ATF4 Gene Network Mediates Cellular Response to the Anticancer PAD
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genome-Wide Analysis of 5-Hmc in the Peripheral Blood of Systemic Lupus Erythematosus Patients Using an Hmedip-Chip
INTERNATIONAL JOURNAL OF MOLECULAR MEDICINE 35: 1467-1479, 2015 Genome-wide analysis of 5-hmC in the peripheral blood of systemic lupus erythematosus patients using an hMeDIP-chip WEIGUO SUI1*, QIUPEI TAN1*, MING YANG1, QIANG YAN1, HUA LIN1, MINGLIN OU1, WEN XUE1, JIEJING CHEN1, TONGXIANG ZOU1, HUANYUN JING1, LI GUO1, CUIHUI CAO1, YUFENG SUN1, ZHENZHEN CUI1 and YONG DAI2 1Guangxi Key Laboratory of Metabolic Diseases Research, Central Laboratory of Guilin 181st Hospital, Guilin, Guangxi 541002; 2Clinical Medical Research Center, the Second Clinical Medical College of Jinan University (Shenzhen People's Hospital), Shenzhen, Guangdong 518020, P.R. China Received July 9, 2014; Accepted February 27, 2015 DOI: 10.3892/ijmm.2015.2149 Abstract. Systemic lupus erythematosus (SLE) is a chronic, Introduction potentially fatal systemic autoimmune disease characterized by the production of autoantibodies against a wide range Systemic lupus erythematosus (SLE) is a typical systemic auto- of self-antigens. To investigate the role of the 5-hmC DNA immune disease, involving diffuse connective tissues (1) and modification with regard to the onset of SLE, we compared is characterized by immune inflammation. SLE has a complex the levels 5-hmC between SLE patients and normal controls. pathogenesis (2), involving genetic, immunologic and envi- Whole blood was obtained from patients, and genomic DNA ronmental factors. Thus, it may result in damage to multiple was extracted. Using the hMeDIP-chip analysis and valida- tissues and organs, especially the kidneys (3). SLE arises from tion by quantitative RT-PCR (RT-qPCR), we identified the a combination of heritable and environmental influences. differentially hydroxymethylated regions that are associated Epigenetics, the study of changes in gene expression with SLE. -

Supplementary Table S1. Upregulated Genes Differentially
Supplementary Table S1. Upregulated genes differentially expressed in athletes (p < 0.05 and 1.3-fold change) Gene Symbol p Value Fold Change 221051_s_at NMRK2 0.01 2.38 236518_at CCDC183 0.00 2.05 218804_at ANO1 0.00 2.05 234675_x_at 0.01 2.02 207076_s_at ASS1 0.00 1.85 209135_at ASPH 0.02 1.81 228434_at BTNL9 0.03 1.81 229985_at BTNL9 0.01 1.79 215795_at MYH7B 0.01 1.78 217979_at TSPAN13 0.01 1.77 230992_at BTNL9 0.01 1.75 226884_at LRRN1 0.03 1.74 220039_s_at CDKAL1 0.01 1.73 236520_at 0.02 1.72 219895_at TMEM255A 0.04 1.72 201030_x_at LDHB 0.00 1.69 233824_at 0.00 1.69 232257_s_at 0.05 1.67 236359_at SCN4B 0.04 1.64 242868_at 0.00 1.63 1557286_at 0.01 1.63 202780_at OXCT1 0.01 1.63 1556542_a_at 0.04 1.63 209992_at PFKFB2 0.04 1.63 205247_at NOTCH4 0.01 1.62 1554182_at TRIM73///TRIM74 0.00 1.61 232892_at MIR1-1HG 0.02 1.61 204726_at CDH13 0.01 1.6 1561167_at 0.01 1.6 1565821_at 0.01 1.6 210169_at SEC14L5 0.01 1.6 236963_at 0.02 1.6 1552880_at SEC16B 0.02 1.6 235228_at CCDC85A 0.02 1.6 1568623_a_at SLC35E4 0.00 1.59 204844_at ENPEP 0.00 1.59 1552256_a_at SCARB1 0.02 1.59 1557283_a_at ZNF519 0.02 1.59 1557293_at LINC00969 0.03 1.59 231644_at 0.01 1.58 228115_at GAREM1 0.01 1.58 223687_s_at LY6K 0.02 1.58 231779_at IRAK2 0.03 1.58 243332_at LOC105379610 0.04 1.58 232118_at 0.01 1.57 203423_at RBP1 0.02 1.57 AMY1A///AMY1B///AMY1C///AMY2A///AMY2B// 208498_s_at 0.03 1.57 /AMYP1 237154_at LOC101930114 0.00 1.56 1559691_at 0.01 1.56 243481_at RHOJ 0.03 1.56 238834_at MYLK3 0.01 1.55 213438_at NFASC 0.02 1.55 242290_at TACC1 0.04 1.55 ANKRD20A1///ANKRD20A12P///ANKRD20A2/// -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

The Gas6 Gene Rs8191974 and Ap3s2 Gene Rs2028299 Are Associated with Type 2 Diabetes in the Northern Chinese Han Population Elena V
Vol. 64, No 2/2017 227–231 https://doi.org/10.18388/abp.2016_1299 Regular paper The Gas6 gene rs8191974 and Ap3s2 gene rs2028299 are associated with type 2 diabetes in the northern Chinese Han population Elena V. Kazakova#, Tianwei Zghuang#, Tingting Li, Qingxiao Fang, Jun Han and Hong Qiao* 1The Fifth Endocrine Department, the Second Affiliated Hospital, Harbin Medical University, Harbin, Heilongjiang, China Previous studies in other countries have shown that INTRODUCTION single nucleotide polymorphisms (SNPs) in the growth arrest-specific gene 6 (Gas6; rs8191974) and adapt- Diabetes mellitus affects more than 300 million indi- er-related protein complex 3 subunit sigma-2 (Ap3s2; viduals worldwide, with increasing prevalence particular- rs2028299) were associated with an increasedrisk for ly in the developing countries (Whiting et al., 2011). In type 2 diabetes mellitus (T2DM). However, the associ- fact, the prevalence of type 2 diabetes mellitus (T2DM) ation of these loci with T2DM has not been examined in China is among the highest in the world. The com- in Chinese populations. We performed a replication bination of insulin resistance in peripheral tissues and study to investigate the association of these suscep- impaired insulin secretion from pancreatic β-cells is be- tibility loci with T2DM in the Chinese population. lieved to contribute to the development and progression We genotyped 1968 Chinese participants (996 with of T2DM. Both, the genetic and environmental factors T2DM and 972controls) for rs8191974 in Gas6 and confer susceptibility to T2DM. In recent years, studies rs2028299 near Ap3s2, and examined their associa- of gene polymorphisms have helped identify a number tion with T2DM using a logistic regression analysis. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -
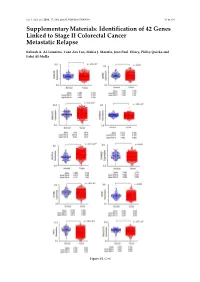
Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S1 of S16 Supplementary Materials: Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse Rabeah A. Al-Temaimi, Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke and Fahd Al-Mulla Figure S1. Cont. Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S2 of S16 Figure S1. Mean expression levels of fourteen genes of significant association with CRC DFS and OS that are differentially expressed in normal colon compared to CRC tissues. Each dot represents a sample. Table S1. Copy number aberrations associated with poor disease-free survival and metastasis in early stage II CRC as predicted by STAC and SPPS combined methodologies with resident gene symbols. CN stands for copy number, whereas CNV is copy number variation. Region Cytoband % of CNV Count of Region Event Gene Symbols Length Location Overlap Genes chr1:113,025,076–113,199,133 174,057 p13.2 CN Loss 0.0 2 AKR7A2P1, SLC16A1 chr1:141,465,960–141,822,265 356,305 q12–q21.1 CN Gain 95.9 1 SRGAP2B MIR5087, LOC10013000 0, FLJ39739, LOC10028679 3, PPIAL4G, PPIAL4A, NBPF14, chr1:144,911,564–146,242,907 1,331,343 q21.1 CN Gain 99.6 16 NBPF15, NBPF16, PPIAL4E, NBPF16, PPIAL4D, PPIAL4F, LOC645166, LOC388692, FCGR1C chr1:177,209,428–177,226,812 17,384 q25.3 CN Gain 0.0 0 chr1:197,652,888–197,676,831 23,943 q32.1 CN Gain 0.0 1 KIF21B chr1:201,015,278–201,033,308 18,030 q32.1 CN Gain 0.0 1 PLEKHA6 chr1:201,289,154–201,298,247 9093 q32.1 CN Gain 0.0 0 chr1:216,820,186–217,043,421 223,235 q41 CN -

Integrative Cross Tissue Analysis of Gene Expression Identifies Novel
bioRxiv preprint doi: https://doi.org/10.1101/108134; this version posted February 27, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 1 Integrative cross tissue analysis of gene expression identifies 2 novel type 2 diabetes genes 1 2 2 3 2 3 Jason M. Torres , Alvaro N. Barbeira , Rodrigo Bonazzola , Andrew P. Morris , Kaanan P. Shah , 4 5,6 7, 2, 4 Heather E. Wheeler , Graeme I. Bell , Nancy J. Cox ⇤, Hae Kyung Im ⇤ 5 1 Committee on Molecular Metabolism and Nutrition, Biological Sciences Division, The University of 6 Chicago, Chicago, IL, USA 7 2 Section of Genetic Medicine, Department of Medicine, The University of Chicago, Chicago, IL, USA 8 3 Institute of Translational Medicine, University of Liverpool, Liverpool, United Kingdom 9 4 Departments of Biology and Computer Science, Loyola University Chicago, Chicago, IL, USA 10 5 Department of Medicine, The University of Chicago, Chicago, IL, USA 11 6 Department of Human Genetics, The University of Chicago, Chicago, IL, USA 12 7 Division of Genetic Medicine, Vanderbilt University, Nashville, TN, USA 13 * Correspondence to: [email protected] and [email protected] 14 Abstract 15 To understand the mechanistic underpinnings of type 2 diabetes (T2D) loci mapped through GWAS, we 16 performed a tissue-specific gene association study in a cohort of over 100K individuals (n 26K, cases ⇡ 17 n 84K) across 44 human tissues using MetaXcan, a summary statistics extension of PrediXcan. -

Modeling Genomic Diversity and Tumor Dependency in Malignant Melanoma
Research Article Modeling Genomic Diversity and Tumor Dependency in Malignant Melanoma William M. Lin,1,3,5 Alissa C. Baker,1,3 Rameen Beroukhim,1,3,5 Wendy Winckler,1,3,5 Whei Feng,1,3,5 Jennifer M. Marmion,7 Elisabeth Laine,8 Heidi Greulich,1,3,5 Hsiuyi Tseng,1,3 Casey Gates,5 F. Stephen Hodi,1 Glenn Dranoff,1 William R. Sellers,1,6 Roman K. Thomas,9,10 Matthew Meyerson,1,3,4,5 Todd R. Golub,2,3,5 Reinhard Dummer,8 Meenhard Herlyn,7 Gad Getz,3,5 and Levi A. Garraway1,3,5 Departments of 1Medical Oncology and 2Pediatric Oncology and 3Center for Cancer Genome Discovery, Dana-Farber Cancer Institute, Harvard Medical School; 4Department of Pathology, Harvard Medical School, Boston, Massachusetts; 5The Broad Institute of M.I.T. and Harvard; 6Novartis Institutes for Biomedical Research, Cambridge, Massachusetts; 7Cancer Biology Division, Wistar Institute, Philadelphia, Pennsylvania; 8Department of Dermatology, University of Zurich Hospital, Zu¨rich, Switzerland; 9Max Planck Institute for Neurological Research with Klaus Joachim Zulch Laboratories of the Max Planck Society and the Medical Faculty of the University of Cologne; and 10Center for Integrated Oncology and Department I for Internal Medicine, University of Cologne, Cologne, Germany Abstract tumorigenesis have been derived from functional studies involving The classification of human tumors based on molecular cultured human cancer cells (e.g., established cell lines, short-term cultures, etc.). Despite their limitations, cancer cell line collections criteria offers tremendous clinical potential; however, dis- cerning critical and ‘‘druggable’’ effectors on a large scale will whose genetic alterations reflect their primary tumor counterparts also require robust experimental models reflective of tumor should provide malleable proxies that facilitate mechanistic genomic diversity. -

1 Novel Expression Signatures Identified by Transcriptional Analysis
ARD Online First, published on October 7, 2009 as 10.1136/ard.2009.108043 Ann Rheum Dis: first published as 10.1136/ard.2009.108043 on 7 October 2009. Downloaded from Novel expression signatures identified by transcriptional analysis of separated leukocyte subsets in SLE and vasculitis 1Paul A Lyons, 1Eoin F McKinney, 1Tim F Rayner, 1Alexander Hatton, 1Hayley B Woffendin, 1Maria Koukoulaki, 2Thomas C Freeman, 1David RW Jayne, 1Afzal N Chaudhry, and 1Kenneth GC Smith. 1Cambridge Institute for Medical Research and Department of Medicine, Addenbrooke’s Hospital, Hills Road, Cambridge, CB2 0XY, UK 2Roslin Institute, University of Edinburgh, Roslin, Midlothian, EH25 9PS, UK Correspondence should be addressed to Dr Paul Lyons or Prof Kenneth Smith, Department of Medicine, Cambridge Institute for Medical Research, Addenbrooke’s Hospital, Hills Road, Cambridge, CB2 0XY, UK. Telephone: +44 1223 762642, Fax: +44 1223 762640, E-mail: [email protected] or [email protected] Key words: Gene expression, autoimmune disease, SLE, vasculitis Word count: 2,906 The Corresponding Author has the right to grant on behalf of all authors and does grant on behalf of all authors, an exclusive licence (or non-exclusive for government employees) on a worldwide basis to the BMJ Publishing Group Ltd and its Licensees to permit this article (if accepted) to be published in Annals of the Rheumatic Diseases and any other BMJPGL products to exploit all subsidiary rights, as set out in their licence (http://ard.bmj.com/ifora/licence.pdf). http://ard.bmj.com/ on September 29, 2021 by guest. Protected copyright. 1 Copyright Article author (or their employer) 2009. -

Identification of C2CD4A As a Human Diabetes Susceptibility Gene with a Role in Β Cell Insulin Secretion
Identification of C2CD4A as a human diabetes susceptibility gene with a role in β cell insulin secretion Taiyi Kuoa, Michael J. Kraakmana, Manashree Damleb,c, Richard Gilla, Mitchell A. Lazarb,c,1, and Domenico Accilia,1 aDepartment of Medicine, Berrie Diabetes Center, Columbia University College of Physicians and Surgeons, New York, NY 10032; bThe Institute for Diabetes, Obesity, and Metabolism, University of Pennsylvania Perelman School of Medicine, Philadelphia, PA 19104; and cDivision of Endocrinology, Diabetes, and Metabolism, Department of Medicine, University of Pennsylvania Perelman School of Medicine, Philadelphia, PA 19104 Contributed by Mitchell A. Lazar, July 31, 2019 (sent for review March 14, 2019; reviewed by Alvin C. Powers and Andrew F. Stewart) Fine mapping and validation of genes causing β cell failure from targets. To circumvent this obstacle, we generated FoxO1- susceptibility loci identified in type 2 diabetes genome-wide asso- GFPVenus (Venus) reporter knockin mice, and utilized 2-photon ciation studies (GWAS) poses a significant challenge. The VPS13C- microscopy to track its subcellular localization in pancreatic β C2CD4A-C2CD4B locus on chromosome 15 confers diabetes suscep- cells. We next performed genome-wide FoxO1 chromatin immu- tibility in every ethnic group studied to date. However, the causative noprecipitation sequencing (ChIP-seq) to identify its genomic gene is unknown. FoxO1 is involved in the pathogenesis of β cell targets as well as superenhancers encompassing FoxO1 sites. A dysfunction, but its link to human diabetes GWAS has not been comparative analysis of human islet and murine β cell super- explored. Here we generated a genome-wide map of FoxO1 super- enhancers revealed C2CD4A, a gene encoding an IL-1β–induced β enhancers in chemically identified cells using 2-photon live-cell nuclear protein (18) embedded among several SNPs conferring imaging to monitor FoxO1 localization. -

As a Model for Lysosomal Storage Disorders Gert De Voer, Dorien Peters, Peter E.M
as a model for lysosomal storage disorders Gert de Voer, Dorien Peters, Peter E.M. Taschner To cite this version: Gert de Voer, Dorien Peters, Peter E.M. Taschner. as a model for lysosomal storage disorders. Biochimica et Biophysica Acta - Molecular Basis of Disease, Elsevier, 2008, 1782 (7-8), pp.433. 10.1016/j.bbadis.2008.04.003. hal-00501575 HAL Id: hal-00501575 https://hal.archives-ouvertes.fr/hal-00501575 Submitted on 12 Jul 2010 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. ÔØ ÅÒÙ×Ö ÔØ Caenorhabditis elegans as a model for lysosomal storage disorders Gert de Voer, Dorien Peters, Peter E.M. Taschner PII: S0925-4439(08)00093-8 DOI: doi: 10.1016/j.bbadis.2008.04.003 Reference: BBADIS 62810 To appear in: BBA - Molecular Basis of Disease Received date: 13 May 2007 Revised date: 23 April 2008 Accepted date: 24 April 2008 Please cite this article as: Gert de Voer, Dorien Peters, Peter E.M. Taschner, Caenorhab- ditis elegans as a model for lysosomal storage disorders, BBA - Molecular Basis of Disease (2008), doi: 10.1016/j.bbadis.2008.04.003 This is a PDF file of an unedited manuscript that has been accepted for publication. -

NIH Public Access Author Manuscript Annu Rev Genomics Hum Genet
NIH Public Access Author Manuscript Annu Rev Genomics Hum Genet. Author manuscript; available in PMC 2009 October 1. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Annu Rev Genomics Hum Genet. 2008 ; 9: 359±386. doi:10.1146/annurev.genom.9.081307.164303. Disorders of Lysosome-related Organelle Biogenesis: Clinical and Molecular Genetics Marjan Huizing1, Amanda Helip-Wooley2, Wendy Westbroek2, Meral Gunay-Aygun2, and William A. Gahl2 Marjan Huizing: [email protected]; Amanda Helip-Wooley: [email protected]; Wendy Westbroek: [email protected]; Meral Gunay-Aygun: [email protected]; William A. Gahl: [email protected] 1 Cell Biology of Metabolic Disorders Unit, National Institutes of Health, Bethesda, Maryland 20892 2 Section on Human Biochemical Genetics, Medical Genetics Branch, National Human Genome Research Institute, National Institutes of Health, Bethesda, Maryland 20892 Abstract Lysosome-related organelles (LROs) are a heterogeneous group of vesicles that share various features with lysosomes, but are distinct in function, morphology, and composition. The biogenesis of LROs employs a common machinery, and genetic defects in this machinery can affect all LROs or only an individual LRO, resulting in a variety of clinical features. In this review, we discuss the main components in LRO biogenesis. We also address the function, composition and resident cell type of the major LROs. Finally, we describe the clinical characteristics of the major human LRO disorders. Keywords Chediak-Higashi syndrome; Griscelli syndrome; Hermansky-Pudlak syndrome; melanosome; platelet INTRODUCTION Lysosomes are membrane-bound cytoplasmic organelles that serve as major degradative compartments in eukaryotic cells (65).