Multimerization of Homo Sapiens TRPA1 Ion Channel Cytoplasmic Domains
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Realizing the Allosteric Potential of the Tetrameric Protein Kinase a RIΑ Holoenzyme
Structure Article Realizing the Allosteric Potential of the Tetrameric Protein Kinase A RIa Holoenzyme Angela J. Boettcher,1,6 Jian Wu,1,6 Choel Kim,2 Jie Yang,1 Jessica Bruystens,1 Nikki Cheung,1 Juniper K. Pennypacker,1,3 Donald A. Blumenthal,4 Alexandr P. Kornev,3,5 and Susan S. Taylor1,3,5,* 1Department of Chemistry and Biochemistry, University of California at San Diego, La Jolla, CA 92093, USA 2Department of Pharmacology, Baylor College of Medicine, Houston, TX 77030, USA 3Department of Pharmacology, University of California at San Diego, La Jolla, CA 92093, USA 4Department of Pharmacology and Toxicology, University of Utah, Salt Lake City, UT 84112, USA 5Howard Hughes Medical Institute, University of California at San Diego, La Jolla, CA 92093, USA 6These authors contributed equally to this work *Correspondence: [email protected] DOI 10.1016/j.str.2010.12.005 SUMMARY the active site cleft in the C subunit in the inactive holoenzyme but is disordered in the dissociated free R subunits (Li et al., PKA holoenzymes containing two catalytic (C) 2000). The linker, as summarized in Figure 1, can be divided subunits and a regulatory (R) subunit dimer are acti- into three segments, the consensus inhibitor site (P-3 to P+1), vated cooperatively by cAMP. While cooperativity the N-linker that joins the inhibitor site to the D/D domain, and involves the two tandem cAMP binding domains in the C-linker that becomes ordered in the heterodimeric holoen- each R-subunit, additional cooperativity is associ- zyme complex. While much has been learned from the structures ated with the tetramer. -

Trpc, Trpv and Vascular Disease | Encyclopedia
TRPC, TRPV and Vascular Disease Subjects: Biochemistry Submitted by: Tarik Smani Hajami Definition Ion channels play an important role in vascular function and pathology. In this review we gave an overview of recent findings and discussed the role of TRPC and TRPV channels as major regulators of cellular remodeling and consequent vascular disorders. Here, we focused on their implication in 4 relevant vascular diseases: systemic and pulmonary artery hypertension, atherosclerosis and restenosis. Transient receptor potentials (TRPs) are non-selective cation channels that are widely expressed in vascular beds. They contribute to the Ca2+ influx evoked by a wide spectrum of chemical and physical stimuli, both in endothelial and vascular smooth muscle cells. Within the superfamily of TRP channels, different isoforms of TRPC (canonical) and TRPV (vanilloid) have emerged as important regulators of vascular tone and blood flow pressure. Additionally, several lines of evidence derived from animal models, and even from human subjects, highlighted the role of TRPC and TRPV in vascular remodeling and disease. Dysregulation in the function and/or expression of TRPC and TRPV isoforms likely regulates vascular smooth muscle cells switching from a contractile to a synthetic phenotype. This process contributes to the development and progression of vascular disorders, such as systemic and pulmonary arterial hypertension, atherosclerosis and restenosis. 1. Introduction Blood vessels are composed essentially of two interacting cell types: endothelial cells (ECs) from the tunica intima lining of the vessel wall and vascular smooth muscle cells (VSMCs) from tunica media of the vascular tube. Blood vessels are a complex network, and they differ according to the tissue to which they belong, having diverse cell expressions, structures and functions [1][2][3]. -

A Perspective on Mechanisms of Protein Tetramer Formation
Biophysical Journal Volume 85 December 2003 3587–3599 3587 A Perspective on Mechanisms of Protein Tetramer Formation Evan T. Powers* and David L. Powersy *Department of Chemistry, The Scripps Research Institute, La Jolla, California 92037; and yDepartment of Mathematics and Computer Science, Clarkson University, Potsdam, New York 13699 ABSTRACT Homotetrameric proteins can assemble by several different pathways, but have only been observed to use one, in which two monomers associate to form a homodimer, and then two homodimers associate to form a homotetramer. To determine why this pathway should be so uniformly dominant, we have modeled the kinetics of tetramerization for the possible pathways as a function of the rate constants for each step. We have found that competition with the other pathways, in which homotetramers can be formed either by the association of two different types of homodimers or by the successive addition of monomers to homodimers and homotrimers, can cause substantial amounts of protein to be trapped as intermediates of the assembly pathway. We propose that this could lead to undesirable consequences for an organism, and that selective pressure may have caused homotetrameric proteins to evolve to assemble by a single pathway. INTRODUCTION Many proteins must be homotetrameric to be functional. ‘‘MDT’’ stands for monomer-dimer-tetramer and ‘‘a’’ and Prominent examples include transcription factors (e.g., p53) ‘‘b’’ indicate the type of homodimer formed. The pathways (Friedman et al., 1993), transport proteins (e.g., trans- that include monomers, homodimers, homotrimers, and thyretin) (Blake et al., 1974), potassium channels (Deutsch, homotetramers will be denoted MDRT, which stands for 2002; Miller, 2000), water channels (Fujiyoshi et al., 2002), monomer-dimer-trimer-tetramer. -

Transient Receptor Potential (TRP) Channels in Haematological Malignancies: an Update
biomolecules Review Transient Receptor Potential (TRP) Channels in Haematological Malignancies: An Update Federica Maggi 1,2 , Maria Beatrice Morelli 2 , Massimo Nabissi 2 , Oliviero Marinelli 2 , Laura Zeppa 2, Cristina Aguzzi 2, Giorgio Santoni 2 and Consuelo Amantini 3,* 1 Department of Molecular Medicine, Sapienza University, 00185 Rome, Italy; [email protected] 2 Immunopathology Laboratory, School of Pharmacy, University of Camerino, 62032 Camerino, Italy; [email protected] (M.B.M.); [email protected] (M.N.); [email protected] (O.M.); [email protected] (L.Z.); [email protected] (C.A.); [email protected] (G.S.) 3 Immunopathology Laboratory, School of Biosciences and Veterinary Medicine, University of Camerino, 62032 Camerino, Italy * Correspondence: [email protected]; Tel.: +30-0737403312 Abstract: Transient receptor potential (TRP) channels are improving their importance in differ- ent cancers, becoming suitable as promising candidates for precision medicine. Their important contribution in calcium trafficking inside and outside cells is coming to light from many papers published so far. Encouraging results on the correlation between TRP and overall survival (OS) and progression-free survival (PFS) in cancer patients are available, and there are as many promising data from in vitro studies. For what concerns haematological malignancy, the role of TRPs is still not elucidated, and data regarding TRP channel expression have demonstrated great variability throughout blood cancer so far. Thus, the aim of this review is to highlight the most recent findings Citation: Maggi, F.; Morelli, M.B.; on TRP channels in leukaemia and lymphoma, demonstrating their important contribution in the Nabissi, M.; Marinelli, O.; Zeppa, L.; perspective of personalised therapies. -

HOOK™ Maleimide Activated Streptavidin for Conjugation of Streptavidin to Sulfhydryl Groups Containing Proteins, Peptides and Ligands
G-Biosciences 1-800-628-7730 1-314-991-6034 [email protected] A Geno Technology, Inc. (USA) brand name HOOK™ Maleimide Activated Streptavidin For conjugation of Streptavidin to sulfhydryl groups containing proteins, peptides and ligands (Cat. #786-1653, 786-1654) think proteins! think G-Biosciences www.GBiosciences.com INTRODUCTION ................................................................................................................. 3 ITEMS SUPPLIED ................................................................................................................ 4 STORAGE CONDITIONS ...................................................................................................... 4 ADDITIONAL ITEMS NEEDED .............................................................................................. 4 IMPORTANT INFORMATION .............................................................................................. 4 PROTOCOL ......................................................................................................................... 4 PREPARATION OF PROTEIN FOR CONJUGATION TO MALEIMIDE ACTIVATED PROTEIN 4 CONJUGATION REACTION ............................................................................................. 5 STORAGE OF CONJUGATED ANTIBODIES/PROTEINS ......................................................... 5 RELATED PRODUCTS .......................................................................................................... 5 Page 2 of 6 INTRODUCTION Streptavidin is a non-glycosylated -

TRPC1 Regulates the Activity of a Voltage-Dependent Nonselective Cation Current in Hippocampal CA1 Neurons
cells Article TRPC1 Regulates the Activity of a Voltage-Dependent Nonselective Cation Current in Hippocampal CA1 Neurons 1, 1 1,2 1,3, Frauke Kepura y, Eva Braun , Alexander Dietrich and Tim D. Plant * 1 Pharmakologisches Institut, BPC-Marburg, Fachbereich Medizin, Philipps-Universität Marburg, Karl-von-Frisch-Straße 2, 35043 Marburg, Germany; [email protected] (F.K.); braune@staff.uni-marburg.de (E.B.); [email protected] (A.D.) 2 Walther-Straub-Institut für Pharmakologie und Toxikologie, Ludwig-Maximilians-Universität München, 80336 München, Germany 3 Center for Mind, Brain and Behavior, Philipps-Universität Marburg, 35032 Marburg, Germany * Correspondence: plant@staff.uni-marburg.de; Tel.: +49-6421-28-65038 Present address: Institut für Bodenkunde und Pflanzenernährung/Institut für angewandte Ökologie, y Hochschule Geisenheim University, Von-Lade-Str. 1, 65366 Geisenheim, Germany. Received: 27 November 2019; Accepted: 14 February 2020; Published: 18 February 2020 Abstract: The cation channel subunit TRPC1 is strongly expressed in central neurons including neurons in the CA1 region of the hippocampus where it forms complexes with TRPC4 and TRPC5. To investigate the functional role of TRPC1 in these neurons and in channel function, we compared current responses to group I metabotropic glutamate receptor (mGluR I) activation and looked +/+ / for major differences in dendritic morphology in neurons from TRPC1 and TRPC1− − mice. mGluR I stimulation resulted in the activation of a voltage-dependent nonselective cation current in both genotypes. Deletion of TRPC1 resulted in a modification of the shape of the current-voltage relationship, leading to an inward current increase. In current clamp recordings, the percentage of neurons that responded to depolarization in the presence of an mGluR I agonist with a plateau / potential was increased in TRPC1− − mice. -

Structure of the Mouse TRPC4 Ion Channel 1 2 Jingjing
bioRxiv preprint doi: https://doi.org/10.1101/282715; this version posted March 15, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Structure of the mouse TRPC4 ion channel 2 3 Jingjing Duan1,2*, Jian Li1,3*, Bo Zeng4*, Gui-Lan Chen4, Xiaogang Peng5, Yixing Zhang1, 4 Jianbin Wang1, David E. Clapham2, Zongli Li6#, Jin Zhang1# 5 6 7 1School of Basic Medical Sciences, Nanchang University, Nanchang, Jiangxi, 330031, China. 8 2Howard Hughes Medical Institute, Janelia Research Campus, Ashburn, VA 20147, USA 9 3Department of Molecular and Cellular Biochemistry, University of Kentucky, Lexington, KY 10 40536, USA. 11 4Key Laboratory of Medical Electrophysiology, Ministry of Education, and Institute of 12 Cardiovascular Research, Southwest Medical University, Luzhou, Sichuan, 646000, China 13 5The Key Laboratory of Molecular Medicine, the Second Affiliated Hospital of Nanchang 14 University, Nanchang 330006, China. 15 6Howard Hughes Medical Institute, Department of Biological Chemistry and Molecular 16 Pharmacology, Harvard Medical School, Boston, MA 02115, USA. 17 18 *These authors contributed equally to this work. 19 # Corresponding authors bioRxiv preprint doi: https://doi.org/10.1101/282715; this version posted March 15, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 20 Abstract 21 Members of the transient receptor potential (TRP) ion channels conduct cations into cells. They 22 mediate functions ranging from neuronally-mediated hot and cold sensation to intracellular 23 organellar and primary ciliary signaling. -

TRPM8 Channels and Dry Eye
UC Berkeley UC Berkeley Previously Published Works Title TRPM8 Channels and Dry Eye. Permalink https://escholarship.org/uc/item/2gz2d8s3 Journal Pharmaceuticals (Basel, Switzerland), 11(4) ISSN 1424-8247 Authors Yang, Jee Myung Wei, Edward T Kim, Seong Jin et al. Publication Date 2018-11-15 DOI 10.3390/ph11040125 Peer reviewed eScholarship.org Powered by the California Digital Library University of California pharmaceuticals Review TRPM8 Channels and Dry Eye Jee Myung Yang 1,2 , Edward T. Wei 3, Seong Jin Kim 4 and Kyung Chul Yoon 1,* 1 Department of Ophthalmology, Chonnam National University Medical School and Hospital, Gwangju 61469, Korea; [email protected] 2 Graduate School of Medical Science and Engineering, Korea Advanced Institute of Science and Technology, Daejeon 34141, Korea 3 School of Public Health, University of California, Berkeley, CA 94720, USA; [email protected] 4 Department of Dermatology, Chonnam National University Medical School and Hospital, Gwangju 61469, Korea; [email protected] * Correspondence: [email protected] Received: 17 September 2018; Accepted: 12 November 2018; Published: 15 November 2018 Abstract: Transient receptor potential (TRP) channels transduce signals of chemical irritation and temperature change from the ocular surface to the brain. Dry eye disease (DED) is a multifactorial disorder wherein the eyes react to trivial stimuli with abnormal sensations, such as dryness, blurring, presence of foreign body, discomfort, irritation, and pain. There is increasing evidence of TRP channel dysfunction (i.e., TRPV1 and TRPM8) in DED pathophysiology. Here, we review some of this literature and discuss one strategy on how to manage DED using a TRPM8 agonist. -

New Natural Agonists of the Transient Receptor Potential Ankyrin 1 (TRPA1
www.nature.com/scientificreports OPEN New natural agonists of the transient receptor potential Ankyrin 1 (TRPA1) channel Coline Legrand, Jenny Meylan Merlini, Carole de Senarclens‑Bezençon & Stéphanie Michlig* The transient receptor potential (TRP) channels family are cationic channels involved in various physiological processes as pain, infammation, metabolism, swallowing function, gut motility, thermoregulation or adipogenesis. In the oral cavity, TRP channels are involved in chemesthesis, the sensory chemical transduction of spicy ingredients. Among them, TRPA1 is activated by natural molecules producing pungent, tingling or irritating sensations during their consumption. TRPA1 can be activated by diferent chemicals found in plants or spices such as the electrophiles isothiocyanates, thiosulfnates or unsaturated aldehydes. TRPA1 has been as well associated to various physiological mechanisms like gut motility, infammation or pain. Cinnamaldehyde, its well known potent agonist from cinnamon, is reported to impact metabolism and exert anti-obesity and anti-hyperglycemic efects. Recently, a structurally similar molecule to cinnamaldehyde, cuminaldehyde was shown to possess anti-obesity and anti-hyperglycemic efect as well. We hypothesized that both cinnamaldehyde and cuminaldehyde might exert this metabolic efects through TRPA1 activation and evaluated the impact of cuminaldehyde on TRPA1. The results presented here show that cuminaldehyde activates TRPA1 as well. Additionally, a new natural agonist of TRPA1, tiglic aldehyde, was identifed -
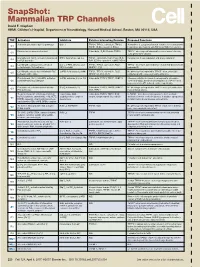
Snapshot: Mammalian TRP Channels David E
SnapShot: Mammalian TRP Channels David E. Clapham HHMI, Children’s Hospital, Department of Neurobiology, Harvard Medical School, Boston, MA 02115, USA TRP Activators Inhibitors Putative Interacting Proteins Proposed Functions Activation potentiated by PLC pathways Gd, La TRPC4, TRPC5, calmodulin, TRPC3, Homodimer is a purported stretch-sensitive ion channel; form C1 TRPP1, IP3Rs, caveolin-1, PMCA heteromeric ion channels with TRPC4 or TRPC5 in neurons -/- Pheromone receptor mechanism? Calmodulin, IP3R3, Enkurin, TRPC6 TRPC2 mice respond abnormally to urine-based olfactory C2 cues; pheromone sensing 2+ Diacylglycerol, [Ca ]I, activation potentiated BTP2, flufenamate, Gd, La TRPC1, calmodulin, PLCβ, PLCγ, IP3R, Potential role in vasoregulation and airway regulation C3 by PLC pathways RyR, SERCA, caveolin-1, αSNAP, NCX1 La (100 µM), calmidazolium, activation [Ca2+] , 2-APB, niflumic acid, TRPC1, TRPC5, calmodulin, PLCβ, TRPC4-/- mice have abnormalities in endothelial-based vessel C4 i potentiated by PLC pathways DIDS, La (mM) NHERF1, IP3R permeability La (100 µM), activation potentiated by PLC 2-APB, flufenamate, La (mM) TRPC1, TRPC4, calmodulin, PLCβ, No phenotype yet reported in TRPC5-/- mice; potentially C5 pathways, nitric oxide NHERF1/2, ZO-1, IP3R regulates growth cones and neurite extension 2+ Diacylglycerol, [Ca ]I, 20-HETE, activation 2-APB, amiloride, Cd, La, Gd Calmodulin, TRPC3, TRPC7, FKBP12 Missense mutation in human focal segmental glomerulo- C6 potentiated by PLC pathways sclerosis (FSGS); abnormal vasoregulation in TRPC6-/- -

Involvement of TRPC4 and 5 Channels in Persistent Firing in Hippocampal CA1 Pyramidal Cells
cells Article Involvement of TRPC4 and 5 Channels in Persistent Firing in Hippocampal CA1 Pyramidal Cells Alberto Arboit 1,2,3, Antonio Reboreda 1,4 and Motoharu Yoshida 1,3,4,5,* 1 German Center for Neurodegenerative Diseases (DZNE), 39120 Magdeburg, Germany; [email protected] (A.A.); [email protected] (A.R.) 2 Otto-von-Guericke University, 39120 Magdeburg, Germany 3 Faculty of Psychology, Ruhr University Bochum (RUB), Universitätsstraße 150, 44801 Bochum, Germany 4 Leibniz Institute for Neurobiology (LIN), 39118 Magdeburg, Germany 5 Center for Behavioral Brain Sciences (CBBS), 39106 Magdeburg, Germany * Correspondence: [email protected] Received: 1 December 2019; Accepted: 1 February 2020; Published: 5 February 2020 Abstract: Persistent neural activity has been observed in vivo during working memory tasks, and supports short-term (up to tens of seconds) retention of information. While synaptic and intrinsic cellular mechanisms of persistent firing have been proposed, underlying cellular mechanisms are not yet fully understood. In vitro experiments have shown that individual neurons in the hippocampus and other working memory related areas support persistent firing through intrinsic cellular mechanisms that involve the transient receptor potential canonical (TRPC) channels. Recent behavioral studies demonstrating the involvement of TRPC channels on working memory make the hypothesis that TRPC driven persistent firing supports working memory a very attractive one. However, this view has been challenged by recent findings that persistent firing in vitro is unchanged in TRPC knock out (KO) mice. To assess the involvement of TRPC channels further, we tested novel and highly specific TRPC channel blockers in cholinergically induced persistent firing in mice CA1 pyramidal cells for the first time. -

Regulation of Ion Channels by Muscarinic Receptors
Regulation of ion channels by muscarinic receptors David A. Brown Department of Neuroscience, Physiology & Pharmacology, University College London, London, WC1E 6BT (UK). Contact address: Professor D.A.Brown, FRS, Department of Neuroscience, Physiology & Pharmacology, University College London, Gower Street, Londin, WC1E 6BT. E-mail: [email protected] Telephone: (+44) (0)20 7679 7297 Mobile (for urgent messages): (+44)(0)7766-236330 Abstract The excitable behaviour of neurons is determined by the activity of their endogenous membrane ion channels. Since muscarinic receptors are not themselves ion channels, the acute effects of muscarinic receptor stimulation on neuronal function are governed by the effects of the receptors on these endogenous neuronal ion channels. This review considers some principles and factors determining the interaction between subtypes and classes of muscarinic receptors with neuronal ion channels, and summarizes the effects of muscarinic receptor stimulation on a number of different channels, the mechanisms of receptor – channel transduction and their direct consequences for neuronal activity. Ion channels considered include potassium channels (voltage-gated, inward rectifier and calcium activated), voltage-gated calcium channels, cation channels and chloride channels. Key words: Ion channels; neuronal excitation and inhibition; pre- and postsynaptic events; muscarinic receptor subtypes; G proteins; transduction mechanisms. Contents. 1. Introduction: some principles of muscarinic receptor – ion channel coupling. 1.2 Some consequences of the indirect link between receptor and ion channel. + The connection between M1Rs and the M-type K channel as a model system 1.2.1 Dynamics of the response 1.2.2 Sensitivity of the response to agonist stimulation 2. Some muscarinic receptor-modulated neural ion channels.