Introduction to Bioinformatics for Computer Scientists
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Bioinformatics 1
Bioinformatics 1 Bioinformatics School School of Science, Engineering and Technology (http://www.stmarytx.edu/set/) School Dean Ian P. Martines, Ph.D. ([email protected]) Department Biological Science (https://www.stmarytx.edu/academics/set/undergraduate/biological-sciences/) Bioinformatics is an interdisciplinary and growing field in science for solving biological, biomedical and biochemical problems with the help of computer science, mathematics and information technology. Bioinformaticians are in high demand not only in research, but also in academia because few people have the education and skills to fill available positions. The Bioinformatics program at St. Mary’s University prepares students for graduate school, medical school or entry into the field. Bioinformatics is highly applicable to all branches of life sciences and also to fields like personalized medicine and pharmacogenomics — the study of how genes affect a person’s response to drugs. The Bachelor of Science in Bioinformatics offers three tracks that students can choose. • Bachelor of Science in Bioinformatics with a minor in Biology: 120 credit hours • Bachelor of Science in Bioinformatics with a minor in Computer Science: 120 credit hours • Bachelor of Science in Bioinformatics with a minor in Applied Mathematics: 120 credit hours Students will take 23 credit hours of core Bioinformatics classes, which included three credit hours of internship or research and three credit hours of a Bioinformatics Capstone course. BS Bioinformatics Tracks • Bachelor of Science -

13 Genomics and Bioinformatics
Enderle / Introduction to Biomedical Engineering 2nd ed. Final Proof 5.2.2005 11:58am page 799 13 GENOMICS AND BIOINFORMATICS Spencer Muse, PhD Chapter Contents 13.1 Introduction 13.1.1 The Central Dogma: DNA to RNA to Protein 13.2 Core Laboratory Technologies 13.2.1 Gene Sequencing 13.2.2 Whole Genome Sequencing 13.2.3 Gene Expression 13.2.4 Polymorphisms 13.3 Core Bioinformatics Technologies 13.3.1 Genomics Databases 13.3.2 Sequence Alignment 13.3.3 Database Searching 13.3.4 Hidden Markov Models 13.3.5 Gene Prediction 13.3.6 Functional Annotation 13.3.7 Identifying Differentially Expressed Genes 13.3.8 Clustering Genes with Shared Expression Patterns 13.4 Conclusion Exercises Suggested Reading At the conclusion of this chapter, the reader will be able to: & Discuss the basic principles of molecular biology regarding genome science. & Describe the major types of data involved in genome projects, including technologies for collecting them. 799 Enderle / Introduction to Biomedical Engineering 2nd ed. Final Proof 5.2.2005 11:58am page 800 800 CHAPTER 13 GENOMICS AND BIOINFORMATICS & Describe practical applications and uses of genomic data. & Understand the major topics in the field of bioinformatics and DNA sequence analysis. & Use key bioinformatics databases and web resources. 13.1 INTRODUCTION In April 2003, sequencing of all three billion nucleotides in the human genome was declared complete. This landmark of modern science brought with it high hopes for the understanding and treatment of human genetic disorders. There is plenty of evidence to suggest that the hopes will become reality—1631 human genetic diseases are now associated with known DNA sequences, compared to the less than 100 that were known at the initiation of the Human Genome Project (HGP) in 1990. -

FUNDAMENTALS of COMPUTING (2019-20) COURSE CODE: 5023 502800CH (Grade 7 for ½ High School Credit) 502900CH (Grade 8 for ½ High School Credit)
EXPLORING COMPUTER SCIENCE NEW NAME: FUNDAMENTALS OF COMPUTING (2019-20) COURSE CODE: 5023 502800CH (grade 7 for ½ high school credit) 502900CH (grade 8 for ½ high school credit) COURSE DESCRIPTION: Fundamentals of Computing is designed to introduce students to the field of computer science through an exploration of engaging and accessible topics. Through creativity and innovation, students will use critical thinking and problem solving skills to implement projects that are relevant to students’ lives. They will create a variety of computing artifacts while collaborating in teams. Students will gain a fundamental understanding of the history and operation of computers, programming, and web design. Students will also be introduced to computing careers and will examine societal and ethical issues of computing. OBJECTIVE: Given the necessary equipment, software, supplies, and facilities, the student will be able to successfully complete the following core standards for courses that grant one unit of credit. RECOMMENDED GRADE LEVELS: 9-12 (Preference 9-10) COURSE CREDIT: 1 unit (120 hours) COMPUTER REQUIREMENTS: One computer per student with Internet access RESOURCES: See attached Resource List A. SAFETY Effective professionals know the academic subject matter, including safety as required for proficiency within their area. They will use this knowledge as needed in their role. The following accountability criteria are considered essential for students in any program of study. 1. Review school safety policies and procedures. 2. Review classroom safety rules and procedures. 3. Review safety procedures for using equipment in the classroom. 4. Identify major causes of work-related accidents in office environments. 5. Demonstrate safety skills in an office/work environment. -

Use of Bioinformatics Resources and Tools by Users of Bioinformatics Centers in India Meera Yadav University of Delhi, [email protected]
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Library Philosophy and Practice (e-journal) Libraries at University of Nebraska-Lincoln 2015 Use of Bioinformatics Resources and Tools by Users of Bioinformatics Centers in India meera yadav University of Delhi, [email protected] Manlunching Tawmbing Saha Institute of Nuclear Physisc, [email protected] Follow this and additional works at: http://digitalcommons.unl.edu/libphilprac Part of the Library and Information Science Commons yadav, meera and Tawmbing, Manlunching, "Use of Bioinformatics Resources and Tools by Users of Bioinformatics Centers in India" (2015). Library Philosophy and Practice (e-journal). 1254. http://digitalcommons.unl.edu/libphilprac/1254 Use of Bioinformatics Resources and Tools by Users of Bioinformatics Centers in India Dr Meera, Manlunching Department of Library and Information Science, University of Delhi, India [email protected], [email protected] Abstract Information plays a vital role in Bioinformatics to achieve the existing Bioinformatics information technologies. Librarians have to identify the information needs, uses and problems faced to meet the needs and requirements of the Bioinformatics users. The paper analyses the response of 315 Bioinformatics users of 15 Bioinformatics centers in India. The papers analyze the data with respect to use of different Bioinformatics databases and tools used by scholars and scientists, areas of major research in Bioinformatics, Major research project, thrust areas of research and use of different resources by the user. The study identifies the various Bioinformatics services and resources used by the Bioinformatics researchers. Keywords: Informaion services, Users, Inforamtion needs, Bioinformatics resources 1. Introduction ‘Needs’ refer to lack of self-sufficiency and also represent gaps in the present knowledge of the users. -

Challenges in Bioinformatics
Yuri Quintana, PhD, delivered a webinar in AllerGen’s Webinars for Research Success series on February 27, 2018, discussing different approaches to biomedical informatics and innovations in big-data platforms for biomedical research. His main messages and a hyperlinked index to his presentation follow. WHAT IS BIOINFORMATICS? Bioinformatics is an interdisciplinary field that develops analytical methods and software tools for understanding clinical and biological data. It combines elements from many fields, including basic sciences, biology, computer science, mathematics and engineering, among others. WHY DO WE NEED BIOINFORMATICS? Chronic diseases are rapidly expanding all over the world, and associated healthcare costs are increasing at an astronomical rate. We need to develop personalized treatments tailored to the genetics of increasingly diverse patient populations, to clinical and family histories, and to environmental factors. This requires collecting vast amounts of data, integrating it, and making it accessible and usable. CHALLENGES IN BIOINFORMATICS Data collection, coordination and archiving: These technologies have evolved to the point Many sources of biomedical data—hospitals, that we can now analyze not only at the DNA research centres and universities—do not have level, but down to the level of proteins. DNA complete data for any particular disease, due in sequencers, DNA microarrays, and mass part to patient numbers, but also to the difficulties spectrometers are generating tremendous of data collection, inter-operability -
Florida Course Code Directory Computer Science Course Information 2019-2020
FLORIDA COURSE CODE DIRECTORY COMPUTER SCIENCE COURSE INFORMATION 2019-2020 Section 1007.2616, Florida Statutes, was amended by the Florida Legislature to include the definition of computer science and a requirement that computer science courses be identified in the Course Code Directory published on the Department of Education’s website. DEFINITION: The study of computers and algorithmic processes, including their principles, hardware and software designs, applications, and their impact on society, and includes computer coding and computer programming. SECONDARY COMPUTER SCIENCE COURSES Middle and high schools in each district, including combination schools in which any of grades 6-12 are taught, must provide an opportunity for students to enroll in a computer science course. If a school district does not offer an identified computer science course the district must provide students access to the course through the Florida Virtual School (FLVS) or through other means. COURSE NUMBER COURSE TITLE 0200000 M/J Computer Science Discoveries 0200010 M/J Computer Science Discoveries 1 0200020 M/J Computer Science Discoveries 2 0200305 Computer Science Discoveries 0200315 Computer Science Principles 0200320 AP Computer Science A 0200325 AP Computer Science A Innovation 0200335 AP Computer Science Principles 0200435 PRE-AICE Computer Studies IGCSE Level 0200480 AICE Computer Science 1 AS Level 0200485 AICE Computer Science 2 A Level 0200490 AICE Information Technology 1 AS Level 0200495 AICE Information Technology 2 A Level 0200800 IB Computer -

Safety and Security Challenge
SAFETY AND SECURITY CHALLENGE TOP SUPERCOMPUTERS IN THE WORLD - FEATURING TWO of DOE’S!! Summary: The U.S. Department of Energy (DOE) plays a very special role in In fields where scientists deal with issues from disaster relief to the keeping you safe. DOE has two supercomputers in the top ten supercomputers in electric grid, simulations provide real-time situational awareness to the whole world. Titan is the name of the supercomputer at the Oak Ridge inform decisions. DOE supercomputers have helped the Federal National Laboratory (ORNL) in Oak Ridge, Tennessee. Sequoia is the name of Bureau of Investigation find criminals, and the Department of the supercomputer at Lawrence Livermore National Laboratory (LLNL) in Defense assess terrorist threats. Currently, ORNL is building a Livermore, California. How do supercomputers keep us safe and what makes computing infrastructure to help the Centers for Medicare and them in the Top Ten in the world? Medicaid Services combat fraud. An important focus lab-wide is managing the tsunamis of data generated by supercomputers and facilities like ORNL’s Spallation Neutron Source. In terms of national security, ORNL plays an important role in national and global security due to its expertise in advanced materials, nuclear science, supercomputing and other scientific specialties. Discovery and innovation in these areas are essential for protecting US citizens and advancing national and global security priorities. Titan Supercomputer at Oak Ridge National Laboratory Background: ORNL is using computing to tackle national challenges such as safe nuclear energy systems and running simulations for lower costs for vehicle Lawrence Livermore's Sequoia ranked No. -

Comparing Bioinformatics Software Development by Computer Scientists and Biologists: an Exploratory Study
Comparing Bioinformatics Software Development by Computer Scientists and Biologists: An Exploratory Study Parmit K. Chilana Carole L. Palmer Amy J. Ko The Information School Graduate School of Library The Information School DUB Group and Information Science DUB Group University of Washington University of Illinois at University of Washington [email protected] Urbana-Champaign [email protected] [email protected] Abstract Although research in bioinformatics has soared in the last decade, much of the focus has been on high- We present the results of a study designed to better performance computing, such as optimizing algorithms understand information-seeking activities in and large-scale data storage techniques. Only recently bioinformatics software development by computer have studies on end-user programming [5, 8] and scientists and biologists. We collected data through semi- information activities in bioinformatics [1, 6] started to structured interviews with eight participants from four emerge. Despite these efforts, there still is a gap in our different bioinformatics labs in North America. The understanding of problems in bioinformatics software research focus within these labs ranged from development, how domain knowledge among MBB computational biology to applied molecular biology and and CS experts is exchanged, and how the software biochemistry. The findings indicate that colleagues play a development process in this domain can be improved. significant role in information seeking activities, but there In our exploratory study, we used an information is need for better methods of capturing and retaining use perspective to begin understanding issues in information from them during development. Also, in bioinformatics software development. We conducted terms of online information sources, there is need for in-depth interviews with 8 participants working in 4 more centralization, improved access and organization of bioinformatics labs in North America. -

Database Technology for Bioinformatics from Information Retrieval to Knowledge Systems
Database Technology for Bioinformatics From Information Retrieval to Knowledge Systems Luis M. Rocha Complex Systems Modeling CCS3 - Modeling, Algorithms, and Informatics Los Alamos National Laboratory, MS B256 Los Alamos, NM 87545 [email protected] or [email protected] 1 Molecular Biology Databases 3 Bibliographic databases On-line journals and bibliographic citations – MEDLINE (1971, www.nlm.nih.gov) 3 Factual databases Repositories of Experimental data associated with published articles and that can be used for computerized analysis – Nucleic acid sequences: GenBank (1982, www.ncbi.nlm.nih.gov), EMBL (1982, www.ebi.ac.uk), DDBJ (1984, www.ddbj.nig.ac.jp) – Amino acid sequences: PIR (1968, www-nbrf.georgetown.edu), PRF (1979, www.prf.op.jp), SWISS-PROT (1986, www.expasy.ch) – 3D molecular structure: PDB (1971, www.rcsb.org), CSD (1965, www.ccdc.cam.ac.uk) Lack standardization of data contents 3 Knowledge Bases Intended for automatic inference rather than simple retrieval – Motif libraries: PROSITE (1988, www.expasy.ch/sprot/prosite.html) – Molecular Classifications: SCOP (1994, www.mrc-lmb.cam.ac.uk) – Biochemical Pathways: KEGG (1995, www.genome.ad.jp/kegg) Difference between knowledge and data (semiosis and syntax)?? 2 Growth of sequence and 3D Structure databases Number of Entries 3 Database Technology and Bioinformatics 3 Databases Computerized collection of data for Information Retrieval Shared by many users Stored records are organized with a predefined set of data items (attributes) Managed by a computer program: the database -
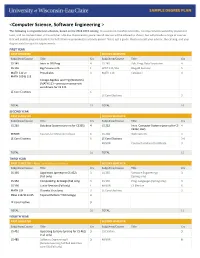
<Computer Science, Software Engineering >
<Computer Science, Software Engineering > The following is a hypothetical schedule, based on the 2018-2019 catalog. It assumes no transferred credits, no requirements waived by placement tests, and no courses taken in the summer. UW-Eau Claire cannot guarantee all courses will be offered as shown, but will provide a range of courses that will enable prepared students to fulfill their requirements in a timely period. This is just a guide. Please consult your advisor, the catalog, and your degree audit for specific requirements. FIRST YEAR FIRST SEMESTER SECOND SEMESTER Subj/Area/Course Title Crs Subj/Area/Course Title Crs CS 145 Intro to OO Prog 4 CS 245 Adv. Prog. Data Structures 4 CS 146 Big Picture in CS 1 WRIT 114/116 Blugold Seminar 5 MATH 112 or Precalculus 4 MATH 114 Calculus I 4 MATH 109 & 113 College Algebra and Trig (Winterim) (MATH 113 – prereq or concurrent enrollment for CS 245 LE Core Electives 6 LE Core Electives 3 TOTAL 15 TOTAL 16 SECOND YEAR FIRST SEMESTER SECOND SEMESTER Subj/Area/Course Title Crs Subj/Area/Course Title Crs CS 260 Database Systems (prereq for CS355) 4 CS 252 Intro. Computer Systems (prereq for CS 4 CS352, 452) MINOR Courses for Minor/Certificate 6 CS 268 Web Systems 3 LE Core Electives 6 LE Core Electives 3-6 MINOR Courses for Minor/Certificate 3 TOTAL 16 TOTAL 15 THIRD YEAR FIRST SEMESTER – Apply for Admission to Major SECOND SEMESTER Subj/Area/Course Title Crs Subj/Area/Course Title Crs CS 335 Algorithms (prereq for CS 452) 3 CS 355 Software Engineering I 3 (Fall only) (Spring only) CS 352 ComputeOrg. -
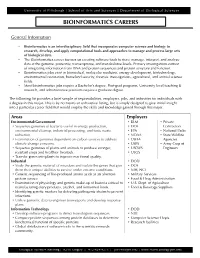
Bioinformatics Careers
University of Pittsburgh | School of Arts and Sciences | Department of Biological Sciences BIOINFORMATICS CAREERS General Information • Bioinformatics is an interdisciplinary field that incorporates computer science and biology to research, develop, and apply computational tools and approaches to manage and process large sets of biological data. • The Bioinformatics career focuses on creating software tools to store, manage, interpret, and analyze data at the genome, proteome, transcriptome, and metabalome levels. Primary investigations consist of integrating information from DNA and protein sequences and protein structure and function. • Bioinformatics jobs exist in biomedical, molecular medicine, energy development, biotechnology, environmental restoration, homeland security, forensic investigations, agricultural, and animal science fields. • Most bioinformatics jobs require a Bachelor’s degree. Post-grad programs, University level teaching & research, and administrative positions require a graduate degree. The following list provides a brief sample of responsibilities, employers, jobs, and industries for individuals with a degree in this major. This is by no means an exhaustive listing, but is simply designed to give initial insight into a particular career field that would employ the skills and knowledge gained through this major. Areas Employers Environmental/Government • BLM • Private • Sequence genomes of bacteria useful in energy production, • DOE Contractors environmental cleanup, industrial processing, and toxic waste • EPA • National Parks reduction. • NOAA • State Wildlife • Examination of genomes dependent on carbon sources to address • USDA Agencies climate change concerns. • USFS • Army Corp of • Sequence genomes of plants and animals to produce stronger, • USFWS Engineers resistant crops and healthier livestock. • USGS • Transfer genes into plants to improve nutritional quality. Industrial • DOD • Study the genetic material of microbes and isolate the genes that give • DOE them their unique abilities to survive under extreme conditions. -

Artificial Intelligence Program 1
Artificial Intelligence Program 1 Artificial Intelligence Program Reid Simmons, Director of the BSAI program (NSH 3213) 15-122 Principles of Imperative Computation 10 (students without credit or a waiver for 15-112, Jean Harpley, Program Coordinator (NSH 1517) Fundamentals of Programming and Computer www.cs.cmu.edu/bs-in-artificial-intelligence (http://www.cs.cmu.edu/bs-in- Science, must take 15-112 before 15-122) artificial-intelligence/) 15-150 Principles of Functional Programming 10 15-210 Parallel and Sequential Data Structures and 12 Overview Algorithms 15-213 Introduction to Computer Systems 12 Carnegie Mellon University has led the world in artificial intelligence 15-251 Great Ideas in Theoretical Computer Science 12 education and innovation since the field was created. It's only natural, then, that the School of Computer Science would offer the nation's first bachelor's degree in Artificial Intelligence, which started in Fall 2018. Artificial Intelligence The new BSAI program gives students the in-depth knowledge needed to transform large amounts of data into actionable decisions. The program All of the following three AI core courses: Units and its curriculum focus on how complex inputs — such as vision, language 07-180 Concepts in Artificial Intelligence 5 and huge databases — can be used to make decisions or enhance human 15-281 Artificial Intelligence: Representation and 12 capabilities. The curriculum includes coursework in computer science, Problem Solving math, statistics, computational modeling, machine learning and symbolic computation. Because Carnegie Mellon is devoted to AI for social good, 10-315 Introduction to Machine Learning (SCS Majors) 12 students will also take courses in ethics and social responsibility, with the plus one of the following AI core courses: option to participate in independent study projects that change the world for 16-385 Computer Vision 12 the better — in areas like healthcare, transportation and education.