the lymphatic endothelial cells, playing an essential role in their differentiation. Many reports have shown that PROX1 is implicated in cancer development and acts as an oncoprotein or suppressor in a tissue-dependent manner. Additionally, the PROX1 expression in many types of tumors has prognostic significance and is associated with patient outcomes. In our previous experimental studies, we showed that PROX1 is present in the thyroid cancer (THC) cells of different origins and has a high impact on follicular thyroid cancer (FTC) phenotypes, regulating migration, invasion, focal adhesion, cytoskeleton reorganization, and angiogenesis. Herein, we discuss the PROX1 transcript and protein structures, the expression pattern of PROX1 in THC specimens, and its epigenetic regulation. Next, we emphasize the biological processes and genes regulated by PROX1 in CGTH-W-1 cells, derived from squamous cell carcinoma of the thyroid gland. Finally, we discuss the interaction of PROX1 with other lymphatic factors. In our review, we aimed to highlight the importance of vascular molecules in cancer development and provide an update on the functionality of PROX1 in THC biology regulation. -

Rattlesnake Genome Supplemental Materials 1 SUPPLEMENTAL
Rattlesnake Genome Supplemental Materials 1 1 SUPPLEMENTAL MATERIALS 2 Table of Contents 3 1. Supplementary Methods …… 2 4 2. Supplemental Tables ……….. 23 5 3. Supplemental Figures ………. 37 Rattlesnake Genome Supplemental Materials 2 6 1. SUPPLEMENTARY METHODS 7 Prairie Rattlesnake Genome Sequencing and Assembly 8 A male Prairie Rattlesnake (Crotalus viridis viridis) collected from a wild population in Colorado was 9 used to generate the genome sequence. This specimen was collected and humanely euthanized according 10 to University of Northern Colorado Institutional Animal Care and Use Committee protocols 0901C-SM- 11 MLChick-12 and 1302D-SM-S-16. Colorado Parks and Wildlife scientific collecting license 12HP974 12 issued to S.P. Mackessy authorized collection of the animal. Genomic DNA was extracted using a 13 standard Phenol-Chloroform-Isoamyl alcohol extraction from liver tissue that was snap frozen in liquid 14 nitrogen. Multiple short-read sequencing libraries were prepared and sequenced on various platforms, 15 including 50bp single-end and 150bp paired-end reads on an Illumina GAII, 100bp paired-end reads on an 16 Illumina HiSeq, and 300bp paired-end reads on an Illumina MiSeq. Long insert libraries were also 17 constructed by and sequenced on the PacBio platform. Finally, we constructed two sets of mate-pair 18 libraries using an Illumina Nextera Mate Pair kit, with insert sizes of 3-5 kb and 6-8 kb, respectively. 19 These were sequenced on two Illumina HiSeq lanes with 150bp paired-end sequencing reads. Short and 20 long read data were used to assemble the previous genome assembly version CroVir2.0 (NCBI accession 21 SAMN07738522). -

Maintenance of Mammary Epithelial Phenotype by Transcription Factor Runx1 Through Mitotic Gene Bookmarking Joshua Rose University of Vermont
University of Vermont ScholarWorks @ UVM Graduate College Dissertations and Theses Dissertations and Theses 2019 Maintenance Of Mammary Epithelial Phenotype By Transcription Factor Runx1 Through Mitotic Gene Bookmarking Joshua Rose University of Vermont Follow this and additional works at: https://scholarworks.uvm.edu/graddis Part of the Biochemistry Commons, and the Genetics and Genomics Commons Recommended Citation Rose, Joshua, "Maintenance Of Mammary Epithelial Phenotype By Transcription Factor Runx1 Through Mitotic Gene Bookmarking" (2019). Graduate College Dissertations and Theses. 998. https://scholarworks.uvm.edu/graddis/998 This Thesis is brought to you for free and open access by the Dissertations and Theses at ScholarWorks @ UVM. It has been accepted for inclusion in Graduate College Dissertations and Theses by an authorized administrator of ScholarWorks @ UVM. For more information, please contact [email protected]. MAINTENANCE OF MAMMARY EPITHELIAL PHENOTYPE BY TRANSCRIPTION FACTOR RUNX1 THROUGH MITOTIC GENE BOOKMARKING A Thesis Presented by Joshua Rose to The Faculty of the Graduate College of The University of Vermont In Partial Fulfillment of the Requirements for the Degree of Master of Science Specializing in Cellular, Molecular, and Biomedical Sciences January, 2019 Defense Date: November 12, 2018 Thesis Examination Committee: Sayyed Kaleem Zaidi, Ph.D., Advisor Gary Stein, Ph.D., Advisor Seth Frietze, Ph.D., Chairperson Janet Stein, Ph.D. Jonathan Gordon, Ph.D. Cynthia J. Forehand, Ph.D. Dean of the Graduate College ABSTRACT Breast cancer arises from a series of acquired mutations that disrupt normal mammary epithelial homeostasis and create multi-potent cancer stem cells that can differentiate into clinically distinct breast cancer subtypes. Despite improved therapies and advances in early detection, breast cancer remains the leading diagnosed cancer in women. -
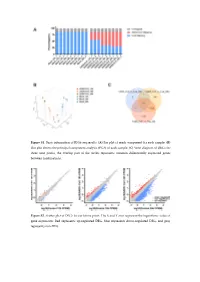
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05