EBV)Episomes in Latently Ulfected B-Lymphocytes by Mediating Their
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Identification of Candidate Biomarkers and Pathways Associated with Type 1 Diabetes Mellitus Using Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2021.06.08.447531; this version posted June 9, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Identification of candidate biomarkers and pathways associated with type 1 diabetes mellitus using bioinformatics analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karnataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2021.06.08.447531; this version posted June 9, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Type 1 diabetes mellitus (T1DM) is a metabolic disorder for which the underlying molecular mechanisms remain largely unclear. This investigation aimed to elucidate essential candidate genes and pathways in T1DM by integrated bioinformatics analysis. In this study, differentially expressed genes (DEGs) were analyzed using DESeq2 of R package from GSE162689 of the Gene Expression Omnibus (GEO). Gene ontology (GO) enrichment analysis, REACTOME pathway enrichment analysis, and construction and analysis of protein-protein interaction (PPI) network, modules, miRNA-hub gene regulatory network and TF-hub gene regulatory network, and validation of hub genes were then performed. A total of 952 DEGs (477 up regulated and 475 down regulated genes) were identified in T1DM. GO and REACTOME enrichment result results showed that DEGs mainly enriched in multicellular organism development, detection of stimulus, diseases of signal transduction by growth factor receptors and second messengers, and olfactory signaling pathway. -

PCR Analysis of Androgen-Regulated Genes in Human Lncap Prostate Cancer Cells
Oncogene (2009) 28, 2051–2063 & 2009 Macmillan Publishers Limited All rights reserved 0950-9232/09 $32.00 www.nature.com/onc ONCOGENOMICS Microarray coupled to quantitative RT–PCR analysis of androgen-regulated genes in human LNCaP prostate cancer cells S Ngan, EA Stronach, A Photiou, J Waxman, S Ali and L Buluwela Department of Oncology, Imperial College London, London, UK The androgen receptor (AR) mediates the growth- 2006). The importance of AR in male development is stimulatory effects of androgens in prostate cancer cells. shown by the androgen insensitivity syndromes char- Identification of androgen-regulated genes in prostate acterized by mutations in the AR gene (Gottlieb et al., cancer cells is therefore of considerable importance for 2004). The prostate is a prototypical androgen-depen- defining the mechanisms of prostate-cancer development dent organ (Cunha et al., 1987; Davies and Eaton, 1991) and progression. Although several studies have used and prostate cancer, which has become the most microarrays to identify AR-regulated genes in prostate commonly diagnosed cancer in males in the western cancer cell lines and in prostate tumours, we present here world and is the second leading cause of male cancer the results of gene expression microarray profiling of the death, is androgen-dependent for its growth (Carter and androgen-responsive LNCaP prostate-cancer cell line Coffey, 1990; McConnell, 1991). Therefore, treatment is treated withR1881 for theidentification of androgen- directed at inhibiting prostate cancer growth by regulated genes. We show that the expression of 319 genes suppressing the action of the endogenous androgen or is stimulated by 24 hafter R1881 addition, witha similar its production. -

Evidence for a Gene Influencing Haematocrit on Chromosome 6Q23–24: Genomewide Scan in the Framingham Heart Study J-P Lin, C J O’Donnell, D Levy, L a Cupples
75 LETTER TO JMG J Med Genet: first published as 10.1136/jmg.2004.021097 on 5 January 2005. Downloaded from Evidence for a gene influencing haematocrit on chromosome 6q23–24: genomewide scan in the Framingham Heart Study J-P Lin, C J O’Donnell, D Levy, L A Cupples ............................................................................................................................... J Med Genet 2005;42:75–79. doi: 10.1136/jmg.2004.021097 or more than 40 years, a number of studies have revealed that high haematocrit (HCT) levels are associated with Key points increased risk for cerebrovascular disease,1–3 cardiovas- F 4–6 7–9 cular disease (CVD), peripheral vascular disease, and all N Elevated haematocrit (HCT) levels are associated with cause mortality.5 10–11 During 34 years of follow up in more increased risk for vascular diseases. We carried out than 5200 individuals, a Framingham investigation demon- genome scans for quantitative trait loci (QTL) on HCT strated that increased HCT was significantly associated with and on haemoglobin (HGB), a correlated trait. increased risks for CVD, coronary heart disease, and N The heritabilities were estimated as 41% for HCT and 5 myocardial infarction in both men and women. A significant 45% for HGB. increase in all cause mortality for individuals with very low or N The genomewide linkage analysis revealed evidence of high HCT was also observed.5 Although HCT levels were related to other vascular risk factors, the risk associated with significant linkage for HCT to chromosome 6q23-24, an elevated HCT persisted after accounting for other risk with a lod score of 3.4 at location 136 cM. -

Prediction of Human Disease Genes by Human-Mouse Conserved Coexpression Analysis
Prediction of Human Disease Genes by Human-Mouse Conserved Coexpression Analysis Ugo Ala1., Rosario Michael Piro1., Elena Grassi1, Christian Damasco1, Lorenzo Silengo1, Martin Oti2, Paolo Provero1*, Ferdinando Di Cunto1* 1 Molecular Biotechnology Center, Department of Genetics, Biology and Biochemistry, University of Turin, Turin, Italy, 2 Department of Human Genetics and Centre for Molecular and Biomolecular Informatics, University Medical Centre Nijmegen, Nijmegen, The Netherlands Abstract Background: Even in the post-genomic era, the identification of candidate genes within loci associated with human genetic diseases is a very demanding task, because the critical region may typically contain hundreds of positional candidates. Since genes implicated in similar phenotypes tend to share very similar expression profiles, high throughput gene expression data may represent a very important resource to identify the best candidates for sequencing. However, so far, gene coexpression has not been used very successfully to prioritize positional candidates. Methodology/Principal Findings: We show that it is possible to reliably identify disease-relevant relationships among genes from massive microarray datasets by concentrating only on genes sharing similar expression profiles in both human and mouse. Moreover, we show systematically that the integration of human-mouse conserved coexpression with a phenotype similarity map allows the efficient identification of disease genes in large genomic regions. Finally, using this approach on 850 OMIM loci characterized by an unknown molecular basis, we propose high-probability candidates for 81 genetic diseases. Conclusion: Our results demonstrate that conserved coexpression, even at the human-mouse phylogenetic distance, represents a very strong criterion to predict disease-relevant relationships among human genes. Citation: Ala U, Piro RM, Grassi E, Damasco C, Silengo L, et al. -

Product Data Sheet
For research purposes only, not for human use Product Data Sheet HEBP2 siRNA (Mouse) Catalog # Source Reactivity Applications CRM5616 Synthetic M RNAi Description siRNA to inhibit HEBP2 expression using RNA interference Specificity HEBP2 siRNA (Mouse) is a target-specific 19-23 nt siRNA oligo duplexes designed to knock down gene expression. Form Lyophilized powder Gene Symbol HEBP2 Alternative Names SOUL; Heme-binding protein 2; Protein SOUL Entrez Gene 56016 (Mouse) SwissProt Q9WU63 (Mouse) Purity > 97% Quality Control Oligonucleotide synthesis is monitored base by base through trityl analysis to ensure appropriate coupling efficiency. The oligo is subsequently purified by affinity-solid phase extraction. The annealed RNA duplex is further analyzed by mass spectrometry to verify the exact composition of the duplex. Each lot is compared to the previous lot by mass spectrometry to ensure maximum lot-to-lot consistency. Components We offers pre-designed sets of 3 different target-specific siRNA oligo duplexes of mouse HEBP2 gene. Each vial contains 5 nmol of lyophilized siRNA. The duplexes can be transfected individually or pooled together to achieve knockdown of the target gene, which is most commonly assessed by qPCR or western blot. Our siRNA oligos are also chemically modified (2’-OMe) at no extra charge for increased stability and enhanced knockdown in vitro and in vivo. Directions for Use We recommends transfection with 100 nM siRNA 48 to 72 hours prior to cell lysis. Application key: E- ELISA, WB- Western blot, IH- Immunohistochemistry, -

Full-Text.Pdf
Systematic Evaluation of Genes and Genetic Variants Associated with Type 1 Diabetes Susceptibility This information is current as Ramesh Ram, Munish Mehta, Quang T. Nguyen, Irma of September 23, 2021. Larma, Bernhard O. Boehm, Flemming Pociot, Patrick Concannon and Grant Morahan J Immunol 2016; 196:3043-3053; Prepublished online 24 February 2016; doi: 10.4049/jimmunol.1502056 Downloaded from http://www.jimmunol.org/content/196/7/3043 Supplementary http://www.jimmunol.org/content/suppl/2016/02/19/jimmunol.150205 Material 6.DCSupplemental http://www.jimmunol.org/ References This article cites 44 articles, 5 of which you can access for free at: http://www.jimmunol.org/content/196/7/3043.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 23, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Systematic Evaluation of Genes and Genetic Variants Associated with Type 1 Diabetes Susceptibility Ramesh Ram,*,† Munish Mehta,*,† Quang T. -

Glutamine-Specific N-Terminal Amidase, a Component of the N- End Rule Pathway
NIH Public Access Author Manuscript Mol Cell. Author manuscript; available in PMC 2010 June 26. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Mol Cell. 2009 June 26; 34(6): 686±695. doi:10.1016/j.molcel.2009.04.032. Glutamine-Specific N-Terminal Amidase, a Component of the N- End Rule Pathway Haiqing Wang1,2,3, Konstantin I. Piatkov1,2, Christopher S. Brower1, and Alexander Varshavsky1,* 1Division of Biology, California Institute of Technology, Pasadena, CA 91125, USA SUMMARY Deamidation of N-terminal Gln by NtQ-amidase, a previously undescribed N-terminal amidohydrolase, is a part of the N-end rule pathway of protein degradation. We detected the activity of NtQ-amidase, termed Ntaq1, in mouse tissues, purified Ntaq1 from bovine brains, identified its gene, and began analyzing this enzyme. Ntaq1 is highly conserved among animals, plants and some fungi, but its sequence is dissimilar to sequences of other amidases. An earlier mutant in the Drosophila Cg8253 gene that we show here to encode NtQ-amidase has defective long-term memory. Other studies identified protein ligands of the uncharacterized human C8orf32 protein that we show here to be the Ntaq1 NtQ-amidase. Remarkably, “high-throughput” studies have recently solved the crystal structure of C8orf32 (Ntaq1). Our site-directed mutagenesis of Ntaq1 and its crystal structure indicate that the active site and catalytic mechanism of NtQ-amidase are similar to those of transglutaminases. Keywords amidohydrolase; proteolysis; ubiquitin; arginylation; tungus INTRODUCTION The N-end rule relates the in vivo half-life of a protein to the identity of its N-terminal residue (reviewed by (Mogk et al., 2007; Tasaki and Kwon, 2007; Varshavsky, 1996, 2008a; Varshavsky, 2008b). -

Genome-Wide Association Study (GWAS)
Linkage disequilibrium based eQTL analysis and comparative evolutionary epigenetic regulation of gene transcription by NING JIANG A thesis submitted to The University of Birmingham for the degree of DOCTOR OF PHILOSOPHY School of Biosciences The University of Birmingham November 2012 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. ABSTRACT The present thesis contains two independent parts of research, which are summarised as below. Part I Genome-Wide Association Study (GWAS) has recently been proposed as a powerful strategy for detecting the many subtle genetic variants that underlie phenotypic variation of complex polygenic traits in population-based samples. One of the main obstacles to successfully using the linkage disequilibrium based methods is knowledge of any underlying population structure. The presence of subgroups within a population can result in spurious association. A robust statistical method is developed to remove the population structure interference in GWAS by incorporating single control marker into testing for significance of genetic association of a polymorphic marker (SNP) with phenotypic variance of a complex trait. The novel approach avoids the need of structure prediction which could be infeasible or inadequate in practice and accounts properly for a varying effect of population stratification on different regions of the genome under study. -

A Grainyhead-Like 2/Ovo-Like 2 Pathway Regulates Renal Epithelial Barrier Function and Lumen Expansion
BASIC RESEARCH www.jasn.org A Grainyhead-Like 2/Ovo-Like 2 Pathway Regulates Renal Epithelial Barrier Function and Lumen Expansion † ‡ | Annekatrin Aue,* Christian Hinze,* Katharina Walentin,* Janett Ruffert,* Yesim Yurtdas,*§ | Max Werth,* Wei Chen,* Anja Rabien,§ Ergin Kilic,¶ Jörg-Dieter Schulzke,** †‡ Michael Schumann,** and Kai M. Schmidt-Ott* *Max Delbrueck Center for Molecular Medicine, Berlin, Germany; †Experimental and Clinical Research Center, and Departments of ‡Nephrology, §Urology, ¶Pathology, and **Gastroenterology, Charité Medical University, Berlin, Germany; and |Berlin Institute of Urologic Research, Berlin, Germany ABSTRACT Grainyhead transcription factors control epithelial barriers, tissue morphogenesis, and differentiation, but their role in the kidney is poorly understood. Here, we report that nephric duct, ureteric bud, and collecting duct epithelia express high levels of grainyhead-like homolog 2 (Grhl2) and that nephric duct lumen expansion is defective in Grhl2-deficient mice. In collecting duct epithelial cells, Grhl2 inactivation impaired epithelial barrier formation and inhibited lumen expansion. Molecular analyses showed that GRHL2 acts as a transcrip- tional activator and strongly associates with histone H3 lysine 4 trimethylation. Integrating genome-wide GRHL2 binding as well as H3 lysine 4 trimethylation chromatin immunoprecipitation sequencing and gene expression data allowed us to derive a high-confidence GRHL2 target set. GRHL2 transactivated a group of genes including Ovol2, encoding the ovo-like 2 zinc finger transcription factor, as well as E-cadherin, claudin 4 (Cldn4), and the small GTPase Rab25. Ovol2 induction alone was sufficient to bypass the requirement of Grhl2 for E-cadherin, Cldn4,andRab25 expression. Re-expression of either Ovol2 or a combination of Cldn4 and Rab25 was sufficient to rescue lumen expansion and barrier formation in Grhl2-deficient collecting duct cells. -

393LN V 393P 344SQ V 393P Probe Set Entrez Gene
393LN v 393P 344SQ v 393P Entrez fold fold probe set Gene Gene Symbol Gene cluster Gene Title p-value change p-value change chemokine (C-C motif) ligand 21b /// chemokine (C-C motif) ligand 21a /// chemokine (C-C motif) ligand 21c 1419426_s_at 18829 /// Ccl21b /// Ccl2 1 - up 393 LN only (leucine) 0.0047 9.199837 0.45212 6.847887 nuclear factor of activated T-cells, cytoplasmic, calcineurin- 1447085_s_at 18018 Nfatc1 1 - up 393 LN only dependent 1 0.009048 12.065 0.13718 4.81 RIKEN cDNA 1453647_at 78668 9530059J11Rik1 - up 393 LN only 9530059J11 gene 0.002208 5.482897 0.27642 3.45171 transient receptor potential cation channel, subfamily 1457164_at 277328 Trpa1 1 - up 393 LN only A, member 1 0.000111 9.180344 0.01771 3.048114 regulating synaptic membrane 1422809_at 116838 Rims2 1 - up 393 LN only exocytosis 2 0.001891 8.560424 0.13159 2.980501 glial cell line derived neurotrophic factor family receptor alpha 1433716_x_at 14586 Gfra2 1 - up 393 LN only 2 0.006868 30.88736 0.01066 2.811211 1446936_at --- --- 1 - up 393 LN only --- 0.007695 6.373955 0.11733 2.480287 zinc finger protein 1438742_at 320683 Zfp629 1 - up 393 LN only 629 0.002644 5.231855 0.38124 2.377016 phospholipase A2, 1426019_at 18786 Plaa 1 - up 393 LN only activating protein 0.008657 6.2364 0.12336 2.262117 1445314_at 14009 Etv1 1 - up 393 LN only ets variant gene 1 0.007224 3.643646 0.36434 2.01989 ciliary rootlet coiled- 1427338_at 230872 Crocc 1 - up 393 LN only coil, rootletin 0.002482 7.783242 0.49977 1.794171 expressed sequence 1436585_at 99463 BB182297 1 - up 393 -

The Pdx1 Bound Swi/Snf Chromatin Remodeling Complex Regulates Pancreatic Progenitor Cell Proliferation and Mature Islet Β Cell
Page 1 of 125 Diabetes The Pdx1 bound Swi/Snf chromatin remodeling complex regulates pancreatic progenitor cell proliferation and mature islet β cell function Jason M. Spaeth1,2, Jin-Hua Liu1, Daniel Peters3, Min Guo1, Anna B. Osipovich1, Fardin Mohammadi3, Nilotpal Roy4, Anil Bhushan4, Mark A. Magnuson1, Matthias Hebrok4, Christopher V. E. Wright3, Roland Stein1,5 1 Department of Molecular Physiology and Biophysics, Vanderbilt University, Nashville, TN 2 Present address: Department of Pediatrics, Indiana University School of Medicine, Indianapolis, IN 3 Department of Cell and Developmental Biology, Vanderbilt University, Nashville, TN 4 Diabetes Center, Department of Medicine, UCSF, San Francisco, California 5 Corresponding author: [email protected]; (615)322-7026 1 Diabetes Publish Ahead of Print, published online June 14, 2019 Diabetes Page 2 of 125 Abstract Transcription factors positively and/or negatively impact gene expression by recruiting coregulatory factors, which interact through protein-protein binding. Here we demonstrate that mouse pancreas size and islet β cell function are controlled by the ATP-dependent Swi/Snf chromatin remodeling coregulatory complex that physically associates with Pdx1, a diabetes- linked transcription factor essential to pancreatic morphogenesis and adult islet-cell function and maintenance. Early embryonic deletion of just the Swi/Snf Brg1 ATPase subunit reduced multipotent pancreatic progenitor cell proliferation and resulted in pancreas hypoplasia. In contrast, removal of both Swi/Snf ATPase subunits, Brg1 and Brm, was necessary to compromise adult islet β cell activity, which included whole animal glucose intolerance, hyperglycemia and impaired insulin secretion. Notably, lineage-tracing analysis revealed Swi/Snf-deficient β cells lost the ability to produce the mRNAs for insulin and other key metabolic genes without effecting the expression of many essential islet-enriched transcription factors. -
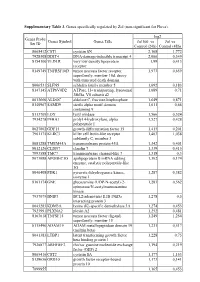
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis