Time-‐Dependent Effects of Antidepressant Treatments On
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Transcriptomic Analysis of Native Versus Cultured Human and Mouse Dorsal Root Ganglia Focused on Pharmacological Targets Short
bioRxiv preprint doi: https://doi.org/10.1101/766865; this version posted September 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Transcriptomic analysis of native versus cultured human and mouse dorsal root ganglia focused on pharmacological targets Short title: Comparative transcriptomics of acutely dissected versus cultured DRGs Andi Wangzhou1, Lisa A. McIlvried2, Candler Paige1, Paulino Barragan-Iglesias1, Carolyn A. Guzman1, Gregory Dussor1, Pradipta R. Ray1,#, Robert W. Gereau IV2, # and Theodore J. Price1, # 1The University of Texas at Dallas, School of Behavioral and Brain Sciences and Center for Advanced Pain Studies, 800 W Campbell Rd. Richardson, TX, 75080, USA 2Washington University Pain Center and Department of Anesthesiology, Washington University School of Medicine # corresponding authors [email protected], [email protected] and [email protected] Funding: NIH grants T32DA007261 (LM); NS065926 and NS102161 (TJP); NS106953 and NS042595 (RWG). The authors declare no conflicts of interest Author Contributions Conceived of the Project: PRR, RWG IV and TJP Performed Experiments: AW, LAM, CP, PB-I Supervised Experiments: GD, RWG IV, TJP Analyzed Data: AW, LAM, CP, CAG, PRR Supervised Bioinformatics Analysis: PRR Drew Figures: AW, PRR Wrote and Edited Manuscript: AW, LAM, CP, GD, PRR, RWG IV, TJP All authors approved the final version of the manuscript. 1 bioRxiv preprint doi: https://doi.org/10.1101/766865; this version posted September 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

4-6 Weeks Old Female C57BL/6 Mice Obtained from Jackson Labs Were Used for Cell Isolation
Methods Mice: 4-6 weeks old female C57BL/6 mice obtained from Jackson labs were used for cell isolation. Female Foxp3-IRES-GFP reporter mice (1), backcrossed to B6/C57 background for 10 generations, were used for the isolation of naïve CD4 and naïve CD8 cells for the RNAseq experiments. The mice were housed in pathogen-free animal facility in the La Jolla Institute for Allergy and Immunology and were used according to protocols approved by the Institutional Animal Care and use Committee. Preparation of cells: Subsets of thymocytes were isolated by cell sorting as previously described (2), after cell surface staining using CD4 (GK1.5), CD8 (53-6.7), CD3ε (145- 2C11), CD24 (M1/69) (all from Biolegend). DP cells: CD4+CD8 int/hi; CD4 SP cells: CD4CD3 hi, CD24 int/lo; CD8 SP cells: CD8 int/hi CD4 CD3 hi, CD24 int/lo (Fig S2). Peripheral subsets were isolated after pooling spleen and lymph nodes. T cells were enriched by negative isolation using Dynabeads (Dynabeads untouched mouse T cells, 11413D, Invitrogen). After surface staining for CD4 (GK1.5), CD8 (53-6.7), CD62L (MEL-14), CD25 (PC61) and CD44 (IM7), naïve CD4+CD62L hiCD25-CD44lo and naïve CD8+CD62L hiCD25-CD44lo were obtained by sorting (BD FACS Aria). Additionally, for the RNAseq experiments, CD4 and CD8 naïve cells were isolated by sorting T cells from the Foxp3- IRES-GFP mice: CD4+CD62LhiCD25–CD44lo GFP(FOXP3)– and CD8+CD62LhiCD25– CD44lo GFP(FOXP3)– (antibodies were from Biolegend). In some cases, naïve CD4 cells were cultured in vitro under Th1 or Th2 polarizing conditions (3, 4). -

Supplementary Table 9. Functional Annotation Clustering Results for the Union (GS3) of the Top Genes from the SNP-Level and Gene-Based Analyses (See ST4)
Supplementary Table 9. Functional Annotation Clustering Results for the union (GS3) of the top genes from the SNP-level and Gene-based analyses (see ST4) Column Header Key Annotation Cluster Name of cluster, sorted by descending Enrichment score Enrichment Score EASE enrichment score for functional annotation cluster Category Pathway Database Term Pathway name/Identifier Count Number of genes in the submitted list in the specified term % Percentage of identified genes in the submitted list associated with the specified term PValue Significance level associated with the EASE enrichment score for the term Genes List of genes present in the term List Total Number of genes from the submitted list present in the category Pop Hits Number of genes involved in the specified term (category-specific) Pop Total Number of genes in the human genome background (category-specific) Fold Enrichment Ratio of the proportion of count to list total and population hits to population total Bonferroni Bonferroni adjustment of p-value Benjamini Benjamini adjustment of p-value FDR False Discovery Rate of p-value (percent form) Annotation Cluster 1 Enrichment Score: 3.8978262119731335 Category Term Count % PValue Genes List Total Pop Hits Pop Total Fold Enrichment Bonferroni Benjamini FDR GOTERM_CC_DIRECT GO:0005886~plasma membrane 383 24.33290978 5.74E-05 SLC9A9, XRCC5, HRAS, CHMP3, ATP1B2, EFNA1, OSMR, SLC9A3, EFNA3, UTRN, SYT6, ZNRF2, APP, AT1425 4121 18224 1.18857065 0.038655922 0.038655922 0.086284383 UP_KEYWORDS Membrane 626 39.77128335 1.53E-04 SLC9A9, HRAS, -

Spatial Distribution of Leading Pacemaker Sites in the Normal, Intact Rat Sinoa
Supplementary Material Supplementary Figure 1: Spatial distribution of leading pacemaker sites in the normal, intact rat sinoatrial 5 nodes (SAN) plotted along a normalized y-axis between the superior vena cava (SVC) and inferior vena 6 cava (IVC) and a scaled x-axis in millimeters (n = 8). Colors correspond to treatment condition (black: 7 baseline, blue: 100 µM Acetylcholine (ACh), red: 500 nM Isoproterenol (ISO)). 1 Supplementary Figure 2: Spatial distribution of leading pacemaker sites before and after surgical 3 separation of the rat SAN (n = 5). Top: Intact SAN preparations with leading pacemaker sites plotted during 4 baseline conditions. Bottom: Surgically cut SAN preparations with leading pacemaker sites plotted during 5 baseline conditions (black) and exposure to pharmacological stimulation (blue: 100 µM ACh, red: 500 nM 6 ISO). 2 a &DUGLDFIoQChDQQHOV .FQM FOXVWHU &DFQDG &DFQDK *MD &DFQJ .FQLS .FQG .FQK .FQM &DFQDF &DFQE .FQM í $WSD .FQD .FQM í .FQN &DVT 5\U .FQM &DFQJ &DFQDG ,WSU 6FQD &DFQDG .FQQ &DFQDJ &DFQDG .FQD .FQT 6FQD 3OQ 6FQD +FQ *MD ,WSU 6FQE +FQ *MG .FQN .FQQ .FQN .FQD .FQE .FQQ +FQ &DFQDD &DFQE &DOP .FQM .FQD .FQN .FQG .FQN &DOP 6FQD .FQD 6FQE 6FQD 6FQD ,WSU +FQ 6FQD 5\U 6FQD 6FQE 6FQD .FQQ .FQH 6FQD &DFQE 6FQE .FQM FOXVWHU V6$1 L6$1 5$ /$ 3 b &DUGLDFReFHSWRUV $GUDF FOXVWHU $GUDD &DY &KUQE &KUP &KJD 0\O 3GHG &KUQD $GUE $GUDG &KUQE 5JV í 9LS $GUDE 7SP í 5JV 7QQF 3GHE 0\K $GUE *QDL $QN $GUDD $QN $QN &KUP $GUDE $NDS $WSE 5DPS &KUP 0\O &KUQD 6UF &KUQH $GUE &KUQD FOXVWHU V6$1 L6$1 5$ /$ 4 c 1HXURQDOPURWHLQV -

Supplementary Material Contents
Supplementary Material Contents Immune modulating proteins identified from exosomal samples.....................................................................2 Figure S1: Overlap between exosomal and soluble proteomes.................................................................................... 4 Bacterial strains:..............................................................................................................................................4 Figure S2: Variability between subjects of effects of exosomes on BL21-lux growth.................................................... 5 Figure S3: Early effects of exosomes on growth of BL21 E. coli .................................................................................... 5 Figure S4: Exosomal Lysis............................................................................................................................................ 6 Figure S5: Effect of pH on exosomal action.................................................................................................................. 7 Figure S6: Effect of exosomes on growth of UPEC (pH = 6.5) suspended in exosome-depleted urine supernatant ....... 8 Effective exosomal concentration....................................................................................................................8 Figure S7: Sample constitution for luminometry experiments..................................................................................... 8 Figure S8: Determining effective concentration ......................................................................................................... -
Identification of Spatially Associated Subpopulations by Combining
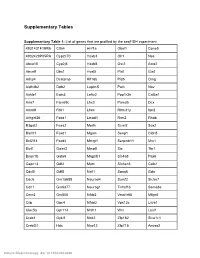
Supplementary Tables Supplementary Table 1: List of genes that are profiled by the seqFISH experiment. 4931431F19Rik Ctla4 Hnf1a Obsl1 Cpne5 4932429P05Rik Cyp2c70 Hoxb3 Olr1 Nes Abca15 Cyp2j5 Hoxb8 Osr2 Acta2 Abca9 Dbx1 Hyal5 Pld1 Gja1 Adcy4 Dcstamp Kif16b Pld5 Omg Aldh3b2 Ddb2 Laptm5 Poln Nov Ankle1 Egln3 Lefty2 Ppp1r3b Col5a1 Ano7 Fam69c Lhx3 Psmd5 Dcx Anxa9 Fbll1 Lhx4 Rbm31y Itpr2 Arhgef26 Foxa1 Lmod1 Rrm2 Rhob B3gat2 Foxa2 Mertk Scml2 Sox2 Barhl1 Foxd1 Mgam Senp1 Cldn5 Bcl2l14 Foxd4 Mmgt1 Serpinb11 Mrc1 Blzf1 Galnt3 Mmp8 Sis Tbr1 Bmpr1b Gata6 Mrgprb1 Slc4a8 Pax6 Capn13 Gdf2 Murc Slc6a16 Calb1 Cdc5l Gdf5 Nell1 Spag6 Gda Cdc6 Gm15688 Neurod4 Sumf2 Slc5a7 Cdh1 Gm6377 Neurog1 Tnfrsf1b Sema3e Cecr2 Gm805 Nfkb2 Vmn1r65 Mfge8 Cilp Gpc4 Nfkbiz Vps13c Lyve1 Clec5a Gpr114 Nhlh1 Wrn Loxl1 Creb1 Gykl1 Nkd2 Zfp182 Slco1c1 Creb3l1 Hdx Nlrp12 Zfp715 Amigo2 Nature Biotechnology: doi:10.1038/nbt.4260 Csf2rb2 Hn1l Npy2r Zfp90 Kcnip2 Supplementary Table 2: List of 43 genes used for cell type mapping. Fbll1 Itpr2 Vps13c Tnfrsf1b Sox2 Hdx Wrn Sumf2 Vmn1r65 Rhob Mrgprb1 Calb1 Pld1 Laptm5 Tbr1 Slc5a7 Abca9 Ankle1 Olr1 Cecr2 Cpne5 Blzf1 Mertk Nell1 Npy2r Cdc5l Slco1c1 Pax6 Cldn5 Cyp2j5 Mfge8 Col5a1 Bmpr1b Rrm2 Gja1 Dcx Spag6 Csf2rb2 Gda Arhgef26 Slc4a8 Gm805 Omg Supplementary Table 3: Astrocyte prediction accuracy, evaluated using fluorescent staining images. We examined and contrasted DAPI, Nissl staining on astrocyte cells. Percentage of cells with no Nissl, and with DAPI staining are recorded (these are accurate instances) % predicted % predicted Total predicted astrocytes with weak astrocytes with astrocytes or no Nissl stain present Nissl stain Cortex Column 1 100% 0% 11 Cortex Column 2 88% 12% 25 Cortex Column 3 87% 13% 24 Cortex Column 4 87% 13% 24 Supplementary Table 4: List of 69 genes used for HMRF. -

Epigenetic Mechanisms Are Involved in the Oncogenic Properties of ZNF518B in Colorectal Cancer
Epigenetic mechanisms are involved in the oncogenic properties of ZNF518B in colorectal cancer Francisco Gimeno-Valiente, Ángela L. Riffo-Campos, Luis Torres, Noelia Tarazona, Valentina Gambardella, Andrés Cervantes, Gerardo López-Rodas, Luis Franco and Josefa Castillo SUPPLEMENTARY METHODS 1. Selection of genomic sequences for ChIP analysis To select the sequences for ChIP analysis in the five putative target genes, namely, PADI3, ZDHHC2, RGS4, EFNA5 and KAT2B, the genomic region corresponding to the gene was downloaded from Ensembl. Then, zoom was applied to see in detail the promoter, enhancers and regulatory sequences. The details for HCT116 cells were then recovered and the target sequences for factor binding examined. Obviously, there are not data for ZNF518B, but special attention was paid to the target sequences of other zinc-finger containing factors. Finally, the regions that may putatively bind ZNF518B were selected and primers defining amplicons spanning such sequences were searched out. Supplementary Figure S3 gives the location of the amplicons used in each gene. 2. Obtaining the raw data and generating the BAM files for in silico analysis of the effects of EHMT2 and EZH2 silencing The data of siEZH2 (SRR6384524), siG9a (SRR6384526) and siNon-target (SRR6384521) in HCT116 cell line, were downloaded from SRA (Bioproject PRJNA422822, https://www.ncbi. nlm.nih.gov/bioproject/), using SRA-tolkit (https://ncbi.github.io/sra-tools/). All data correspond to RNAseq single end. doBasics = TRUE doAll = FALSE $ fastq-dump -I --split-files SRR6384524 Data quality was checked using the software fastqc (https://www.bioinformatics.babraham. ac.uk /projects/fastqc/). The first low quality removing nucleotides were removed using FASTX- Toolkit (http://hannonlab.cshl.edu/fastxtoolkit/). -

1 Fibrillar Αβ Triggers Microglial Proteome Alterations and Dysfunction in Alzheimer Mouse 1 Models 2 3 4 Laura Sebastian
bioRxiv preprint doi: https://doi.org/10.1101/861146; this version posted December 2, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Fibrillar triggers microglial proteome alterations and dysfunction in Alzheimer mouse 2 models 3 4 5 Laura Sebastian Monasor1,10*, Stephan A. Müller1*, Alessio Colombo1, Jasmin König1,2, Stefan 6 Roth3, Arthur Liesz3,4, Anna Berghofer5, Takashi Saito6,7, Takaomi C. Saido6, Jochen Herms1,4,8, 7 Michael Willem9, Christian Haass1,4,9, Stefan F. Lichtenthaler 1,4,5# & Sabina Tahirovic1# 8 9 1 German Center for Neurodegenerative Diseases (DZNE) Munich, 81377 Munich, Germany 10 2 Faculty of Chemistry, Technical University of Munich, Garching, Germany 11 3 Institute for Stroke and Dementia Research (ISD), Ludwig-Maximilians Universität München, 12 81377 Munich, Germany 13 4 Munich Cluster for Systems Neurology (SyNergy), Munich, Germany 14 5 Neuroproteomics, School of Medicine, Klinikum Rechts der Isar, Technical University of Munich, 15 Munich, Germany 16 6 Laboratory for Proteolytic Neuroscience, RIKEN Center for Brain Science Institute, Wako, 17 Saitama 351-0198, Japan 18 7 Department of Neurocognitive Science, Nagoya City University Graduate School of Medical 19 Science, Nagoya, Aichi 467-8601, Japan 20 8 Center for Neuropathology and Prion Research, Ludwig-Maximilians-Universität München, 81377 21 Munich, Germany 22 9 Biomedical Center (BMC), Ludwig-Maximilians Universität München, 81377 Munich, Germany 23 10 Graduate School of Systemic Neuroscience, Ludwig-Maximilians-University Munich, Munich, 24 Germany. 25 *Contributed equally 26 #Correspondence: [email protected] and [email protected] 27 28 Running title: 29 Microglial proteomic signatures of AD 30 Keywords: Alzheimer’s disease / microglia / proteomic signatures / neuroinflammation / 31 phagocytosis 32 1 bioRxiv preprint doi: https://doi.org/10.1101/861146; this version posted December 2, 2019. -

1 1 2 3 Cell Type-Specific Transcriptomics of Hypothalamic
1 2 3 4 Cell type-specific transcriptomics of hypothalamic energy-sensing neuron responses to 5 weight-loss 6 7 Fredrick E. Henry1,†, Ken Sugino1,†, Adam Tozer2, Tiago Branco2, Scott M. Sternson1,* 8 9 1Janelia Research Campus, Howard Hughes Medical Institute, 19700 Helix Drive, Ashburn, VA 10 20147, USA. 11 2Division of Neurobiology, Medical Research Council Laboratory of Molecular Biology, 12 Cambridge CB2 0QH, UK 13 14 †Co-first author 15 *Correspondence to: [email protected] 16 Phone: 571-209-4103 17 18 Authors have no competing interests 19 1 20 Abstract 21 Molecular and cellular processes in neurons are critical for sensing and responding to energy 22 deficit states, such as during weight-loss. AGRP neurons are a key hypothalamic population 23 that is activated during energy deficit and increases appetite and weight-gain. Cell type-specific 24 transcriptomics can be used to identify pathways that counteract weight-loss, and here we 25 report high-quality gene expression profiles of AGRP neurons from well-fed and food-deprived 26 young adult mice. For comparison, we also analyzed POMC neurons, an intermingled 27 population that suppresses appetite and body weight. We find that AGRP neurons are 28 considerably more sensitive to energy deficit than POMC neurons. Furthermore, we identify cell 29 type-specific pathways involving endoplasmic reticulum-stress, circadian signaling, ion 30 channels, neuropeptides, and receptors. Combined with methods to validate and manipulate 31 these pathways, this resource greatly expands molecular insight into neuronal regulation of 32 body weight, and may be useful for devising therapeutic strategies for obesity and eating 33 disorders. -

Supplementary Information
Supplementary Information Ten-eleven translocation 1 Mediated-DNA Hydroxymethylation is Required for Myelination and Remyelination in the Mouse Brain Ming Zhang1, Jian Wang1, Kaixiang Zhang1, Guozhen Lu1, Yuming Liu1, Keke Ren1, Wenting Wang1, Dazhuan Xin2, Lingli Xu3, Honghui Mao1, Junlin Xing1, Xingchun Gao4, Weilin Jin5, Kalen 2 6 1, 2, 1, Berry , Katsuhiko Mikoshiba , Shengxi Wu *, Q. Richard Lu * and Xianghui Zhao * 1 Supplementary figures: Supplementary Fig. 1 Distribution pattern of genomic 5hmCs is associated with the transition from NPCs to OPCs (a-c) Examples of OL lineage-specific genes expressed in different OL stages. Fragments per kilobase of transcript per million mapped reads (FPKMs) of representative genes highly expressed in OPCs (a), iOLs (b), and OLs (c). Diagrams were made from http://jiaqianwulab. org/braincell/RNASeq.html. (d) The level of 5hmC in certain gene loci is identified by qPCR assay. Validation of loci specific 5hmC modification in OPC and NPC cultures by qPCR analysis of OPC associated genes, immature OLs (iOL) associated genes, mOL associated genes and negative regulators of OPC differentiation. Data are Means ± SEM (n = 3 independent cultures per group). *, p<0.05, **, p<0.01, ***, p<0.001, compared to NPC. Two-tailed unpaired t test. Cspg4: t = -5.867, df = 4, p = 0.004; Slc22a3: t = -10.100, df = 4, p = 0.001; Wnt10a: t = -20.501, df = 4, p = 0.000033; Kndc1: t = -13.045, df = 4, p = 0.000199; Olfr279: t = -4.221, df = 4, p = 0.013; Zdhhc12: t = -5.173, df = 4, p = 0.007; Mag: t = -32.333, df = 4, p = 0.000005; Elovl7: t = -4.274, df = 4, p = 0.013; Arl2: t = -4.319, df = 4, p = 0.012; Cdc42ep2: t = -3.081, df = 4, p = 0.037; Ngf: t = 27.294, df = 4, p = 0.037; Zfp28: t = 5.877, df = 4, p = 0.004.