TRIBE Editing Reveals Specific Mrna Targets of Eif4e-BP in Drosophila and in Mammals
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

4-6 Weeks Old Female C57BL/6 Mice Obtained from Jackson Labs Were Used for Cell Isolation
Methods Mice: 4-6 weeks old female C57BL/6 mice obtained from Jackson labs were used for cell isolation. Female Foxp3-IRES-GFP reporter mice (1), backcrossed to B6/C57 background for 10 generations, were used for the isolation of naïve CD4 and naïve CD8 cells for the RNAseq experiments. The mice were housed in pathogen-free animal facility in the La Jolla Institute for Allergy and Immunology and were used according to protocols approved by the Institutional Animal Care and use Committee. Preparation of cells: Subsets of thymocytes were isolated by cell sorting as previously described (2), after cell surface staining using CD4 (GK1.5), CD8 (53-6.7), CD3ε (145- 2C11), CD24 (M1/69) (all from Biolegend). DP cells: CD4+CD8 int/hi; CD4 SP cells: CD4CD3 hi, CD24 int/lo; CD8 SP cells: CD8 int/hi CD4 CD3 hi, CD24 int/lo (Fig S2). Peripheral subsets were isolated after pooling spleen and lymph nodes. T cells were enriched by negative isolation using Dynabeads (Dynabeads untouched mouse T cells, 11413D, Invitrogen). After surface staining for CD4 (GK1.5), CD8 (53-6.7), CD62L (MEL-14), CD25 (PC61) and CD44 (IM7), naïve CD4+CD62L hiCD25-CD44lo and naïve CD8+CD62L hiCD25-CD44lo were obtained by sorting (BD FACS Aria). Additionally, for the RNAseq experiments, CD4 and CD8 naïve cells were isolated by sorting T cells from the Foxp3- IRES-GFP mice: CD4+CD62LhiCD25–CD44lo GFP(FOXP3)– and CD8+CD62LhiCD25– CD44lo GFP(FOXP3)– (antibodies were from Biolegend). In some cases, naïve CD4 cells were cultured in vitro under Th1 or Th2 polarizing conditions (3, 4). -

Eukaryotic Translation Initiation Factor 3 Plays Distinct Roles at The
RESEARCH ARTICLE Eukaryotic translation initiation factor 3 plays distinct roles at the mRNA entry and exit channels of the ribosomal preinitiation complex Colin Echeverrı´aAitken1, Petra Beznoskova´ 2, Vladislava Vlcˇkova2, Wen-Ling Chiu3†, Fujun Zhou1, Leosˇ Shivaya Vala´ sˇek2*, Alan G Hinnebusch3*, Jon R Lorsch1* 1Laboratory on the Mechanism and Regulation of Protein Synthesis, Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institutes of Health, Bethesda, United States; 2Laboratory of Regulation of Gene Expression, Institute of Microbiology ASCR, Prague, Czech Republic; 3Laboratory of Gene Regulation and Development, Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institutes of Health, Bethesda, United States Abstract Eukaryotic translation initiation factor 3 (eIF3) is a central player in recruitment of the pre-initiation complex (PIC) to mRNA. We probed the effects on mRNA recruitment of a library of S. cerevisiae eIF3 functional variants spanning its 5 essential subunits using an in vitro-reconstituted *For correspondence: valasekl@ system. Mutations throughout eIF3 disrupt its interaction with the PIC and diminish its ability to biomed.cas.cz (LSV); [email protected] (AGH); jon. accelerate recruitment to a native yeast mRNA. Alterations to the eIF3a CTD and eIF3b/i/g . [email protected] (JRL) significantly slow mRNA recruitment, and mutations within eIF3b/i/g destabilize eIF2 GTP Met- tRNAi binding to the PIC. Using model mRNAs lacking contacts with the 40S entry or exit channels, Present address: we uncovered a critical role for eIF3 requiring the eIF3a NTD, in stabilizing mRNA interactions at †PharmaEssentia Corp., Taipei, the exit channel, and an ancillary role at the entry channel requiring residues of the eIF3a CTD. -

Characterization of the Small RNA Transcriptomes of Cell Protrusions and Cell Bodies of Highly Metastatic Hepatocellular Carcinoma Cells Via RNA Sequencing
ONCOLOGY LETTERS 22: 568, 2021 Characterization of the small RNA transcriptomes of cell protrusions and cell bodies of highly metastatic hepatocellular carcinoma cells via RNA sequencing WENPIN CAI1*, JINGZHANG JI2*, BITING WU2*, KAIXUAN HAO2, PING REN2, YU JIN2, LIHONG YANG2, QINGCHAO TONG2 and ZHIFA SHEN2 1Department of Laboratory Medicine, Wen Zhou Traditional Chinese Medicine Hospital; 2Zhejiang Provincial Key Laboratory of Medical Genetics, Key Laboratory of Laboratory Medicine, Ministry of Education, School of Laboratory Medicine and Life Sciences, Wenzhou Medical University, Wenzhou, Zhejiang 325035, P.R. China Received July 11, 2020; Accepted February 23, 2021 DOI: 10.3892/ol.2021.12829 Abstract. Increasing evidence suggest that hepatocellular differentially expressed miRNAs and circRNAs. The interac‑ carcinoma (HCC) HCCLM3 cells initially develop pseudo‑ tion maps between miRNAs and circRNAs were constructed, podia when they metastasize, and microRNAs (miRNAs/miRs) and signaling pathway maps were analyzed to determine the and circular RNAs (circRNAs) have been demonstrated to molecular mechanism and regulation of the differentially serve important roles in the development, progression and expressed miRNAs and circRNAs. Taken together, the results metastasis of cancer. The present study aimed to isolate the of the present study suggest that the Boyden chamber assay cell bodies (CBs) and cell protrusions (CPs) from HCCLM3 can be used to effectively isolate the somatic CBs and CPs of cells, and screen the miRNAs and circRNAs associated with HCC, which can be used to screen the miRNAs and circRNAs HCC infiltration and metastasis in CBs and CPs. The Boyden associated with invasion and metastasis of HCC. chamber assay has been confirmed to effectively isolate the CBs and CPs from HCCLM3 cells via observation of microtu‑ Introduction bule immunofluorescence, DAPI staining and nuclear protein H3 western blotting. -

Systematically Profiling the Expression of Eif3 Subunits in Glioma Reveals
Chai et al. Cancer Cell Int (2019) 19:155 https://doi.org/10.1186/s12935-019-0867-1 Cancer Cell International PRIMARY RESEARCH Open Access Systematically profling the expression of eIF3 subunits in glioma reveals the expression of eIF3i has prognostic value in IDH-mutant lower grade glioma Rui‑Chao Chai1,4,6†, Ning Wang2†, Yu‑Zhou Chang3, Ke‑Nan Zhang1,6, Jing‑Jun Li1,6, Jun‑Jie Niu5, Fan Wu1,6*, Yu‑Qing Liu1,6* and Yong‑Zhi Wang1,3,4,6* Abstract Background: Abnormal expression of the eukaryotic initiation factor 3 (eIF3) subunits plays critical roles in tumo‑ rigenesis and progression, and also has potential prognostic value in cancers. However, the expression and clinical implications of eIF3 subunits in glioma remain unknown. Methods: Expression data of eIF3 for patients with gliomas were obtained from the Chinese Glioma Genome Atlas (CGGA) (n 272) and The Cancer Genome Atlas (TCGA) (n 595). Cox regression, the receiver operating characteristic (ROC) curves= and Kaplan–Meier analysis were used to study= the prognostic value. Gene oncology (GO) and gene set enrichment analysis (GSEA) were utilized for functional prediction. Results: In both the CGGA and TCGA datasets, the expression levels of eIF3d, eIF3e, eIF3f, eIF3h and eIF3l highly were associated with the IDH mutant status of gliomas. The expression of eIF3b, eIF3i, eIF3k and eIF3m was increased with the tumor grade, and was associated with poorer overall survival [All Hazard ratio (HR) > 1 and P < 0.05]. By contrast, the expression of eIF3a and eIF3l was decreased in higher grade gliomas and was associated with better overall sur‑ vival (Both HR < 1 and P < 0.05). -

The J-Subunit of Human Translation Initiation Factor Eif3 Is Required for the Stable Binding of Eif3 and Its Subcomplexes to 40 S Ribosomal Subunits in Vitro*
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 279, No. 10, Issue of March 5, pp. 8946–8956, 2004 © 2004 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. The j-Subunit of Human Translation Initiation Factor eIF3 Is Required for the Stable Binding of eIF3 and Its Subcomplexes to 40 S Ribosomal Subunits in Vitro* Received for publication, November 21, 2003, and in revised form, December 18, 2003 Published, JBC Papers in Press, December 19, 2003, DOI 10.1074/jbc.M312745200 Christopher S. Fraser‡, Jennifer Y. Lee, Greg L. Mayeur, Martin Bushell§, Jennifer A. Doudna¶, and John W. B. Hersheyʈ From the Department of Biological Chemistry, School of Medicine, University of California, Davis, California 95616 Eukaryotic initiation factor 3 (eIF3) is a 12-subunit protein subunits, named in order of decreasing molecular protein complex that plays a central role in binding of weight as recommended (4): eIF3a, eIF3b, eIF3c, eIF3d, eIF3l, initiator methionyl-tRNA and mRNA to the 40 S riboso- eIF3e, eIF3f, eIF3g, eIF3h, eIF3i, eIF3j, and eIF3k (5, 6). Spe- mal subunit to form the 40 S initiation complex. The cific functions for mammalian eIF3 have been identified by a molecular mechanisms by which eIF3 exerts these func- variety of in vitro experiments. It binds directly to 40 S ribo- tions are poorly understood. To learn more about the somal subunits in the absence of other initiation components structure and function of eIF3 we have expressed and (1), and affects the association/dissociation of ribosomes (7–10). purified individual human eIF3 subunits or complexes It promotes the binding of Met-tRNA and mRNA to the 40 S of eIF3 subunits using baculovirus-infected Sf9 cells. -

Relevance of Translation Initiation in Diffuse Glioma Biology and Its
cells Review Relevance of Translation Initiation in Diffuse Glioma Biology and its Therapeutic Potential Digregorio Marina 1, Lombard Arnaud 1,2, Lumapat Paul Noel 1, Scholtes Felix 1,2, Rogister Bernard 1,3 and Coppieters Natacha 1,* 1 Laboratory of Nervous System Disorders and Therapy, GIGA-Neurosciences Research Centre, University of Liège, 4000 Liège, Belgium; [email protected] (D.M.); [email protected] (L.A.); [email protected] (L.P.N.); [email protected] (S.F.); [email protected] (R.B.) 2 Department of Neurosurgery, CHU of Liège, 4000 Liège, Belgium 3 Department of Neurology, CHU of Liège, 4000 Liège, Belgium * Correspondence: [email protected] Received: 18 October 2019; Accepted: 26 November 2019; Published: 29 November 2019 Abstract: Cancer cells are continually exposed to environmental stressors forcing them to adapt their protein production to survive. The translational machinery can be recruited by malignant cells to synthesize proteins required to promote their survival, even in times of high physiological and pathological stress. This phenomenon has been described in several cancers including in gliomas. Abnormal regulation of translation has encouraged the development of new therapeutics targeting the protein synthesis pathway. This approach could be meaningful for glioma given the fact that the median survival following diagnosis of the highest grade of glioma remains short despite current therapy. The identification of new targets for the development of novel therapeutics is therefore needed in order to improve this devastating overall survival rate. This review discusses current literature on translation in gliomas with a focus on the initiation step covering both the cap-dependent and cap-independent modes of initiation. -
Lncrna Gclnc1 Promotes Gastric Carcinogenesis and May Act As a Modular Scaffold of WDR5 and KAT2A Complexes to Specify the Histone Modifi Cation Pattern
Published OnlineFirst May 4, 2016; DOI: 10.1158/2159-8290.CD-15-0921 RESEARCH ARTICLE LncRNA GClnc1 Promotes Gastric Carcinogenesis and May Act as a Modular Scaffold of WDR5 and KAT2A Complexes to Specify the Histone Modifi cation Pattern Tian-Tian Sun 1 , Jie He 2 , Qian Liang 1 , Lin-Lin Ren 1 , Ting-Ting Yan 1 , Ta-Chung Yu 1 , Jia-Yin Tang 1 , Yu-Jie Bao 1 , Ye H u 1 , Yanwei Lin 1 , 3 , Danfeng Sun 1 , 3 , Ying-Xuan Chen 1 , Jie Hong 1 , Haoyan Chen 1 , Weiping Zou 3 , and Jing-Yuan Fang 1 Downloaded from cancerdiscovery.aacrjournals.org on September 27, 2021. © 2016 American Association for Cancer Research. Published OnlineFirst May 4, 2016; DOI: 10.1158/2159-8290.CD-15-0921 ABSTRACT Long noncoding RNAs (lncRNA) play a role in carcinogenesis. However, the func- tion of lncRNAs in human gastric cancer remains largely unknown. In this study, we identifi ed a novel lncRNA, GClnc1, which was upregulated and associated with tumorigenesis, tumor size, metastasis, and poor prognosis in gastric cancer. GClnc1 affected gastric cancer cell proliferation, invasiveness, and metastasis in multiple gastric cancer models. Mechanistically, GClnc1 bound WDR5 (a key component of histone methyltransferase complex) and KAT2A histone acetyltransferase, acted as a modular scaffold of WDR5 and KAT2A complexes, coordinated their localization, specifi ed the histone modifi cation pattern on the target genes, including SOD2 , and consequently altered gastric cancer cell biology. Thus, GClnc1 is mechanistically, functionally, and clinically oncogenic in gastric cancer. Targeting GClnc1 and its pathway may be meaningful for treating patients with gastric cancer. -

Duke University Dissertation Template
Regulation of Eukaryotic Translation Initiation by Signal Transduction by Mayya Shveygert Department of Molecular Genetics and Microbiology Duke University Date:_______________________ Approved: ___________________________ Matthias Gromeier, Supervisor ___________________________ Christopher Nicchitta ___________________________ Jeffrey Rathmell ___________________________ Micah Luftig Dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Molecular Genetics and Microbiology in the Graduate School of Duke University 2012 i v ABSTRACT Regulation of Eukaryotic Translation Initiation by Signal Transduction by Mayya Shveygert Department of Molecular Genetics and Microbiology Duke University Date:_______________________ Approved: ___________________________ Matthias Gromeier, Supervisor ___________________________ Christopher Nicchitta ___________________________ Jeffrey Rathmell ___________________________ Micah Luftig An abstract of a dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Molecular Genetics and Microbiology in the Graduate School of Duke University 2012 Copyright by Mayya Shveygert 2012 Abstract Eukaryotic translation initiation is a rate-limiting step of protein synthesis and is controlled by signal transduction in response to various extracellular cues and stresses. This thesis is focused on the eukaryotic initiation factor 4G (eIF4G), which participates in multiple steps of initiation: (i) eIF4G interaction with polyA-binding protein (PABP) links the 5’ pre-initiation complex to the 3’ poly(A) tail of mRNAs; (ii) eIF4G recruits 40S ribosomal subunit to mRNAs through interactions with the m7-G cap binding protein eIF4E; (iii) eIF4G binds to the eIF4E kinase, mitogen activated protein kinase (MAPK) interacting kinase 1 (Mnk1), modulating eIF4E phosphorylation. I studied how eIF4G function is affected by viral infection, mitogenic stimulation and during mitosis. -

EIF3G Is Associated with Narcolepsy Across Ethnicities
European Journal of Human Genetics (2015) 23, 1573–1580 & 2015 Macmillan Publishers Limited All rights reserved 1018-4813/15 www.nature.com/ejhg ARTICLE EIF3G is associated with narcolepsy across ethnicities Anja Holm1,2, Ling Lin3, Juliette Faraco3, Sara Mostafavi4, Alexis Battle4, Xiaowei Zhu5, Douglas F Levinson5, Fang Han6, Steen Gammeltoft1, Poul Jennum2, Emmanuel Mignot3 and Birgitte R Kornum*,1 Type 1 narcolepsy, an autoimmune disease affecting hypocretin (orexin) neurons, is strongly associated with HLA-DQB1*06:02. Among polymorphisms associated with the disease is single-nucleotide polymorphism rs2305795 (c.*638G4A) located within the P2RY11 gene. P2RY11 is in a region of synteny conserved in mammals and zebrafish containing PPAN, EIF3G and DNMT1 (DNA methyltransferase 1). As mutations in DNMT1 cause a rare dominant form of narcolepsy in association with deafness, cerebellar ataxia and dementia, we questioned whether the association with P2RY11 in sporadic narcolepsy could be secondary to linkage disequilibrium with DNMT1. Based on genome-wide association data from two cohorts of European and Chinese ancestry, we found that the narcolepsy association signal drops sharply between P2RY11/EIF3G and DNMT1, suggesting that the association with narcolepsy does not extend into the DNMT1 gene region. Interestingly, using transethnic mapping, we identified a novel single-nucleotide polymorphism rs3826784 (c.596–260A4G) in the EIF3G gene also associated with narcolepsy. The disease-associated allele increases EIF3G mRNA expression. EIF3G is located in the narcolepsy risk locus and EIF3G expression correlates with PPAN and P2RY11 expression. This suggests shared regulatory mechanisms that might be affected by the polymorphism and are of relevance to narcolepsy. -
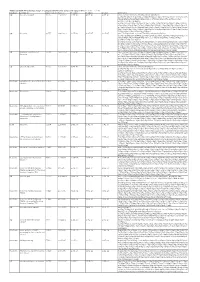
Additional Tables.Xlsx
Additional Table 5 Enriched pathways of upregulated DEGs after spinal cord injury in WT (P < 0.05, q < 0.05) Database Description Ratio of DEGs Ratio of P -value P adjust q value gene name GO ribosomal subunit 87/755 214/12262 1.03E-49 4.61E-47 4.20E-47 Gm6576/Gm10036/Gm5786/Mrps28/Gm9493/Rpl13- ps3/Rpl7l1/Mrpl10/Mrpl44/Rps20/Mrpl51/Mrps11/Mpv17l2/Mrps7/Gm10020/Gm10073 /Mrpl2/Mrpl52/Rpl22/Mrpl4/Mrps24/Zcchc17/Rpl28/Mrpl34/Rpl29/Mrps36/Rpl9- ps6/Hba-a2/Hba-a1/Rpl23a- ps3/Rpl15/Rack1/Mrps18a/Mrpl28/Rpl10a/Mrpl12/Rpl18/Rpl35a/Mrpl41/Mrps12/Rpl36 a/Rps9/Rps18/Rps5/Rps27a/Rps24/Rplp2/Rpl14/Mrpl30/Rps6/Rpl6/Rps3/Rps4x/Rpsa/R pl7/Rpl27a/Rps23/Rps10/Mrps21/Rpl36/Rpl30/Rps17/Mrpl42/Rpl35/Rpl27/Rplp0/Rps28 /Rpl31/Rpl39/Rps13/Rpl13/Rps11/Rpl8/Rps3a1/Fau/Rpl9/Rpl3/Rpl26/Rps15/Rpl13a/Uba 52/Rpl24/Rpl19/Rps14/Rpl4/Rps27/Rplp1 GO ribosome 93/755 248/12262 1.67E-49 4.61E-47 4.20E-47 Gm6576/Pnpt1/Gm10036/Gm5786/Mrps28/Gm9493/Rpl13- ps3/Rpl7l1/Mrpl10/Mrpl44/Rps20/Mrpl51/Mrps11/Mpv17l2/Mrps7/Gm10020/Gm10073 /Mrpl2/Mrpl52/Rpl22/Mrpl4/Mrps24/Zcchc17/Rpl28/Mrpl34/Rpl29/Mrps36/Rpl9- ps6/Hba-a2/Hba-a1/Rpl23a- ps3/Rpl15/Rack1/Eif3h/Mrps18a/Mrpl28/Mrps33/Rpl10a/Mrpl12/Rpl18/Rpl35a/Btf3/Mrp l41/Mrps12/Rpl36a/Rps9/Rps18/Rps5/Rps27a/Rps24/Rplp2/Rpl14/Mrpl30/Rps6/Rpl6/R ps3/Rps4x/Rpsa/Rpl7/Rpl27a/Rps23/Rps10/Mrps21/Rpl36/Rpl30/Rps17/Mrpl42/Rpl35/R pl27/Rplp0/Rps28/Rpl31/Rpl39/Rps13/Rpl13/Rps11/Rpl8/Rps3a1/Fau/Rpl9/Rpl3/Rpl26/ Rps15/Rpl13a/Uba52/Rpl41/Rpl24/Rpl19/Rps14/Rpl4/Ndufa7/Rps27/Rplp1 GO structural constituent of 76/733 169/11958 1.65E-47 1.16E-44 1.12E-44 Gm6576/Gm10036/Gm5786/Rpl7l1/Mrpl10/Rps20/Mrpl51/Mrps11/Mrps7/Gm10020/G -

Original Article Overexpression of Eif3e Is Correlated with Colon Tumor Development and Poor Prognosis
Int J Clin Exp Pathol 2014;7(10):6462-6474 www.ijcep.com /ISSN:1936-2625/IJCEP0002023 Original Article Overexpression of eIF3e is correlated with colon tumor development and poor prognosis Zhi Li1*, Shengtao Lin2*, Tao Jiang3, Jingtao Wang2, Huijun Lu4, Huamei Tang4, Mujian Teng5, Junwei Fan2 1Department of Anorectal Surgery, Qianfoshan Hospital Affiliated to Shandong University, 16766 Jingshi Road, Jinan 250014, Shandong, China; 2Department of General Surgery, Shanghai Jiaotong University Affiliated First People’s Hospital, 85 Wujin Road, Shanghai 200080, China; 3Department of Anal-Colorectal Surgery, General Hospital of Ningxia Medical University, 804 South Shengli Road, Yinchuan 750004, China; 4Department of Pathology, Shanghai Jiaotong University Affiliated First People’s Hospital, 85 Wujin Road, Shanghai 200080, China; 5Department of General Surgery, Qianfoshan Hospital Affiliated to Shandong University, 16766 Jingshi Road, Jinan 250014, Shandong, China. *Equal contributors. Received August 21, 2014; Accepted September 15, 2014; Epub September 15, 2014; Published October 1, 2014 Abstract: EIF3e is a component of the eukaryotic translation initiation factor 3 (eIF-3) complexes, which is an es- sential factor for initiation of protein synthesis in mammalian cells. Translational control plays key roles in the complex mechanism of cancer development and progression. However, the clinical significance of eIF3e in colon cancer remains to be elucidated. We analyzed the eIF3e expression in a tissue microarray (TMA), which contained 173 colon cancer tissues paired with adjacent normal mucosa and lymph node metastasis. The expression of eIF3e was significantly elevated in colon cancer tissues in comparison with those in adjacent normal mucosa (P < 0.001) and lymph node metastasis (P < 0.001).