Mouse Nol4 Knockout Project (CRISPR/Cas9)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Table S1. Detailed Clinical Features of Individuals with IDDCA and LADCI Syndromes
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Med Genet Table S1. Detailed clinical features of individuals with IDDCA and LADCI syndromes Lodder E., De Nittis P., Koopman C. et al., 2016 Family A Family B Family C Family D Family E Family F Individual 1 2 3 4 5 6 7 8 9 Gender, Age (years) F, 22 F, 20 F, 6 F, 11 M, 9 F, 12 F, 13 M, 8 M, 23 c.249G>A,r.249_250 c.249G>A,r.249_250 Nucleotide change c.249+1G>T/ c.249+3G>T/ c.249+3G>T/ c.906C>G/ c.242C>T/ c.242C>T/ c.242C>T/ ins249+1_249+25/ ins249+1_249+25/ (NM_006578.3) c.249+1G>T c.249+3G>T c.249+3G>T c.906C>G c.242C>T c.242C>T c.242C>T c.994C>T c.994C>T Amino acid change p.Asp84Valfs*52/ p.Asp84Valfs*52/ p.Asp84Leufs*31/ p.Asp84Valfs*31/ p.Asp84Valfs*31/ p.Tyr302*/ p.(Ser81Leu)/ p.(Ser81Leu)/ p.(Ser81Leu)/ (NP_006569.1) p.(Arg332*) p.(Arg332*) p.Asp84Leufs*31 p.Asp84Valfs*31 p.Asp84Valfs*31 p.Tyr302* p.(Ser81Leu) p.(Ser81Leu) p.(Ser81Leu) 3580 g (50th 2751 g (15 th Birth weight NA NA NA 2845 g (15th NA NA NA percentile) percentile) percentile) Ethnicity Italy Italy Jordan Puerto Rico Puerto Rico India Morocco Morocco Brazil Consanguinity − − + + + − − − + Altered speech + + NR + + + + + NA development - Verbal NA NA nonverbal unremarkable unremarkable NA NA NA NA understanding - Lexical production NA NA nonverbal delayed delayed nonverbal delayed delayed NA Intellectual + + + + + + mild mild mild disability (ID) Epilepsy + + + - -

Pancreatic Intraductal Tubulopapillary Neoplasm Is Genetically Distinct from Intraductal Papillary Mucinous Neoplasm and Ductal Adenocarcinoma
Modern Pathology (2017) 30, 1760–1772 1760 © 2017 USCAP, Inc All rights reserved 0893-3952/17 $32.00 Pancreatic intraductal tubulopapillary neoplasm is genetically distinct from intraductal papillary mucinous neoplasm and ductal adenocarcinoma Olca Basturk1, Michael F Berger1, Hiroshi Yamaguchi2, Volkan Adsay3, Gokce Askan1, Umesh K Bhanot1, Ahmet Zehir1, Fatima Carneiro4, Seung-Mo Hong5, Giuseppe Zamboni6, Esra Dikoglu7, Vaidehi Jobanputra7,8, Kazimierz O Wrzeszczynski7, Serdar Balci3, Peter Allen9, Naoki Ikari10, Shoko Takeuchi10, Hiroyuki Akagawa10, Atsushi Kanno11, Tooru Shimosegawa11, Takanori Morikawa12, Fuyuhiko Motoi12, Michiaki Unno12, Ryota Higuchi13, Masakazu Yamamoto13, Kyoko Shimizu14, Toru Furukawa15 and David S Klimstra1 1Department of Pathology, Memorial Sloan Kettering Cancer Center, New York, NY, USA; 2Department of Pathology, Tokyo Medical University, Tokyo, Japan; 3Department of Pathology, Emory University, Atlanta, GA, USA; 4Department of Pathology, Centro Hospitalar São João/Faculty of Medicine of Porto University and Institute for Research and Innovation in Health/Institute of Molecular Pathology and Immunology of the University of Porto (Ipatimup), Porto, Portugal; 5Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea; 6Department of Pathology, University of Verona, Ospedale S.C.-Don Calabria-Negrar, Verona, Italy; 7New York Genome Center, Molecular Diagnostics, New York, NY, USA; 8Department of Pathology, Colombia University Medical Center, New York, NY, USA; 9Department -

UC San Diego UC San Diego Electronic Theses and Dissertations
UC San Diego UC San Diego Electronic Theses and Dissertations Title Insights from reconstructing cellular networks in transcription, stress, and cancer Permalink https://escholarship.org/uc/item/6s97497m Authors Ke, Eugene Yunghung Ke, Eugene Yunghung Publication Date 2012 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California UNIVERSITY OF CALIFORNIA, SAN DIEGO Insights from Reconstructing Cellular Networks in Transcription, Stress, and Cancer A dissertation submitted in the partial satisfaction of the requirements for the degree Doctor of Philosophy in Bioinformatics and Systems Biology by Eugene Yunghung Ke Committee in charge: Professor Shankar Subramaniam, Chair Professor Inder Verma, Co-Chair Professor Web Cavenee Professor Alexander Hoffmann Professor Bing Ren 2012 The Dissertation of Eugene Yunghung Ke is approved, and it is acceptable in quality and form for the publication on microfilm and electronically ________________________________________________________________ ________________________________________________________________ ________________________________________________________________ ________________________________________________________________ Co-Chair ________________________________________________________________ Chair University of California, San Diego 2012 iii DEDICATION To my parents, Victor and Tai-Lee Ke iv EPIGRAPH [T]here are known knowns; there are things we know we know. We also know there are known unknowns; that is to say we know there -

Genetic Heterogeneity in Van Der Woude Syndrome: Identification of NOL4 and IRF6 Haplotype from the Noncoding Region As Candidates in Two Families
RESEARCH ARTICLE Genetic heterogeneity in Van der Woude syndrome: Identification of NOL4 and IRF6 haplotype from the noncoding region as candidates in two families Priyanka Kumari1, Akhtar Ali2, Subodh Kumar Singh3, Amit Chaurasia4, Rajiva Raman1 1Cytogenetics Laboratory, Department of Zoology, Institute of Science, Banaras Hindu University, Varanasi-221005, Uttar Pradesh, India 2Centre for Genetic Disorders, Institute of Science, Banaras Hindu University, Varanasi- 221005, Uttar Pradesh, India 3G. S. Memorial Plastic Surgery Hospital and Trauma Centre, Varanasi-221010, Uttar Pradesh, India 4Premas Life Sciences, Pvt. Ltd., New Delhi, India Running Title: Non coding region variants are involved in VWS Address for correspondence: Prof. Rajiva Raman, Cytogenetics Laboratory, Department of Zoology, Banaras Hindu University, Varanasi-221005, Uttar Pradesh, India. e-mail: [email protected], [email protected] Abstract Van der Woude Syndrome (VWS) shows autosomal dominant pattern of inheritance with 2 known candidate genes, IRF6 and GRHL3. Employing genome wide linkage analyses on 2 VWS affected families, we report co-segregation of an intronic rare variant in NOL4 in one family, and a haplotype consisting of 3 variants in non coding region of IRF6 (introns1, 8 and 3’UTR) in the other family. Using mouse as well as human embryos as model, we demonstrate expression of NOL4 in the lip and palate primordia during development. Luciferase as well as miRNA-transfection assays show decline in the expression of mutant NOL4 construct due to creation of a binding site for hsa-miR-4796-5p. In family 2, the non-coding region IRF6 haplotype turns out to be the candidate possibly by diminishing its IRF6 expression to half of its normal activity. -

A Network Pharmacology Exploring the Mechanism of a Chinese Medicine Pair for the Treatment of Gout
A Network Pharmacology Exploring the Mechanism of a Chinese Medicine Pair for the Treatment of Gout Wenjia Zhao ( [email protected] ) Shanxi hospital of traditional Chinese medicine https://orcid.org/0000-0001-9945-3517 Wei Liu First Teaching Hospital of Tianjin University of Traditional Chinese Medicine Yuanhao Wu First Teaching Hospital of Tianjin University of Traditional Chinese Medicine Original Article Keywords: targets prediction, pathways analysis, action mechanism, gout Posted Date: February 9th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-174064/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/18 Abstract Background: Gout is a crystalline-related arthropathy aroused by monosodium urate aggradation, which directly have a bearing on the disturbance of purine excretory and/or hyperuricemia, and seriously affects human health. In China, Dioscoreae Hypoglaucae Rhizoma (DH) and Smilacis Glabrae Rhixoma (SG) are widely used as medicinal pairs to prevent and treat gout. However, there is still a lack of research on the mechanism of DS (DH and SG). The aim of this study is to identify the absorbable components, potential targets and related therapeutic pathways of DS by means of network pharmacology. Material/Methods: We searched the effective chemical components of DS from TCMSP database, and explored the target of DS from GeneCards, a Bioinformatics Analysis Tool for Molecular mechANism of Traditional Chinese Medicine (BATMAN-TCM) and STITCH database. Databases such as Online Mendelian Inheritance in Man (OMIM), Therapeutic Target Database (TTD) and Pharmacogenomics Knowledgebase (PharmGkb) were used to query treatment targets for gout. -
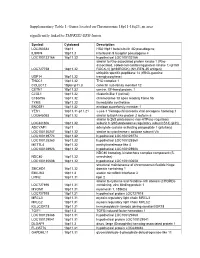
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

393LN V 393P 344SQ V 393P Probe Set Entrez Gene
393LN v 393P 344SQ v 393P Entrez fold fold probe set Gene Gene Symbol Gene cluster Gene Title p-value change p-value change chemokine (C-C motif) ligand 21b /// chemokine (C-C motif) ligand 21a /// chemokine (C-C motif) ligand 21c 1419426_s_at 18829 /// Ccl21b /// Ccl2 1 - up 393 LN only (leucine) 0.0047 9.199837 0.45212 6.847887 nuclear factor of activated T-cells, cytoplasmic, calcineurin- 1447085_s_at 18018 Nfatc1 1 - up 393 LN only dependent 1 0.009048 12.065 0.13718 4.81 RIKEN cDNA 1453647_at 78668 9530059J11Rik1 - up 393 LN only 9530059J11 gene 0.002208 5.482897 0.27642 3.45171 transient receptor potential cation channel, subfamily 1457164_at 277328 Trpa1 1 - up 393 LN only A, member 1 0.000111 9.180344 0.01771 3.048114 regulating synaptic membrane 1422809_at 116838 Rims2 1 - up 393 LN only exocytosis 2 0.001891 8.560424 0.13159 2.980501 glial cell line derived neurotrophic factor family receptor alpha 1433716_x_at 14586 Gfra2 1 - up 393 LN only 2 0.006868 30.88736 0.01066 2.811211 1446936_at --- --- 1 - up 393 LN only --- 0.007695 6.373955 0.11733 2.480287 zinc finger protein 1438742_at 320683 Zfp629 1 - up 393 LN only 629 0.002644 5.231855 0.38124 2.377016 phospholipase A2, 1426019_at 18786 Plaa 1 - up 393 LN only activating protein 0.008657 6.2364 0.12336 2.262117 1445314_at 14009 Etv1 1 - up 393 LN only ets variant gene 1 0.007224 3.643646 0.36434 2.01989 ciliary rootlet coiled- 1427338_at 230872 Crocc 1 - up 393 LN only coil, rootletin 0.002482 7.783242 0.49977 1.794171 expressed sequence 1436585_at 99463 BB182297 1 - up 393 -
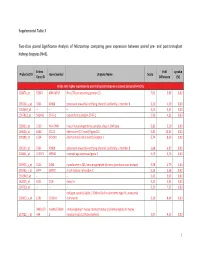
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at -

Epigenetic Variability in Cells of Normal Cytology Is Associated with the Risk of Future Morphological Transformation
Teschendorff et al. Genome Medicine 2012, 4:24 http://genomemedicine.com/content/4/3/24 RESEARCH Open Access Epigenetic variability in cells of normal cytology is associated with the risk of future morphological transformation Andrew E Teschendorff1*, Allison Jones2, Heidi Fiegl3, Alexandra Sargent4, Joanna J Zhuang1,2, Henry C Kitchener4 and Martin Widschwendter2* Abstract Background: Recently, it has been proposed that epigenetic variation may contribute to the risk of complex genetic diseases like cancer. We aimed to demonstrate that epigenetic changes in normal cells, collected years in advance of the first signs of morphological transformation, can predict the risk of such transformation. Methods: We analyzed DNA methylation (DNAm) profiles of over 27,000 CpGs in cytologically normal cells of the uterine cervix from 152 women in a prospective nested case-control study. We used statistics based on differential variability to identify CpGs associated with the risk of transformation and a novel statistical algorithm called EVORA (Epigenetic Variable Outliers for Risk prediction Analysis) to make predictions. Results: We observed many CpGs that were differentially variable between women who developed a non-invasive cervical neoplasia within 3 years of sample collection and those that remained disease-free. These CpGs exhibited heterogeneous outlier methylation profiles and overlapped strongly with CpGs undergoing age-associated DNA methylation changes in normal tissue. Using EVORA, we demonstrate that the risk of cervical neoplasia can be predicted in blind test sets (AUC = 0.66 (0.58 to 0.75)), and that assessment of DNAm variability allows more reliable identification of risk-associated CpGs than statistics based on differences in mean methylation levels. -

Computational Studies of the Genome Dynamics of Mammalian Transposable Elements and Their Relationships to Genes
COMPUTATIONAL STUDIES OF THE GENOME DYNAMICS OF MAMMALIAN TRANSPOSABLE ELEMENTS AND THEIR RELATIONSHIPS TO GENES by Ying Zhang M.Sc., Katholieke Universiteit Leuven (BELGIUM), 2004 B.E., Harbin Institute of Technology (CHINA), 1993 A THESIS SUBMITTED IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY in THE FACULTY OF GRADUATE STUDIES (Genetics) THE UNIVERSITY OF BRITISH COLUMBIA (Vancouver) May, 2012 © Ying Zhang, 2012 Abstract Sequences derived from transposable elements (TEs) comprise nearly 40 - 50% of the genomic DNA of most mammalian species, including mouse and human. However, what impact they may exert on their hosts is an intriguing question. Originally considered as merely genomic parasites or “selfish DNA”, these mobile elements show their detrimental effects through a variety of mechanisms, from physical DNA disruption to epigenetic regulation. On the other hand, evidence has been mounting to suggest that TEs sometimes may also play important roles by participating in essential biological processes in the host cell. The dual-roles of TE-host interactions make it critical for us to understand the relationship between TEs and the host, which may ultimately help us to better understand both normal cellular functions and disease. This thesis encompasses my three genome-wide computational studies of TE-gene dynamics in mammals. In the first, I identified high levels of TE insertional polymorphisms among inbred mouse strains, and systematically analyzed their distributional features and biological effects, through mining tens of millions of mouse genomic DNA sequences. In the second, I examined the properties of TEs located in introns, and identified key factors, such as the distance to the intron-exon boundary, insertional orientation, and proximity to splice sites, that influence the probability that TEs will be retained in genes. -

Supplementary Material Peptide-Conjugated Oligonucleotides Evoke Long-Lasting Myotonic Dystrophy Correction in Patient-Derived C
Supplementary material Peptide-conjugated oligonucleotides evoke long-lasting myotonic dystrophy correction in patient-derived cells and mice Arnaud F. Klein1†, Miguel A. Varela2,3,4†, Ludovic Arandel1, Ashling Holland2,3,4, Naira Naouar1, Andrey Arzumanov2,5, David Seoane2,3,4, Lucile Revillod1, Guillaume Bassez1, Arnaud Ferry1,6, Dominic Jauvin7, Genevieve Gourdon1, Jack Puymirat7, Michael J. Gait5, Denis Furling1#* & Matthew J. A. Wood2,3,4#* 1Sorbonne Université, Inserm, Association Institut de Myologie, Centre de Recherche en Myologie, CRM, F-75013 Paris, France 2Department of Physiology, Anatomy and Genetics, University of Oxford, South Parks Road, Oxford, UK 3Department of Paediatrics, John Radcliffe Hospital, University of Oxford, Oxford, UK 4MDUK Oxford Neuromuscular Centre, University of Oxford, Oxford, UK 5Medical Research Council, Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge, UK 6Sorbonne Paris Cité, Université Paris Descartes, F-75005 Paris, France 7Unit of Human Genetics, Hôpital de l'Enfant-Jésus, CHU Research Center, QC, Canada † These authors contributed equally to the work # These authors shared co-last authorship Methods Synthesis of Peptide-PMO Conjugates. Pip6a Ac-(RXRRBRRXRYQFLIRXRBRXRB)-CO OH was synthesized and conjugated to PMO as described previously (1). The PMO sequence targeting CUG expanded repeats (5′-CAGCAGCAGCAGCAGCAGCAG-3′) and PMO control reverse (5′-GACGACGACGACGACGACGAC-3′) were purchased from Gene Tools LLC. Animal model and ASO injections. Experiments were carried out in the “Centre d’études fonctionnelles” (Faculté de Médecine Sorbonne University) according to French legislation and Ethics committee approval (#1760-2015091512001083v6). HSA-LR mice are gift from Pr. Thornton. The intravenous injections were performed by single or multiple administrations via the tail vein in mice of 5 to 8 weeks of age. -

BCL6 – Regulated by Ahr/ARNT and Wild-Type MEF2B – Drives Expression of Germinal Center Markers MYBL1 and LMO2
Non-Hodgkin Lymphoma SUPPLEMENTARY APPENDIX BCL6 – regulated by AhR/ARNT and wild-type MEF2B – drives expression of germinal center markers MYBL1 and LMO2 Jie Ding, 1 Wilhelm G Dirks, 1 Stefan Ehrentraut, 1 Robert Geffers, 2 Roderick AF MacLeod, 1 Stefan Nagel, 1 Claudia Pom - merenke, 1 Julia Romani, 1 Michaela Scherr, 3 Lea AI Vaas, 1 Margarete Zaborski, 1 Hans G Drexler, 1 and Hilmar Quent - meier 1 1Leibniz-Institute DSMZ-German Collection of Microorganisms and Cell Cultures, Braunschweig; 2Helmholtz Centre for Infection Re - search, Genome Analysis Research Group, Braunschweig; and 3Medical School Hannover, Department of Hematology, Hemostasis, Oncology and Stem Cell Transplantation, Germany ©2015 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol.2014.120048 Manuscript received on November 13, 2014. Manuscript accepted on March 4, 2015. Correspondence: [email protected] SUPPLEMENTAL METHODS Gene expression analyses RNA was prepared using the RNeasy Mini kit including the RNase Free DNase Set (Qiagen). TaqMan probes (Applied Biosystems) were used to quantify human AhR (Hs00169233_m1), ARNT (Hs01121918_m1), BCL6 (Hs00153368_m1), CGGBP1 (Hs00383191_m1), CHMP2B (Hs01045897_m1), CTXN3 (Hs00416961_m1), CYP1A1 (Hs01054797_g1), GAPDH (Hs02758991_g1), ITM2B (Hs00222753_m1), LMO2 (Hs00153473_m1), MEF2B (Hs04188747_m1), MYBL1 (Hs00277143_m1), RB1 (Hs01078066_m1), SOCS1 (Hs00705164s1) and SOCS2 (Hs00919620_m1) expression levels with TATA box binding protein (TBP) as endogenous control. Relative expression levels were calculated using the Ct-method. Preparation of recombinant retroviral supernatants and retroviral transduction Retroviral supernatants were generated by calcium phosphate co-transfection of 293T cells using MSCV-BCL6-IRES-GFP (obtained from Addgene, Cambridge, MA, USA (http://www.addgene.org) or MSCV-IRES-GFP (vector control), M57 for gag/pol and pMD.g.