University of Ghana College of Basic and Applied Sciences by Shirley Victoria Simpson (10551058) This Thesis Is Submitted To
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Reporting of Diseases and Conditions Regulation, Amendment, M.R. 289/2014
THE PUBLIC HEALTH ACT LOI SUR LA SANTÉ PUBLIQUE (C.C.S.M. c. P210) (c. P210 de la C.P.L.M.) Reporting of Diseases and Conditions Règlement modifiant le Règlement sur la Regulation, amendment déclaration de maladies et d'affections Regulation 289/2014 Règlement 289/2014 Registered December 23, 2014 Date d'enregistrement : le 23 décembre 2014 Manitoba Regulation 37/2009 amended Modification du R.M. 37/2009 1 The Reporting of Diseases and 1 Le présent règlement modifie le Conditions Regulation , Manitoba Règlement sur la déclaration de maladies et Regulation 37/2009, is amended by this d'affections , R.M. 37/2009. regulation. 2 Schedules A and B are replaced with 2 Les annexes A et B sont remplacées Schedules A and B to this regulation. par les annexes A et B du présent règlement. Coming into force Entrée en vigueur 3 This regulation comes into force on 3 Le présent règlement entre en vigueur January 1, 2015, or on the day it is registered le 1 er janvier 2015 ou à la date de son under The Statutes and Regulations Act , enregistrement en vertu de Loi sur les textes whichever is later. législatifs et réglementaires , si cette date est postérieure. December 19, 2014 Minister of Health/La ministre de la Santé, 19 décembre 2014 Sharon Blady 1 SCHEDULE A (Section 1) 1 The following diseases are diseases requiring contact notification in accordance with the disease-specific protocol. Common name Scientific or technical name of disease or its infectious agent Chancroid Haemophilus ducreyi Chlamydia Chlamydia trachomatis (including Lymphogranuloma venereum (LGV) serovars) Gonorrhea Neisseria gonorrhoeae HIV Human immunodeficiency virus Syphilis Treponema pallidum subspecies pallidum Tuberculosis Mycobacterium tuberculosis Mycobacterium africanum Mycobacterium canetti Mycobacterium caprae Mycobacterium microti Mycobacterium pinnipedii Mycobacterium bovis (excluding M. -

Melioidosis: an Emerging Infectious Disease
Review Article www.jpgmonline.com Melioidosis: An emerging infectious disease Raja NS, Ahmed MZ,* Singh NN** Department of Medical ABSTRACT Microbiology, University of Malaya Medical Center, Kuala Lumpur, Infectious diseases account for a third of all the deaths in the developing world. Achievements in understanding Malaysia, *St. the basic microbiology, pathogenesis, host defenses and expanded epidemiology of infectious diseases have Bartholomew’s Hospital, resulted in better management and reduced mortality. However, an emerging infectious disease, melioidosis, West Smithfield, London, is becoming endemic in the tropical regions of the world and is spreading to non-endemic areas. This article UK and **School of highlights the current understanding of melioidosis including advances in diagnosis, treatment and prevention. Biosciences, Cardiff Better understanding of melioidosis is essential, as it is life-threatening and if untreated, patients can succumb University, Cardiff, UK to it. Our sources include a literature review, information from international consensus meetings on melioidosis Correspondence: and ongoing discussions within the medical and scientific community. N. S. Raja, E-mail: [email protected] Received : 21-2-2005 Review completed : 20-3-2005 Accepted : 30-5-2005 PubMed ID : 16006713 KEY WORDS: Melioidosis, Burkholderia pseudomallei, Infection J Postgrad Med 2005;51:140-5 he name melioidosis [also known as Whitmore dis- in returning travellers to Europe from endemic areas.[14] The T ease] is taken from the Greek word ‘melis’ meaning geographic area of the prevalence of the organism is bound to distemper of asses and ‘eidos’ meaning resembles glanders. increase as the awareness increases. Melioidosis is a zoonotic disease caused by Pseudomonas pseudomallei [now known as Burkholderia pseudomallei], a B. -

MALDI-TOF MS for the Identification of Anaerobic Bacteria
University Medical Center Groningen Department Medical Microbiology and Infection Prevention Linda Veloo Expert Center Anaerobic Infections The Netherlands www.mmb-umcg.eu © by author ESCMID Online Lecture Library Matrix Assisted Laser Desorption/Ionization time-of-flight Mass Spectrometry (MALDI-TOF MS) Time of flight tube Target at 15-25 kV Detector Ion source © by author ESCMID- “Time of flight” Online of individual Lecture proteins is converted Library into mass information. - Spectrum is produced - Database is built Veloo et al. Anaerobe 2011; 17:211-212 Workflow Direct spotting of bacteria on target using a toothpick Add HCCA matrix Data acquisition © Databy analyses author ESCMID Online Lecture Library Log score: <1.7 no reliable identification 1.7 – 2.0 reliable genus identification ≥ 2.0 reliable species identification Obtained spectrum is unique for bacterial species Intensity © by author ESCMID Online Lecture Library Anaerobic culture Phenotypic pure culture primary incubation 2 days 2-7 days aerotolerance identification MALDI-TOF MS 1 day 2-14 days primary incubation 2-7 days © by author MALDI-TOF MS testing minutes ESCMID Online Lecture Library How many anaerobic bacteria can be identified using MALDI-TOF MS? UMCG 2011/2012 Total no. of strains 1000 Species ID 650 65% Genus ID 149 15% No ID © 201by author20% ESCMID Online Lecture Library Performance differs per genus Genus* % species ID % genus ID Clostridium sp. (n=149) 97 3 B. fragilis sp. (n=179) 97 3 Parabacteroides sp. (n=14) 93 7 GPAC (n=133) 85 15 Prevotella sp.(n=83) 78 12 Propionibacterium sp. (n=129) 64 36 Actinomyces sp. (n=28) 57 43 Fusobacterium sp. -

Microbiota of Human Precolostrum and Its Potential Role As a Source Of
www.nature.com/scientificreports OPEN Microbiota of human precolostrum and its potential role as a source of bacteria to the infant mouth Received: 24 October 2018 Lorena Ruiz1,2, Rodrigo Bacigalupe3, Cristina García-Carral2,4, Alba Boix-Amoros3, Accepted: 2 April 2019 Héctor Argüello5, Camilla Beatriz Silva2,6, Maria de los Angeles Checa7, Alex Mira3 & Published: xx xx xxxx Juan M. Rodríguez 2 Human milk represents a source of bacteria for the initial establishment of the oral (and gut) microbiomes in the breastfed infant, however, the origin of bacteria in human milk remains largely unknown. While some evidence points towards a possible endogenous enteromammary route, other authors have suggested that bacteria in human milk are contaminants from the skin or the breastfed infant mouth. In this work 16S rRNA sequencing and bacterial culturing and isolation was performed to analyze the microbiota on maternal precolostrum samples, collected from pregnant women before delivery, and on oral samples collected from the corresponding infants. The structure of both ecosystems demonstrated a high proportion of taxa consistently shared among ecosystems, Streptococcus spp. and Staphylococcus spp. being the most abundant. Whole genome sequencing on those isolates that, belonging to the same species, were isolated from both the maternal and infant samples in the same mother-infant pair, evidenced that in 8 out of 10 pairs both isolates were >99.9% identical at nucleotide level. The presence of typical oral bacteria in precolostrum before contact with the newborn indicates that they are not a contamination from the infant, and suggests that at least some oral bacteria reach the infant’s mouth through breastfeeding. -

Avoidance of Mechanisms of Innate Immune Response by Neisseria Gonorrhoeae
ADVANCEMENTS OF MICROBIOLOGY – POSTĘPY MIKROBIOLOGII 2019, 58, 4, 367–373 DOI: 10.21307/PM–2019.58.4.367 AVOIDANCE OF MECHANISMS OF INNATE IMMUNE RESPONSE BY NEISSERIA GONORRHOEAE Jagoda Płaczkiewicz* Department of Virology, Institute of Microbiology, Faculty of Biology, University of Warsaw Submitted in July, accepted in October 2019 Abstract: Neisseria gonorrhoeae (gonococcus) is a Gram-negative bacteria and an etiological agent of the sexually transmitted disease – gonorrhea. N. gonorrhoeae possesses many mechanism to evade the innate immune response of the human host. Most are related to serum resistance and avoidance of complement killing. However the clinical symptoms of gonorrhea are correlated with a significant pres- ence of neutrophils, whose response is also insufficient and modulated by gonococci. 1. Introduction. 2. Adherence ability. 3. Serum resistance and complement system. 4. Neutrophils. 4.1. Phagocytosis. 4.1.1. Oxygen- dependent intracellular killing. 4.1.2. Oxygen-independent intracellular killing. 4.2. Neutrophil extracellular traps. 4.3. Degranulation. 4.4. Apoptosis. 5. Summary UNIKANIE MECHANIZMÓW WRODZONEJ ODPOWIEDZI IMMUNOLOGICZNEJ PRZEZ NEISSERIA GONORRHOEAE Streszczenie: Neisseria gonorrhoeae (gonokok) to Gram-ujemna dwoinka będąca czynnikiem etiologicznym choroby przenoszonej drogą płciową – rzeżączki. N. gonorrhoeae posiada liczne mechanizmy umożliwiające jej unikanie wrodzonej odpowiedzi immunologicznej gospodarza. Większość z nich związana jest ze zdolnością gonokoków do manipulowania układem dopełniacza gospodarza oraz odpor- nością tej bakterii na surowicę. Jednakże symptomy infekcji N. gonorrhoeae wynikają między innymi z obecności licznych neutrofili, których aktywność jest modulowana przez gonokoki. 1. Wprowadzenie. 2. Zdolność adherencji. 3. Surowica i układ dopełniacza. 4. Neutrofile. 4.1. Fagocytoza. 4.1.1. Wewnątrzkomórkowe zabijanie zależne od tlenu. 4.1.2. -

Reportable Disease Surveillance in Virginia, 2013
Reportable Disease Surveillance in Virginia, 2013 Marissa J. Levine, MD, MPH State Health Commissioner Report Production Team: Division of Surveillance and Investigation, Division of Disease Prevention, Division of Environmental Epidemiology, and Division of Immunization Virginia Department of Health Post Office Box 2448 Richmond, Virginia 23218 www.vdh.virginia.gov ACKNOWLEDGEMENT In addition to the employees of the work units listed below, the Office of Epidemiology would like to acknowledge the contributions of all those engaged in disease surveillance and control activities across the state throughout the year. We appreciate the commitment to public health of all epidemiology staff in local and district health departments and the Regional and Central Offices, as well as the conscientious work of nurses, environmental health specialists, infection preventionists, physicians, laboratory staff, and administrators. These persons report or manage disease surveillance data on an ongoing basis and diligently strive to control morbidity in Virginia. This report would not be possible without the efforts of all those who collect and follow up on morbidity reports. Divisions in the Virginia Department of Health Office of Epidemiology Disease Prevention Telephone: 804-864-7964 Environmental Epidemiology Telephone: 804-864-8182 Immunization Telephone: 804-864-8055 Surveillance and Investigation Telephone: 804-864-8141 TABLE OF CONTENTS INTRODUCTION Introduction ......................................................................................................................................1 -

Laboratory Manual for Diagnosis of Sexually Transmitted And
Department of AIDS Control LaborLaboraattororyy ManualManual fforor DiagnosisDiagnosis ofof SeSexxuallyually TTrransmitansmittteded andand RRepreproductivoductivee TTrractact InInffectionsections FOREWORD Sexually Transmitted Infections (STIs) and Reproductive Tract Infections (RTIs) are diseases of major global concern. About 6% of Indian population is reported to be having STIs. In addition to having high levels of morbidity, they also facilitate transmission of HIV infection. Thus control of STIs goes hand in hand with control of HIV/AIDS. Countrywide strengthening of laboratories by helping them to adopt uniform standardized protocols is very important not only for case detection and treatment, but also to have reliable epidemiological information which will help in evaluation and monitoring of control efforts. It is also essential to have good referral services between primary level of health facilities and higher levels. This manual aims to bring in standard testing practices among laboratories that serve health facilities involved in managing STIs and RTIs. While generic procedures such as staining, microscopy and culture have been dealt with in detail, procedures that employ specific manufacturer defined kits have been left to the laboratories to follow the respective protocols. An introduction to quality system essentials and quality control principles has also been included in the manual to sensitize the readers on the importance of quality assurance and quality management system, which is very much the need of the hour. Manual of Operating Procedures for Diagnosis of STIs/RTIs i PREFACE Sexually Transmitted Infections (STIs) are the most common infectious diseases worldwide, with over 350 million new cases occurring each year, and have far-reaching health, social, and economic consequences. -
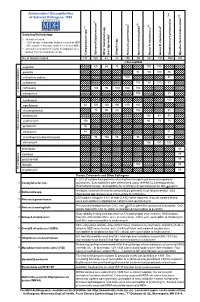
Antimicrobial Susceptibilities of Selected Pathogens, 1999
✔ Antimicrobial Susceptibilities * * † 7 of Selected Pathogens, 1999 8 e † a 2 ✔ ✔ † 3 ✔ † 4 5 6 culosis * r 1 urium spp. m L spp. Sampling Methodology 2 L † all isolates tested * ~ 20% sample of statewide isolates received at MDH spp. ~10% sample of statewide isolates received at MDH Salmonella ** all isolates tested from 7-county metropolitan area oup A streptococci ✔ oup B streptococci r isolates from a normally sterile site r Other (non-typhoidal) G Campylobacter Salmonella typhi Shigella Neisseria gonorrhoeae Neisseria meningitidis G Streptococcus pneumoni Mycobacterium tube No. of Isolates Tested 131 160 43 20 250 55 162 192 559 163 123456 123456% Susceptib123456le 123456123456 123456 123451623456 123451623456 ampicillin 1234561234566012345686 123456 15123456123456 100 100 123456123456 123451623451623451623451623456 123456 penicillin 123456123456123456123456123456 98100 100 76 123456 123456123456123456123456123456123456123456123456 123456 123451623451623451623456 123451623451623456 123456 cefuroxime sodium 123456123456123456123456 100123456123456123456 81 123456 123456123456123456123456123456123456123456123456 123456 cefotaxime 123451623451623451623451623456 100 100100 83 123456 123456123456123456123456123456 123456123456123456123456 123456 123456123456123456123456 ceftriaxone 123456 100 95 100 100 100 123451623451623451623456 -lactam antibiotics 123456123456123456123456123456 123456123456123456123456 β 123451623451623451623451623456 123451623456 123456 meropenem 123456123456123456123456123456 100 123456123456 83 123456 123456123456123456123456123456123456123456123456 -

Microbial NAD Metabolism: Lessons from Comparative Genomics
Dartmouth College Dartmouth Digital Commons Dartmouth Scholarship Faculty Work 9-2009 Microbial NAD Metabolism: Lessons from Comparative Genomics Francesca Gazzaniga Rebecca Stebbins Sheila Z. Chang Mark A. McPeek Dartmouth College Charles Brenner Carver College of Medicine Follow this and additional works at: https://digitalcommons.dartmouth.edu/facoa Part of the Biochemistry Commons, Genetics and Genomics Commons, Medicine and Health Sciences Commons, and the Microbiology Commons Dartmouth Digital Commons Citation Gazzaniga, Francesca; Stebbins, Rebecca; Chang, Sheila Z.; McPeek, Mark A.; and Brenner, Charles, "Microbial NAD Metabolism: Lessons from Comparative Genomics" (2009). Dartmouth Scholarship. 1191. https://digitalcommons.dartmouth.edu/facoa/1191 This Article is brought to you for free and open access by the Faculty Work at Dartmouth Digital Commons. It has been accepted for inclusion in Dartmouth Scholarship by an authorized administrator of Dartmouth Digital Commons. For more information, please contact [email protected]. MICROBIOLOGY AND MOLECULAR BIOLOGY REVIEWS, Sept. 2009, p. 529–541 Vol. 73, No. 3 1092-2172/09/$08.00ϩ0 doi:10.1128/MMBR.00042-08 Copyright © 2009, American Society for Microbiology. All Rights Reserved. Microbial NAD Metabolism: Lessons from Comparative Genomics Francesca Gazzaniga,1,2 Rebecca Stebbins,1,2 Sheila Z. Chang,1,2 Mark A. McPeek,2 and Charles Brenner1,3* Departments of Genetics and Biochemistry and Norris Cotton Cancer Center, Dartmouth Medical School, Lebanon, New Hampshire -

Detection of Tick-Borne Pathogens of the Genera Rickettsia, Anaplasma and Francisella in Ixodes Ricinus Ticks in Pomerania (Poland)
pathogens Article Detection of Tick-Borne Pathogens of the Genera Rickettsia, Anaplasma and Francisella in Ixodes ricinus Ticks in Pomerania (Poland) Lucyna Kirczuk 1 , Mariusz Piotrowski 2 and Anna Rymaszewska 2,* 1 Department of Hydrobiology, Faculty of Biology, Institute of Biology, University of Szczecin, Felczaka 3c Street, 71-412 Szczecin, Poland; [email protected] 2 Department of Genetics and Genomics, Faculty of Biology, Institute of Biology, University of Szczecin, Felczaka 3c Street, 71-412 Szczecin, Poland; [email protected] * Correspondence: [email protected] Abstract: Tick-borne pathogens are an important medical and veterinary issue worldwide. Environ- mental monitoring in relation to not only climate change but also globalization is currently essential. The present study aimed to detect tick-borne pathogens of the genera Anaplasma, Rickettsia and Francisella in Ixodes ricinus ticks collected from the natural environment, i.e., recreational areas and pastures used for livestock grazing. A total of 1619 specimens of I. ricinus were collected, including ticks of all life stages (adults, nymphs and larvae). The study was performed using the PCR technique. Diagnostic gene fragments msp2 for Anaplasma, gltA for Rickettsia and tul4 for Francisella were ampli- fied. No Francisella spp. DNA was detected in I. ricinus. DNA of A. phagocytophilum was detected in 0.54% of ticks and Rickettsia spp. in 3.69%. Nucleotide sequence analysis revealed that only one species of Rickettsia, R. helvetica, was present in the studied tick population. The present results are a Citation: Kirczuk, L.; Piotrowski, M.; part of a large-scale analysis aimed at monitoring the level of tick infestation in Northwest Poland. -

Antigen Detection Assay for the Diagnosis of Melioidosis
PI: Title: Antigen Detection assay for the Diagnosis of Melioidosis Received: 12/05/2013 FOA: PA10-124 Council: 05/2014 Competition ID: ADOBE-FORMS-B1 FOA Title: NIAID ADVANCED TECHNOLOGY STTR (NIAID-AT-STTR [R41/R42]) 2 R42 AI102482-03 Dual: Accession Number: 3650491 IPF: 3966401 Organization: INBIOS INTERNATIONAL, INC. Former Number: Department: IRG/SRG: ZRG1 IDM-V (12)B AIDS: N Expedited: N Subtotal Direct Costs Animals: N New Investigator: N (excludes consortium F&A) Humans: Y Early Stage Investigator: N Year 3: Clinical Trial: N Year 4: Current HS Code: E4 Year 5: HESC: N Senior/Key Personnel: Organization: Role Category: Always follow your funding opportunity's instructions for application format. Although this application demonstrates good grantsmanship, time has passed since the grantee applied. The sample may not reflect the latest format or rules. NIAID posts new samples periodically: https://www.niaid.nih.gov/grants-contracts/sample-applications The text of the application is copyrighted. You may use it only for nonprofit educational purposes provided the document remains unchanged and the PI, the grantee organization, and NIAID are credited. Note on Section 508 conformance and accessibility: We have reformatted these samples to improve accessibility for people with disabilities and users of assistive technology. If you have trouble accessing the content, please contact the NIAID Office of Knowledge and Educational Resources at [email protected]. Additions for Review Accepted Publication Accepted manuscript news Post-submission supplemental material. Information about manuscript accepted for publication. OMB Number: 4040-0001 Expiration Date: 06/30/2011 APPLICATION FOR FEDERAL ASSISTANCE 3. DATE RECEIVED BY STATE State Application Identifier SF 424 (R&R) 1. -

Non-Coding Rnas of the Q Fever Agent, Coxiella Burnetii
University of Montana ScholarWorks at University of Montana Graduate Student Theses, Dissertations, & Professional Papers Graduate School 2015 Non-coding RNAs of the Q fever agent, Coxiella burnetii Indu Ramesh Warrier The University of Montana Follow this and additional works at: https://scholarworks.umt.edu/etd Let us know how access to this document benefits ou.y Recommended Citation Warrier, Indu Ramesh, "Non-coding RNAs of the Q fever agent, Coxiella burnetii" (2015). Graduate Student Theses, Dissertations, & Professional Papers. 4620. https://scholarworks.umt.edu/etd/4620 This Dissertation is brought to you for free and open access by the Graduate School at ScholarWorks at University of Montana. It has been accepted for inclusion in Graduate Student Theses, Dissertations, & Professional Papers by an authorized administrator of ScholarWorks at University of Montana. For more information, please contact [email protected]. NON-CODING RNAS OF THE Q FEVER AGENT, COXIELLA BURNETII By INDU RAMESH WARRIER M.Sc (Med), Kasturba Medical College, Manipal, India, 2010 Dissertation presented in partial fulfillment of the requirements for the degree of Doctor of Philosophy Cellular, Molecular and Microbial Biology The University of Montana Missoula, MT August, 2015 Approved by: Sandy Ross, Dean of The Graduate School Graduate School Michael F. Minnick, Chair Division of Biological Sciences Stephen J. Lodmell Division of Biological Sciences Scott D. Samuels Division of Biological Sciences Scott Miller Division of Biological Sciences Keith Parker Department of Biomedical and Pharmaceutical Sciences Warrier, Indu, PhD, Summer 2015 Cellular, Molecular and Microbial Biology Non-coding RNAs of the Q fever agent, Coxiella burnetii Chairperson: Michael F. Minnick Coxiella burnetii is an obligate intracellular bacterial pathogen that undergoes a biphasic developmental cycle, alternating between a small cell variant (SCV) and a large cell variant (LCV).