Are Molecular Tools Solving the Challenges Posed by Detection Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Use of Vectorette and Subvectorette PCR to Isolate Transgene Flanking DNA
Downloaded from genome.cshlp.org on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Use of Vectorette and Subvectorette PCR to Isolate Transgene Flanking DNA Maxine J. Allen, Andrew Collick, and Alec J. Jeffreys Department of Genetics, University of Leicester, Leicester LE1 7RH, UK Vectorette PCR permits the specific PCR allows specific amplification of structures and in the recovery of un- amplification of DNA segments flank- DNA fragments between two regions of known bacterial sequences. ~4-6~ In this ing a known DNA sequence. It en- known sequence to which primers can paper we detail its use in the systematic ables the application of the PCR be directed. Often, however, molecular isolation of 5'- and 3'-flanking junction where sequence information is only biological techniques require the isola- sequences around mouse transgenes of available for one primer site. We now tion and subsequent sequence analysis the minisatellite MS32. MS32 is a hu- show that vectorette PCR can be used of unknown regions of DNA. PCR can be man minisatellite composed of a tan- for the systematic mapping and re- used to retrieve this unknown sequence dem array of 29-bp repeat-unit sequence trieval of transgene flanking DNA. if it is flanking a region of known se- variants. The interspersion of these We also show that the sequence of quence. In this case, a primer can be de- sequence variants can be assayed in large vectorette PCR fragments can signed in either orientation from the the minisatellite variant repeat (MVR)- be obtained without cloning, by the known sequence. -

Download At: Amplification May Be Performed During Library Con- Fenyolab/Tipseqhunter (See Table 6)
Steranka et al. Mobile DNA (2019) 10:8 https://doi.org/10.1186/s13100-019-0148-5 METHODOLOGY Open Access Transposon insertion profiling by sequencing (TIPseq) for mapping LINE-1 insertions in the human genome Jared P. Steranka1,2, Zuojian Tang3,4, Mark Grivainis3,4, Cheng Ran Lisa Huang2, Lindsay M. Payer1, Fernanda O. R. Rego5, Thiago Luiz Araujo Miller5,6, Pedro A. F. Galante5, Sitharam Ramaswami7, Adriana Heguy7, David Fenyö3,4, Jef D. Boeke4* and Kathleen H. Burns1,2* Abstract Background: Transposable elements make up a significant portion of the human genome. Accurately locating these mobile DNAs is vital to understand their role as a source of structural variation and somatic mutation. To this end, laboratories have developed strategies to selectively amplify or otherwise enrich transposable element insertion sites in genomic DNA. Results: Here we describe a technique, Transposon Insertion Profiling by sequencing (TIPseq), to map Long INterspersed Element 1 (LINE-1, L1) retrotransposon insertions in the human genome. This method uses vectorette PCR to amplify species-specific L1 (L1PA1) insertion sites followed by paired-end Illumina sequencing. In addition to providing a step-by-step molecular biology protocol, we offer users a guide to our pipeline for data analysis, TIPseqHunter. Our recent studies in pancreatic and ovarian cancer demonstrate the ability of TIPseq to identify invariant (fixed), polymorphic (inherited variants),aswellassomatically-acquiredL1insertionsthat distinguish cancer genomes from a patient’s constitutional make-up. Conclusions: TIPseq provides an approach for amplifying evolutionarily young, active transposable element insertion sites from genomic DNA. Our rationale and variations on this protocol may be useful to those mapping L1 and other mobile elements in complex genomes. -

Loop-Mediated Isothermal Amplification (LAMP)
cells Review Loop-Mediated Isothermal Amplification (LAMP): The Better Sibling of PCR? Marianna Soroka, Barbara Wasowicz * and Anna Rymaszewska Department of Genetics and Genomics, Institute of Biology, University of Szczecin, 3c Felczaka St., 71-412 Szczecin, Poland; [email protected] (M.S.); [email protected] (A.R.) * Correspondence: [email protected] Abstract: In 1998, when the PCR technique was already popular, a Japanese company called Eiken Chemical Co., Ltd. designed a method known as the loop-mediated isothermal amplification of DNA (LAMP). The method can produce up to 109 copies of the amplified DNA within less than an hour. It is also highly specific due to the use of two to three pairs of primers (internal, external, and loop), which recognise up to eight specific locations on the DNA or RNA targets. Furthermore, the Bst DNA polymerase most used in LAMP shows a high strand displacement activity, which eliminates the DNA denaturation stage. One of the most significant advantages of LAMP is that it can be conducted at a stable temperature, for instance, in a dry block heater or an incubator. The products of LAMP can be detected much faster than in standard techniques, sometimes only requiring analysis with the naked eye. The following overview highlights the usefulness of LAMP and its effectiveness in various fields; it also considers the superiority of LAMP over PCR and presents RT-LAMP as a rapid diagnostic tool for SARS-CoV-2. Keywords: LAMP method; isothermal amplification; SARS-CoV-2 detection Citation: Soroka, M.; Wasowicz, B.; Rymaszewska, A. -

Methods and Compositions for Whole Genome Amplification and Genotyping
(19) & (11) EP 2 264 188 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 22.12.2010 Bulletin 2010/51 C12Q 1/68 (2006.01) (21) Application number: 10175210.3 (22) Date of filing: 17.06.2004 (84) Designated Contracting States: • Steemers, Frank AT BE BG CH CY CZ DE DK EE ES FI FR GB GR Encinitas, CA 92024 (US) HU IE IT LI LU MC NL PL PT RO SE SI SK TR (74) Representative: Murphy, Colm Damien et al (30) Priority: 20.06.2003 US 600634 Ipulse 08.10.2003 US 681800 26 Mallinson Road London (62) Document number(s) of the earlier application(s) in SW11 1BP (GB) accordance with Art. 76 EPC: 04776759.5 / 1 636 337 Remarks: This application was filed on 03-09-2010 as a (71) Applicant: Illumina, Inc. divisional application to the application mentioned San Diego, California 92121-1975 (US) under INID code 62. (72) Inventors: • Gunderson, Kevin Encinitas, CA 92024 (US) (54) Methods and compositions for whole genome amplification and genotyping (57) There is disclosed a method for indicating the presence of single nucleotide polymorphisms (SNPs) of interest in typable loci of a genome, comprising the steps of: (a) representationally amplifying a native genome to ob- tain a representative population of genome fragments comprising said typable loci; (b) contacting said fragments with an array of different immobilized nucleic acid probes under conditions where- in probe-fragment hybrids are formed, wherein said fragments have a concentration of at least 1 Pg/ul of DNA and wherein said immobilized nucleic acid probes comprise a region that is complementary to typable loci of a ge- nome; (c) contacting said probe-fragment hybrids with a modi- fication enzyme, whereby if the typable locus of a fragment contains a SNP of interest, a detection moiety is attached to the hybridized probe; and (d) detecting the probe, thereby indicating the presence of said single nucleotide polymorphisms of interest at typable loci of said genome. -

(12) United States Patent (10) Patent No.: US 9,546,389 B2 Li (45) Date of Patent: Jan
USOO9546,389 B2 (12) United States Patent (10) Patent No.: US 9,546,389 B2 Li (45) Date of Patent: Jan. 17, 2017 (54) METHODS AND SYSTEMS FOR NUCLEIC (56) References Cited ACIDAMPLIFICATION U.S. PATENT DOCUMENTS (71) Applicant: Coyote Bioscience Co., Ltd., Beijing 5,759,821. A * 6/1998 Teasdale .............. C12O 1/6806 (CN) 435,912 2004/0209331 A1* 10, 2004 Ririe ....................... BO1L 3,505 435,912 (72) Inventor: Xiang Li, Beijing (CN) 2008/0085541 A1 4/2008 Spangler 2012/0308990 A1* 12/2012 TeMaat ..................... BO1L, 7,52 (73) Assignee: COYOTE BIOSCIENCE CO.,LTD., 435/3 Beijing (CN) 2013/0022963 A1 1/2013 Exner et al. FOREIGN PATENT DOCUMENTS (*) Notice: Subject to any disclaimer, the term of this patent is extended or adjusted under 35 CN 101597652. A 12/2009 U.S.C. 154(b) by 0 days. CN 103074349 A 5, 2013 JP EP 1069 190 A2 * 1 2001 ............. C12O 1/686 WO WO-2012109604 A1 8, 2012 (21) Appl. No.: 14/963,986 OTHER PUBLICATIONS (22) Filed: Dec. 9, 2015 Callahan et al. Use of a portable real-time reverse transcriptase (65) Prior Publication Data polymerase chain reaction assay for rapid detection of foot-and mouth disease virus. JAVMA 220: 1636-1642 (2002).* US 2016/O115513 A1 Apr. 28, 2016 Gilbert et al. Typing of bovine viral diarrhea viruses directly from blood of persistently infected cattle by multiplex PCR. Journal of Clinical Microbiology 37:2020-2023 (1999).* International search report and written opinion dated Feb. 26, 2015 Related U.S. Application Data for PCT Application No. CN2014/094914. -

Biochemistry 24 Polymerase Chain Reaction
Biochemical Techniques Biochemistry 24 Polymerase Chain Reaction Description of Module Subject Name Biochemistry Paper Name 12 Biochemical Techniques Module Name/Title 24 Polymerase Chain Reaction Biochemical Techniques Biochemistry 24 Polymerase Chain Reaction 1. Objectives 1.1 To understand principle of Polymerase chain reaction (PCR) technique. 1.2 To explain how this techniques is performed. 1.3 What are helpful hints which help in successful performance of PCR technique? 1.4 What are potential applications of PCR? 1.5 What are different variants of PCR? 2.0 Introduction and principle Polymerase chain reaction (PCR) can be recognized as a revolutionary technology for which the inventor Kary B. Mullis was awarded with the prestigious Nobel Prize in 1993 for chemistry along with Michael Smith. The secret story of any living organism which gives it unique characteristics is coded within the large and complexed DNA of that individual. To understand the story, it was needed to bring the DNA outside the living cell into the laboratory, this gave birth to the concept of PCR. Kary Mullis conceptualized of a machine through which from a single molecule of DNA we can get multiples of its copy after the end, like a Xerox machine does. With this in 1983 Kary Mullis developed PCR which is now a most generalized technique used in medical or biological science research labs for a range of applications. Examples are its use in DNA cloning, phylogeny, functional analysis of genes, diagnosis of diseases, genetic fingerprints for forensic sciences as paternity test where the aim is to amplify a single or a few copies of a specific DNA segment in generating thousands to millions of copies of that specific DNA sequence. -

Molecular Tools for Nucleic Acid Analysis
Molecular Tools for Nucleic Acid Analysis Deirdre O’ Meara Doctoral Thesis Department of Biotechnology Royal Institute of Technology Stockholm 2001 © Deirdre O’ Meara Department of Biotechnology Royal Institute of Technology, KTH SCFAB 106 91 Stockholm Sweden Printed at Universitetsservice US AB Box 700 14 100 44 Stockholm, Sweden ISBN 91-7283-161-8 Deirdre O’ Meara (2001): Molecular Tools for Nucleic Acid Analysis. Department of Biotechnology, Royal Institute of Technology, KTH, SCFAB, Stockholm, Sweden. ISBN 91-7283-161-8 Abstract Nucleic acid technology has assumed an essential role in various areas of in vitro diagnostics ranging from infectious disease diagnosis to human genetics. An important requirement of such molecular methods is that they achieve high sensitivity and specificity with a fast turnaround time in a cost-effective manner. To this end, in this thesis we have focused on the development of sensitive nucleic acid strategies that facilitate automation and high-throughput analysis. The success of nucleic acid diagnostics in the clinical setting depends heavily on the method used for purification of the nucleic acid target from biological samples. Here we have focused on developing strategies for hybridisation capture of such templates. Using biosensor technology we observed that the hybridisation efficiency could be improved using contiguous oligonucleotide probes which acted co-operatively. By immobilising one of the probes and annealing the second probe in solution, we achieved a marked increase in target capture due to a base stacking effect between nicked oligonucleotides and/or due to the opening up of secondary structure. Such co- operatively interacting modular probes were then combined with bio-magnetic bead technology to develop a capture system for the extraction of hepatitis C RNA from serum. -

M.Sc. (MICROBIOLOGY)
SAURASHTRA UNIVERSITY RAJKOT (ACCREDITED GRADE “A” BY NAAC) FACULTY OF SCIENCE Syllabus for M. Sc. (MICROBIOLOGY) Choice Based Credit System With Effect From: 2016-17 Department of Microbiology Course Structure and Scheme of Examination For Choice Based Credit System (CBCS) (Total 96 credits) Effective from June 2016 M. Sc. Microbiology Program Outcomes (PO) PO1: Academic Competence Understanding of the subject with respect to structure, diversity, metabolisms and applications of the microorganisms is extensively developed. It also involves structure and function of biological molecules. At the end, the students gain depth of scientific knowledge regarding microorganisms. PO2: Critical thinking Students are expected to develop skills in conducting practical, designing experiments and analyzing the data reaching to conclusions. The theoretical base is developed on the various aspects of microorganisms and ability to solve the problems. PO3: Research and development Students gain ideas and competence regarding research and data analysis which will be highly useful to them in higher studies and exploring job opportunities. They acquire abilities for self direction and originality in tackling and solving problems and implementing tasks at professional levels. PO4: Personal and Behavioural Competence Professional skills concerning biochemical analysis, carrying out experiments and developing ability in specific areas of microorganisms are also developed. PO5: Effective Communication 1 During this programme, the students develop the ability for articulation of ideas, scientific writing and report reporting, conference presentations and delivering seminars. They also develop conversational competence and effective verbal and written. PO6: Social Competence The students will develop abilities to manage projects in order to achieve objectives and ability to work in team. -

Lecture 37: Polymerase Chain Reaction
Proteomics & Genomics Dr. Vikash Kumar Dubey Lecture 37: Polymerase Chain Reaction We have already studied basics of DNA/RNA structure and recombinant DNA technology in previous classes. Polymerase Chain Reaction (PCR) is another revolutionary method developed by Kary Mullis and Michael Smith. Both shared Nobel Prize in Chemistry for the work in 1993. PCR is based on ability of DNA polymerase to synthesize complementary strand to the template strand. As DNA polymerase can add a nucleotide only onto a 3'-OH group, it needs an artificial DNA strand (called DNA primer) of about 18 to 25 nucleotides complementary to 3’ end of the DNA template. As shown below, each polynucleotide has a free 3’ –OH group and 5’ phosphate group. Moreover, a DNA strand has complimentary sequence, already paired by hydrogen IIT Guwahati Page 1 of 9 Proteomics & Genomics Dr. Vikash Kumar Dubey bonding. Thus, primer can bind only when DNA strands are separated. This is generally done by heating. The primers anneal to the single-stranded DNA template at specific temperature (depends on primer sequence) and then DNA-Polymerase binds to this double stranded DNA produced. The again reaction mixture is heated to 72°C (extension); a temperature optimum for DNA- polymerase functions. This starts synthesis of the new DNA strand. Than reaction mixture is cooled to lower temperature for short term storage, if required. This completes one cycle. After first cycle, one DNA molecule has become two. After multiple cycle of the PCR reaction, the specific sequence will be accumulated in billions of copies. The PCR reaction requires the following components: DNA template: DNA template is DNA target sequence. -
Polymerase Chain Reaction. Wikipedia. Last Modified on 13
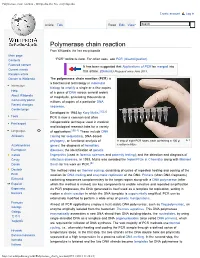
Polymerase chain reaction - Wikipedia, the free encyclopedia Create account Log in Article Talk Read Edit View Polymerase chain reaction From Wikipedia, the free encyclopedia Main page Contents "PCR" redirects here. For other uses, see PCR (disambiguation). Featured content It has been suggested that Applications of PCR be merged into Current events this article. (Discuss) Proposed since June 2013. Random article Donate to Wikipedia The polymerase chain reaction (PCR) is a biochemical technology in molecular Interaction biology to amplify a single or a few copies Help of a piece of DNA across several orders About Wikipedia of magnitude, generating thousands to Community portal millions of copies of a particular DNA Recent changes sequence. Contact page Developed in 1983 by Kary Mullis,[1][2] Tools PCR is now a common and often Print/export indispensable technique used in medical and biological research labs for a variety Languages of applications.[3][4] These include DNA Afrikaans cloning for sequencing, DNA-based phylogeny, or functional analysis of A strip of eight PCR tubes, each containing a 100 μl Azərbaycanca genes; the diagnosis of hereditary reaction mixture Български diseases; the identification of genetic Català fingerprints (used in forensic sciences and paternity testing); and the detection and diagnosis of Česky infectious diseases. In 1993, Mullis was awarded the Nobel Prize in Chemistry along with Michael Dansk Smith for his work on PCR.[5] Deutsch The method relies on thermal cycling, consisting of cycles of repeated heating and cooling of the Eesti reaction for DNA melting and enzymatic replication of the DNA. Primers (short DNA fragments) Ελληνικά containing sequences complementary to the target region along with a DNA polymerase (after Español which the method is named) are key components to enable selective and repeated amplification. -

(PCR-RFLP) and Gel Electrophoresis – Valuable Tool for Genotyping and Genetic Fingerprinting
18 Restriction Fragment Length Polymorphism Analysis of PCR-Amplified Fragments (PCR-RFLP) and Gel Electrophoresis – Valuable Tool for Genotyping and Genetic Fingerprinting Henrik Berg Rasmussen Institute of Biological Psychiatry, Mental Health Centre Sct. Hans, Copenhagen University Hospitals, Roskilde, Denmark 1. Introrduction PCR-restriction fragment length polymorphism (RFLP)-based analysis, also known as cleaved amplified polymorphic sequence (CAPS), is a popular technique for genetic analysis. It has been applied for the detection of intraspecies as well as interspecies variation. There exist several techniques that are related with PCR-RFLP and also involve gel electrophoresis including techniques for DNA fingerprinting and expression profiling. This chapter describes PCR-RFLP and related techniques. 1.1 Genetic variation There are different types of genetic variations. The so-called small-scale genetic variation includes single nucleotide polymorphisms (SNPs), multi-nucleotide polymorphisms (MNPs) and microindels. MNPs are multiple, consecutive nucleotide variations of a single common length such as double nucleotide polymorphisms (DNPs) and triple nucleotide polymorphisms (TNPs) with two and three variable nucleotides, respectively (Figure 1). Microindels are deletions, duplications and combinations thereof involving the gain or loss of 1 to 50 nucleotides (Gonzalez et al., 2007). The human genome contains more than 3 million SNPs located with an average distance of approximately 1000 bp (International HapMap Consortium 2005, Levy et al., 2007). The frequency of DNPs and TNPs, the most common forms of MNPs, amounts to ∼1% of the total number of SNPs (Rosenfeld et al., 2010). Most likely, the genome-wide occurrence of small insertions and deletions has been underestimated, probably reflecting an inaccuracy of the current sequencing techniques (Krawitz et al., 2010). -

Magdalena Sztyler Phd 2014
MOLECULAR ANALYSIS OF MICROBIAL COMMUNITIES FROM OIL INDUSTRY ENVIRONMENTS Magdalena K. Sztyler A thesis submitted in partial fulfilment of the requirements for the award of the degree of Doctor of Philosophy of the University of Portsmouth Institute of Biomedical and Biomolecular Sciences School of Pharmacy and Biomedical Sciences May 2014 “Więcej się można nauczyć podróżując...” Olga A. Jackowska Rodzicom AUTHORS’ DECLARATION Whilst registered as a candidate for the above degree, I have not been registered for any other research award. The results and conclusions embodied in this thesis are the work of the named candidate and have not been submitted for any other academic award. Magdalena K. Sztyler I ABSTRACT The effects of microbiologically influenced corrosion (MIC) can be very expensive to correct, dangerous to workers and its mechanisms are poorly understood. Understanding these processes is important so that they can be monitored and mitigated (Koch et al., 2001). It is now accepted that for the assessment of biocorrosion risks, the most powerful approach is to detect functional genes encoding the enzymes that play an important part in material deterioration (Schadt et al., 2004). The main aim of this study was to identify the microbial community present in corroded and non-corroded systems, and to detect genes that might be implicated in corrosion processes, particularly iron corrosion, so that a biochip could be designed for risk assessment of oil environments. In this thesis the microbial populations and their actives were assessed using sequencing and hybridisation techniques for three oil field sites, generating information that can help identify MIC risk. The final section of the thesis describes the development and design of functional gene probes, identified from hybridisation studies that might be included in a biochip for risk assessment in oil field environments.