2-S2.0-84958164785.Pdf (413.0Kb)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Vampire Fish Rides Giant Catfishes in The
FREE MEALS ON LONG-DISTANCE CRUISERS: THE VAMPIRE FISH RIDES GIANT CATFISHES IN THE AMAZON Jansen Zuanon*, Ivan Sazima** Biota Neotropica v5 (n1) – http://www.biotaneotropica.org.br/v5n1/pt/abstract?article+BN03005012005 Recebido em 16/08/04 - Versão revisada recebida em 21/01/05 - Publicado em 10/02/05. *CPBA, Caixa Postal 478, INPA-Instituto Nacional de Pesquisas da Amazônia,69083-970 Manaus, Amazonas, Brasil **Departamento de Zoologia e Museu de História Natural, Caixa Postal 6109, Universidade Estadual de Campinas, 13083-970 Campinas, São Paulo, Brazil (www.unicamp.br) **Corresponding author. Tel: + 55-19-3788 7292; fax: +55-19-3289 3124; e-mail: [email protected] Abstract - The trichomycterid catfishes known as candirus are renowned for their blood feeding, but information on their habits under natural conditions is very fragmentary and generally restricted to hosts or habitats. We recorded an undescribed species of the vandelliine genus Paracanthopoma riding the giant jau catfish, Zungaro zungaro (Pimelodidae), in the upper Amazon. The candirus were found on the host’s caudal and pectoral fins, as well as the base of the dorsal fin, with their snouts buried up to the eyes in the tough skin of the catfish host. All of them had small amounts of partly digested blood in the distal part of the gut. Along the host’s dorsal fin base we found a few additional tiny holes, most of them healed. We suggest that Paracanthopoma feeds on the gill chamber of its hosts, and that the individuals we found were taking a ride partly buried into the host’s skin. -

Water Diversion in Brazil Threatens Biodiversit
See discussions, stats, and author profiles for this publication at: https://www.researchgate.net/publication/332470352 Water diversion in Brazil threatens biodiversity Article in AMBIO A Journal of the Human Environment · April 2019 DOI: 10.1007/s13280-019-01189-8 CITATIONS READS 0 992 12 authors, including: Vanessa Daga Valter Monteiro de Azevedo-Santos Universidade Federal do Paraná 34 PUBLICATIONS 374 CITATIONS 17 PUBLICATIONS 248 CITATIONS SEE PROFILE SEE PROFILE Fernando Pelicice Philip Fearnside Universidade Federal de Tocantins Instituto Nacional de Pesquisas da Amazônia 68 PUBLICATIONS 2,890 CITATIONS 612 PUBLICATIONS 20,906 CITATIONS SEE PROFILE SEE PROFILE Some of the authors of this publication are also working on these related projects: Freshwater microscrustaceans from continental Ecuador and Galápagos Islands: Integrative taxonomy and ecology View project Conservation policy View project All content following this page was uploaded by Philip Fearnside on 11 May 2019. The user has requested enhancement of the downloaded file. The text that follows is a PREPRINT. O texto que segue é um PREPRINT. Please cite as: Favor citar como: Daga, Vanessa S.; Valter M. Azevedo- Santos, Fernando M. Pelicice, Philip M. Fearnside, Gilmar Perbiche-Neves, Lucas R. P. Paschoal, Daniel C. Cavallari, José Erickson, Ana M. C. Ruocco, Igor Oliveira, André A. Padial & Jean R. S. Vitule. 2019. Water diversion in Brazil threatens biodiversity: Potential problems and alternatives. Ambio https://doi.org/10.1007/s13280-019- 01189-8 . (online version published 27 April 2019) ISSN: 0044-7447 (print version) ISSN: 1654-7209 (electronic version) Copyright: Royal Swedish Academy of Sciences & Springer Science+Business Media B.V. -

2017 JMIH Program Book Web Version 6-26-17.Pub
Organizing Societies American Elasmobranch Society 33rd Annual Meeting President: Dean Grubbs Treasurer: Cathy Walsh Secretary: Jennifer Wyffels Editor and Webmaster: David Shiffman Immediate Past President: Chris Lowe American Society of Ichthyologists and Herpetologists 97th Annual Meeting President: Carole Baldwin President Elect: Brian Crother Past President: Maureen A. Donnelly Prior Past President: Larry G. Allen Treasurer: F. Douglas Martin Secretary: Prosanta Chakrabarty Editor: Christopher Beachy Herpetologists’ League 75th Annual Meeting President: David M. Green Immediate Past President: James Spotila Vice-President: David Sever Treasurer: Laurie Mauger Secretary: Renata Platenburg Publications Secretary: Ken Cabarle Communications Secretary: Wendy Palin Herpetologica Editor: Stephen Mullin Herpetological Monographs Editor: Michael Harvey Society for the Study of Amphibians and Reptiles 60th Annual Meeting President: Richard Shine President-Elect: Marty Crump Immediate Past-President: Aaron Bauer Secretary: Marion R. Preest Treasurer: Kim Lovich Publications Secretary: Cari-Ann Hickerson Thank you to our generous sponsor We would like to thank the following: Local Hosts David Hillis, University of Texas at Austin, LHC Chair Dean Hendrickson, University of Texas at Austin Becca Tarvin, University of Texas at Austin Anne Chambers, University of Texas at Austin Christopher Peterson, University of Texas at Austin Volunteers We wish to thank the following volunteers who have helped make the Joint Meeting of Ichthyologists and Herpetologists -

Dedication Donald Perrin De Sylva
Dedication The Proceedings of the First International Symposium on Mangroves as Fish Habitat are dedicated to the memory of University of Miami Professors Samuel C. Snedaker and Donald Perrin de Sylva. Samuel C. Snedaker Donald Perrin de Sylva (1938–2005) (1929–2004) Professor Samuel Curry Snedaker Our longtime collaborator and dear passed away on March 21, 2005 in friend, University of Miami Professor Yakima, Washington, after an eminent Donald P. de Sylva, passed away in career on the faculty of the University Brooksville, Florida on January 28, of Florida and the University of Miami. 2004. Over the course of his diverse A world authority on mangrove eco- and productive career, he worked systems, he authored numerous books closely with mangrove expert and and publications on topics as diverse colleague Professor Samuel Snedaker as tropical ecology, global climate on relationships between mangrove change, and wetlands and fish communities. Don pollutants made major scientific contributions in marine to this area of research close to home organisms in south and sedi- Florida ments. One and as far of his most afield as enduring Southeast contributions Asia. He to marine sci- was the ences was the world’s publication leading authority on one of the most in 1974 of ecologically important inhabitants of “The ecology coastal mangrove habitats—the great of mangroves” (coauthored with Ariel barracuda. His 1963 book Systematics Lugo), a paper that set the high stan- and Life History of the Great Barracuda dard by which contemporary mangrove continues to be an essential reference ecology continues to be measured. for those interested in the taxonomy, Sam’s studies laid the scientific bases biology, and ecology of this species. -

Estudo Das Relações Filogenéticas De Trichomycteridae (Teleostei, Siluriformes) Com Base Em Evidências Cromossômicas E Moleculares
Universidade Estadual Paulista Instituto de Biociências Departamento de Morfologia Luciana Ramos Sato Estudo das relações filogenéticas de Trichomycteridae (Teleostei, Siluriformes) com base em evidências cromossômicas e moleculares. Tese apresentada ao curso de Pós Graduação em Ciências Biológicas, Área de Concentração: Genética, para obtenção do título de Doutor. Orientador: Profo Dr. Claudio de Oliveira Botucatu-SP 2007 Livros Grátis http://www.livrosgratis.com.br Milhares de livros grátis para download. FICHA CATALOGRÁFICA ELABORADA PELA SEÇÃO TÉCNICA DE AQUISIÇÃO E TRATAMENTO DA INFORMAÇÃO DIVISÃO TÉCNICA DE BIBLIOTECA E DOCUMENTAÇÃO - CAMPUS DE BOTUCATU - UNESP BIBLIOTECÁRIA RESPONSÁVEL: SELMA MARIA DE JESUS Sato, Luciana Ramos. Estudo das relações filogenéticas de Trichomycteridae (Teleostei, Silurifomes) com base em evidências cromossômicas e moleculares / Luciana Ramos Sato. – Botucatu : [s.n.], 2007. Tese (doutorado) – Universidade Estadual Paulista, Instituto de Biociências de Botucatu 2007. Orientador: Cláudio de Oliveira Assunto CAPES: 20200008 1. Peixe de água doce - Filogenia 2. Peixe - Genética 3. Biologia molecular CDD 597.15 Palavras-chave: Citogenética; DNA mitocondrial; Filogenia; Siluriformes; Trichomycteridae “A classificação por descendência não pode ser inventada por biólogos, ela pode apenas ser descoberta”. (Theodosius Dobzhansky) Dedico este trabalho, Aos meus pais Sato e Irene e à minha irmã Eliana Agradecimentos Gostaria de agradecer a todos que contribuíram para a realização deste trabalho, e em especial: A Deus por estar sempre presente guiando meus caminhos. Ao professor Cláudio de Oliveira pela orientação, transmissão de conhecimento e acima de tudo pela paciência durante todos os meus anos em Botucatu. Aos professores Dr. Fausto Foresti, Dr. César Martins e Dra Adriane Wasko por todos os ensinamentos. Ao Departamento de Morfologia e seus funcionários, e em especial a D. -

Amazon Alive: a Decade of Discoveries 1999-2009
Amazon Alive! A decade of discovery 1999-2009 The Amazon is the planet’s largest rainforest and river basin. It supports countless thousands of species, as well as 30 million people. © Brent Stirton / Getty Images / WWF-UK © Brent Stirton / Getty Images The Amazon is the largest rainforest on Earth. It’s famed for its unrivalled biological diversity, with wildlife that includes jaguars, river dolphins, manatees, giant otters, capybaras, harpy eagles, anacondas and piranhas. The many unique habitats in this globally significant region conceal a wealth of hidden species, which scientists continue to discover at an incredible rate. Between 1999 and 2009, at least 1,200 new species of plants and vertebrates have been discovered in the Amazon biome (see page 6 for a map showing the extent of the region that this spans). The new species include 637 plants, 257 fish, 216 amphibians, 55 reptiles, 16 birds and 39 mammals. In addition, thousands of new invertebrate species have been uncovered. Owing to the sheer number of the latter, these are not covered in detail by this report. This report has tried to be comprehensive in its listing of new plants and vertebrates described from the Amazon biome in the last decade. But for the largest groups of life on Earth, such as invertebrates, such lists do not exist – so the number of new species presented here is no doubt an underestimate. Cover image: Ranitomeya benedicta, new poison frog species © Evan Twomey amazon alive! i a decade of discovery 1999-2009 1 Ahmed Djoghlaf, Executive Secretary, Foreword Convention on Biological Diversity The vital importance of the Amazon rainforest is very basic work on the natural history of the well known. -

Convergent Evolution of Weakly Electric Fishes from Floodplain Habitats in Africa and South America
Environmental Biology of Fishes 49: 175–186, 1997. 1997 Kluwer Academic Publishers. Printed in the Netherlands. Convergent evolution of weakly electric fishes from floodplain habitats in Africa and South America Kirk O. Winemiller & Alphonse Adite Department of Wildlife and Fisheries Sciences, Texas A&M University, College Station, TX 77843, U.S.A. Received 19.7.1995 Accepted 27.5.1996 Key words: diet, electrogenesis, electroreception, foraging, morphology, niche, Venezuela, Zambia Synopsis An assemblage of seven gymnotiform fishes in Venezuela was compared with an assemblage of six mormyri- form fishes in Zambia to test the assumption of convergent evolution in the two groups of very distantly related, weakly electric, noctournal fishes. Both assemblages occur in strongly seasonal floodplain habitats, but the upper Zambezi floodplain in Zambia covers a much larger area. The two assemblages had broad diet overlap but relatively narrow overlap of morphological attributes associated with feeding. The gymnotiform assemblage had greater morphological variation, but mormyriforms had more dietary variation. There was ample evidence of evolutionary convergence based on both morphology and diet, and this was despite the fact that species pairwise morphological similarity and dietary similarity were uncorrelated in this dataset. For the most part, the two groups have diversified in a convergent fashion within the confines of their broader niche as nocturnal invertebrate feeders. Both assemblages contain midwater planktivores, microphagous vegetation- dwellers, macrophagous benthic foragers, and long-snouted benthic probers. The gymnotiform assemblage has one piscivore, a niche not represented in the upper Zambezi mormyriform assemblage, but present in the form of Mormyrops deliciousus in the lower Zambezi and many other regions of Africa. -

Chirping and Asymmetric Jamming Avoidance Responses in the Electric Fish Distocyclus Conirostris Jacquelyn M
© 2018. Published by The Company of Biologists Ltd | Journal of Experimental Biology (2018) 221, jeb178913. doi:10.1242/jeb.178913 SHORT COMMUNICATION Chirping and asymmetric jamming avoidance responses in the electric fish Distocyclus conirostris Jacquelyn M. Petzold1,2, JoséA. Alves-Gomes3 and G. Troy Smith1,2,* ABSTRACT of two of more EODs creates a periodic amplitude modulation Electrosensory systems of weakly electric fish must accommodate (beat). Beat frequency is equal to the difference between the EOD competing demands of sensing the environment (electrolocation) frequencies (EODfs) of the two interacting fish. Fish use the beat and receiving social information (electrocommunication). The and the relative geometry of the interacting signals to estimate jamming avoidance response (JAR) is a behavioral strategy thought conspecific EODfs, which convey important social information to reduce electrosensory interference from conspecific signals close (Smith, 2013; Dunlap, et al., 2017). However, slow beats (<10 Hz) in frequency. We used playback experiments to characterize electric created by interactions between fish with similar EODfs can impair organ discharge frequency (EODf), chirping behavior and the JAR of the electrolocation function of the EOD by masking localized EOD Distocyclus conirostris, a gregarious electric fish species. EODs of D. distortions (Heiligenberg, 1973; Matsubara and Heiligenberg, conirostris had low frequencies (∼80–200 Hz) that shifted in response 1978). The JAR is a stereotyped response in which an electric to playback stimuli. Fish consistently lowered EODf in response to fish increases or decreases its EODf to increase beat frequency and higher-frequency stimuli but inconsistently raised or lowered EODf in thereby reduce or eliminate the interference caused by slow beats response to lower-frequency stimuli. -

Information Sheet on Ramsar Wetlands (RIS) – 2009-2012 Version Available for Download From
Information Sheet on Ramsar Wetlands (RIS) – 2009-2012 version Available for download from http://www.ramsar.org/ris/key_ris_index.htm. Categories approved by Recommendation 4.7 (1990), as amended by Resolution VIII.13 of the 8th Conference of the Contracting Parties (2002) and Resolutions IX.1 Annex B, IX.6, IX.21 and IX. 22 of the 9th Conference of the Contracting Parties (2005). Notes for compilers: 1. The RIS should be completed in accordance with the attached Explanatory Notes and Guidelines for completing the Information Sheet on Ramsar Wetlands. Compilers are strongly advised to read this guidance before filling in the RIS. 2. Further information and guidance in support of Ramsar site designations are provided in the Strategic Framework and guidelines for the future development of the List of Wetlands of International Importance (Ramsar Wise Use Handbook 14, 3rd edition). A 4th edition of the Handbook is in preparation and will be available in 2009. 3. Once completed, the RIS (and accompanying map(s)) should be submitted to the Ramsar Secretariat. Compilers should provide an electronic (MS Word) copy of the RIS and, where possible, digital copies of all maps. 1. Name and address of the compiler of this form: FOR OFFICE USE ONLY. DD MM YY Beatriz de Aquino Ribeiro - Bióloga - Analista Ambiental / [email protected], (95) Designation date Site Reference Number 99136-0940. Antonio Lisboa - Geógrafo - MSc. Biogeografia - Analista Ambiental / [email protected], (95) 99137-1192. Instituto Chico Mendes de Conservação da Biodiversidade - ICMBio Rua Alfredo Cruz, 283, Centro, Boa Vista -RR. CEP: 69.301-140 2. -

Curriculum Vitae 21 May 2012 Christopher B
Curriculum Vitae 21 May 2012 Christopher B. Braun Department of Psychology Hunter College Biopsychology Program City University of New York 695 Park Avenue New York, NY 10021 Phone: (212) 772-5554; Fax: (212) 772-5620 [email protected] www.urban.hunter.cuny.edu/~cbraun Education: Postdoctoral (Neuroethology) Parmly Hearing Institute, Loyola University Chicago 2001 Ph.D. (Neurosciences) University of California at San Diego 1997 M.S. (Neurosciences) University of California at San Diego 1993 B.A. (Evolutionary Biology) Hampshire College. 1991 Positions Held: 2008- Research Associate, Vertebrate Zoology, American Museum of Natural History. 2007- Associate Professor, Department of Psychology, Hunter College, and Biopsychology and Ecology Evolution and Behavior programs, City University of New York. 2001-2006 Assistant Professor, Department of Psychology, Hunter College, and Biopsychology and Ecology Evolution and Behavior programs, City University of New York. 1997-2001 Postdoctoral Research Associate, Parmly Hearing Institute, Loyola University Chicago. 1999 Lecturer in Psychology, Department of Psychology, Loyola University Chicago. 1998-2000 Postdoctoral Fellow, Neuroscience and Aging Institute, Stritch School of Medicine, Loyola University Chicago. 1991-1997 Graduate Research Assistant, University of California at San Diego. 1991 Research Assistant to Dr. W. Wheeler, Curator, American Museum of Natural History. Grants and Research Support: 2012-2013 PSC-CUNY #65638-00 43 2008-2013 RUI: Collaborative Research: The Origin and Diversification of Hearing in Malagasy-South Asian Cichlids. NSF # 0749984. 2007 Endocrine disruptors and fin-ray morphology in teleost fish: Organizational and activational effects, PSC-CUNY Equipment Grant, Co-investigator. 2007-2008 Enhanced Audition and Temporal Resolution. C. Braun Principal Investigator. PSC-CUNY #69494-00 38 2005-2006 Evolution of Hearing Specializations. -

(Hymenoptera: Chalcidoidea) De La Región Neotropical
Biota Colombiana 4 (2) 123 - 145, 2003 Lista de los géneros y especies de la familia Chalcididae (Hymenoptera: Chalcidoidea) de la región Neotropical Diana C. Arias1 y Gerard Delvare2 1 Instituto de Investigación de Recursos Biológicos “Alexander von Humboldt”, AA 8693, Bogotá, D.C., Colombia. [email protected], [email protected] 2 Departamento de Faunística y Taxonomía del CIRAD, Montpellier, Francia. [email protected] Palabras Clave: Insecta, Hymenoptera, Chalcidoidea, Chalcididae, Parasitoide, Avispas Patonas, Neotrópico El orden Hymenoptera se ha dividido tradicional- La superfamilia Chalcidoidea se caracteriza por presentar mente en dos subórdenes “Symphyta” y Apocrita, este úl- en el ala anterior una venación reducida, tan solo están timo a su vez dividido en dos grupos con categoría de sec- presentes la vena submarginal, la vena marginal, la vena ción o infraorden dependiendo de los autores, denomina- estigmal y la vena postmarginal. Adicionalmente el pronoto dos “Parasitica” o también conocidos como Terebrantes y no se extiende hasta la tégula debido a que el prepecto Aculeata (Gauld & Bolton 1988). Gauld & Hanson (1995) (esclerito, en forma de sillín o herradura) se extiende hasta abandonan esta clasificación reconociendo únicamente la tégula y separa el mesopleurón del pronoto. Otra caracte- superfamilias dentro del orden. Sin embargo muchos auto- rística de este superfamilia es la presencia de un espiráculo res siguen utilizando la división tradicional porque consi- mesotorácico visible, además algunos especimenes presen- deran que es un medio práctico para separar grandes gru- tan estructuras sensoriales en uno o más de los pos de Hymenoptera en el aspecto biológico. flagelómeros. Finalmente algunas familias exhiben coloraciones metálicas (Gibson 1993). -
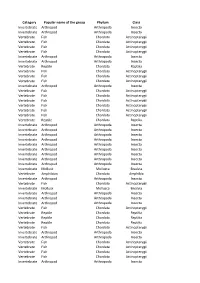
Category Popular Name of the Group Phylum Class Invertebrate
Category Popular name of the group Phylum Class Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Vertebrate Fish Chordata Actinopterygii Vertebrate Fish Chordata Actinopterygii Vertebrate Fish Chordata Actinopterygii Vertebrate Fish Chordata Actinopterygii Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Vertebrate Reptile Chordata Reptilia Vertebrate Fish Chordata Actinopterygii Vertebrate Fish Chordata Actinopterygii Vertebrate Fish Chordata Actinopterygii Invertebrate Arthropod Arthropoda Insecta Vertebrate Fish Chordata Actinopterygii Vertebrate Fish Chordata Actinopterygii Vertebrate Fish Chordata Actinopterygii Vertebrate Fish Chordata Actinopterygii Vertebrate Fish Chordata Actinopterygii Vertebrate Fish Chordata Actinopterygii Vertebrate Reptile Chordata Reptilia Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Invertebrate Mollusk Mollusca Bivalvia Vertebrate Amphibian Chordata Amphibia Invertebrate Arthropod Arthropoda Insecta Vertebrate Fish Chordata Actinopterygii Invertebrate Mollusk Mollusca Bivalvia Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Invertebrate Arthropod Arthropoda Insecta Vertebrate