Simple Quantitative Tests to Validate Sampling from Thermodynamic
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A High-Order Boris Integrator
A high-order Boris integrator Mathias Winkela,∗, Robert Speckb,a, Daniel Ruprechta aInstitute of Computational Science, University of Lugano, Switzerland. bJ¨ulichSupercomputing Centre, Forschungszentrum J¨ulich,Germany. Abstract This work introduces the high-order Boris-SDC method for integrating the equations of motion for electrically charged particles in an electric and magnetic field. Boris-SDC relies on a combination of the Boris-integrator with spectral deferred corrections (SDC). SDC can be considered as preconditioned Picard iteration to com- pute the stages of a collocation method. In this interpretation, inverting the preconditioner corresponds to a sweep with a low-order method. In Boris-SDC, the Boris method, a second-order Lorentz force integrator based on velocity-Verlet, is used as a sweeper/preconditioner. The presented method provides a generic way to extend the classical Boris integrator, which is widely used in essentially all particle-based plasma physics simulations involving magnetic fields, to a high-order method. Stability, convergence order and conserva- tion properties of the method are demonstrated for different simulation setups. Boris-SDC reproduces the expected high order of convergence for a single particle and for the center-of-mass of a particle cloud in a Penning trap and shows good long-term energy stability. Keywords: Boris integrator, time integration, magnetic field, high-order, spectral deferred corrections (SDC), collocation method 1. Introduction Often when modeling phenomena in plasma physics, for example particle dynamics in fusion vessels or particle accelerators, an externally applied magnetic field is vital to confine the particles in the physical device [1, 2]. In many cases, such as instabilities [3] and high-intensity laser plasma interaction [4], the magnetic field even governs the microscopic evolution and drives the phenomena to be studied. -

Vibration Odes 1
Vibration ODEs 1 Vibration problems lead to differential equations with solutions that oscillate in time, typically in a damped or undamped sinusoidal fashion. Such solutions put certain demands on the numerical methods compared to other phenomena whose solutions are monotone or very smooth. Both the frequency and amplitude of the oscillations need to be accurately handled by the numerical schemes. The forthcom- ing text presents a range of different methods, from classical ones (Runge-Kutta and midpoint/Crank-Nicolson methods), to more modern and popular symplectic (ge- ometric) integration schemes (Leapfrog, Euler-Cromer, and Störmer-Verlet meth- ods), but with a clear emphasis on the latter. Vibration problems occur throughout mechanics and physics, but the methods discussed in this text are also fundamen- tal for constructing successful algorithms for partial differential equations of wave nature in multiple spatial dimensions. 1.1 Finite Difference Discretization Many of the numerical challenges faced when computing oscillatory solutions to ODEs and PDEs can be captured by the very simple ODE u00 C u D 0. This ODE is thus chosen as our starting point for method development, implementation, and analysis. 1.1.1 A Basic Model for Vibrations The simplest model of a vibrating mechanical system has the following form: u00 C !2u D 0; u.0/ D I; u0.0/ D 0; t 2 .0; T : (1.1) Here, ! and I are given constants. Section 1.12.1 derives (1.1) from physical prin- ciples and explains what the constants mean. The exact solution of (1.1)is u.t/ D I cos.!t/ : (1.2) © The Author(s) 2017 1 H.P. -

A Position Based Approach to Ragdoll Simulation
LINKÖPING UNIVERSITY Department of Electrical Engineering Master Thesis A Position Based Approach to Ragdoll Simulation Fiammetta Pascucci LITH-ISY-EX- -07/3966- -SE June 2007 LINKÖPING UNIVERSITY Department of Electrical Engineering Master Thesis A position Based Approach to Ragdoll Simulation Fiammetta Pascucci LITH-ISY-EX- -07/3966- -SE June 2007 Supervisor: Ingemar Ragnemalm Examinator: Ingemar Ragnemalm Abstract Create the realistic motion of a character is a very complicated work. This thesis aims to create interactive animation for characters in three dimen- sions using position based approach. Our character is pictured from ragdoll, which is a structure of system particles where all particles are linked by equidistance con- straints. The goal of this thesis is observed the fall in the space of our ragdoll after creating all constraints, as structure, contact and environment constraints. The structure constraint represents all joint constraints which have one, two or three Degree of Freedom (DOF). The contact constraints are represented by collisions between our ragdoll and other objects in the space. Finally, the environment constraints are represented by means of the wall con- straint. The achieved results allow to have a realist fall of our ragdoll in the space. Keywords: Particle System, Verlet’s Integration, Skeleton Animation, Joint Con- straint, Environment Constraint, Skinning, Collision Detection. Acknowledgments I would to thanks all peoples who aid on making my master thesis work. In particular, thanks a lot to the person that most of all has believed in me. This person has always encouraged to go forward, has always given a lot love and help me in difficult movement. -

Exponential Integration for Hamiltonian Monte Carlo
Exponential Integration for Hamiltonian Monte Carlo Wei-Lun Chao1 [email protected] Justin Solomon2 [email protected] Dominik L. Michels2 [email protected] Fei Sha1 [email protected] 1Department of Computer Science, University of Southern California, Los Angeles, CA 90089 2Department of Computer Science, Stanford University, 353 Serra Mall, Stanford, California 94305 USA Abstract butions that generate random walks in sample space. A fraction of these walks aligned with the target distribution is We investigate numerical integration of ordinary kept, while the remainder is discarded. Sampling efficiency differential equations (ODEs) for Hamiltonian and quality depends critically on the proposal distribution. Monte Carlo (HMC). High-quality integration is For this reason, designing good proposals has long been a crucial for designing efficient and effective pro- subject of intensive research. posals for HMC. While the standard method is leapfrog (Stormer-Verlet)¨ integration, we propose For distributions over continuous variables, Hamiltonian the use of an exponential integrator, which is Monte Carlo (HMC) is a state-of-the-art sampler using clas- robust to stiff ODEs with highly-oscillatory com- sical mechanics to generate proposal distributions (Neal, ponents. This oscillation is difficult to reproduce 2011). The method starts by introducing auxiliary sam- using leapfrog integration, even with carefully pling variables. It then treats the negative log density of selected integration parameters and precondition- the variables’ joint distribution as the Hamiltonian of parti- ing. Concretely, we use a Gaussian distribution cles moving in sample space. Final positions on the motion approximation to segregate stiff components of trajectories are used as proposals. the ODE. -

Advanced Programming in Engineering Lecture Notes
Advanced Programming in Engineering lecture notes P.J. Visser, S. Luding, S. Srivastava March 13, 2011 Changes 13/3/2011 Added Section 7.4 to 7.7. Added Section 4.4 7/3/2011 Added chapter Chapter 7 23/2/2011 Corrected the equation for the average temperature in terms of micro- scopic quantities, Eq. 3.14. Corrected the equation for the speed distribution in 2D, Eq. 3.19. Added links to sources of `true' random numbers available through the web as footnotes in Section 5.1. Added examples of choices for parameters of the LCG and LFG in Section 5.1.1 and 5.1.2, respectively. Added the chi-square test and correlation functions as ways for testing random numbers in Section 5.1.4. Improved the derivation of the average squared distance of particles in a random walk being linear with time, in Section 5.2.1. Added a discussion of the meaning of the terms in the master equation, in Section 5.2.5. Small changes. 19/2/2011 Corrected the Maxwell-Boltzmann distribution for the speed, Eq. 3.19. 14/2/2011 1 2 Added Chapter 6. 6/2/2011 Verified Eq. 3.15 and added footnote discussing its meaning in case of periodic boundary conditions. Added formula for the radial distribution function in Section 3.4.2, Eq. 3.21. 31/1/2011 Added Chapter 5. 9/1/2010 Corrected labels on the axes in Fig. 3.10, changed U · " and r · σ to U=" and r/σ, respectively. Added truncated version of the Lennard-Jones potential in Section 3.3.1. -

Transport Properties of Fluids: Symplectic Integrators and Their
Fluid Phase Equilibria 183–184 (2001) 351–361 Transport properties of fluids: symplectic integrators and their usefulness J. Ratanapisit a,b, D.J. Isbister c, J.F. Ely b,∗ a Chemical Engineering and Petroleum Refining Department, Colorado School of Mines, Golden, CO 80401, USA b Department of Chemical Engineering, Prince of Songla University, Hat-Yai, Songkla 90110, Thailand c School of Physics, University College, University of New South Wales, ADFA, Canberra, ACT 2600, Australia Abstract An investigation has been carried out into the effectiveness of using symplectic/operator splitting generated algorithms for the evaluation of transport coefficients in Lennard–Jones fluids. Equilibrium molecular dynamics is used to revisit the Green–Kubo calculation of these transport coefficients through integration of the appropriate correlation functions. In particular, an extensive series of equilibrium molecular dynamic simulations have been per- formed to investigate the accuracy, stability and efficiency of second-order explicit symplectic integrators: position Verlet, velocity Verlet, and the McLauchlan–Atela algorithms. Comparisons are made to nonsymplectic integrators that include the fourth-order Runge–Kutta and fourth-order Gear predictor–corrector methods. These comparisons were performed based on several transport properties of Lennard–Jones fluids: self-diffusion, shear viscosity and thermal conductivity. Because transport properties involve long time simulations to obtain accurate evaluations of their numerical values, they provide an excellent basis to study the accuracy and stability of the SI methods. To our knowledge, previous studies on the SIs have only looked at the thermodynamic energy using a simple model fluid. This study presents realistic, but perhaps the simplest simulations possible to test the effect of the integrators on the three main transport properties. -

Verlet with Collisions for Mass Spring Model Simulations
Verlet with Collisions for Mass Spring Model Simulations Maciej Kot1 and Hiroshi Nagahashi2 1Imaging Science and Engineering Laboratory, Tokyo Institute of Technology, Tokyo, Japan 2Laboratory for Future Interdisciplinary Research of Science and Technology, Tokyo Institute of Technology, Tokyo, Japan Keywords: Mass Spring Models, Deformable Objects, Collisions, Verlet, Friction. Abstract: In this paper we study the problem of the interaction of soft bodies modeled with mass spring models (MSM) and static elements of the environment. We show that in such setup it is possible to couple standard time evolution of MSMs with collision responses in a way, that does not require complex processing for multi collision situations while successfully preventing object inter-penetration. Moreover we show how to achieve similar energy dissipation for models with different resolutions when the friction is present. 1 INTRODUCTION ulation and we show how to couple it with collision responses. One of our goals is to design a collision Mass spring models are a popular choice for the rep- handling technique which preserves the energy of the resentation of soft bodies in computer graphics and system (affects stability of the simulation), prevents virtual reality applications. For a long time they were object penetration, and if the friction is present, obeys preferred over the finite element method (FEM) in the Newton’s law of dry friction. We achieve this goal real time applications, as they tend to offer better completely for collisions between MSM and static el- computational efficiency (although recent FEM im- ements of the environment and partially for MSM- plementations are quite fast as well (Sin et al., 2013)), MSM collisions; in case of MSM-MSM collisions our but they are believed to be less accurate in terms of method does not preserve energy in certain situations, physical plausibility (Nealen et al., 2006). -

A Semi-Analytical Approach to Molecular Dynamics
Journal of Computational Physics 303 (2015) 336–354 Contents lists available at ScienceDirect Journal of Computational Physics www.elsevier.com/locate/jcp A semi-analytical approach to molecular dynamics a,b, c,d Dominik L. Michels ∗, Mathieu Desbrun a Computer Science Department, Stanford University, 353 Serra Mall, MC 9515, Stanford, CA 94305, USA b Max Planck Institute for Informatics, Campus E1 4, 66123 Saarbrücken, Germany c Department of Computing and Mathematical Sciences, California Institute of Technology, 1200 E. California Blvd., MC 305-16, Pasadena, CA 91125, USA d Centre de Recherche INRIA Sophia-Antipolis Méditerranée, 2004 Route des Lucioles, BP 93, 06902 Sophia Antipolis Cedex, France a r t i c l e i n f o a b s t r a c t Article history: Despite numerous computational advances over the last few decades, molecular dynamics Received 12 July 2014 still favors explicit (and thus easily-parallelizable) time integrators for large scale numerical Received in revised form 28 September simulation. As a consequence, computational efficiency in solving its typically stiff 2015 oscillatory equations of motion is hampered by stringent stability requirements on the Accepted 2 October 2015 time step size. In this paper, we present a semi-analytical integration scheme that offers Available online 13 October 2015 a total speedup of a factor 30 compared to the Verlet method on typical MD simulation Keywords: by allowing over three orders of magnitude larger step sizes. By efficiently approximating Energy conservation the exact integration of the strong (harmonic) forces of covalent bonds through matrix Explicit integration functions, far improved stability with respect to time step size is achieved without Exponential integrators sacrificing the explicit, symplectic, time-reversible, or fine-grained parallelizable nature of Fast Multipole Method the integration scheme. -

Molecular Dynamics Simulation
Molecular Dynamics Simulation Table of Contents Theory I: Solving Newtons Laws for collection of particles ........................................................ 1 Figure 1: Diagram defining particle-particle interaction variables. ................................................ 2 Simple Molecular Dynamics Simulation ................................................................................. 3 Experimental Parameters (particle) ........................................................................................ 3 Experimental Parameters (space-domain) ................................................................................ 3 Experimental Parameters (initial conditions) ............................................................................ 4 Experimental Parameters (time-domain) ................................................................................. 4 Display Parameters ............................................................................................................. 4 Theory II: Numeric integration of Newton's laws Part I ............................................................. 4 Figure 2: Euler vs. Semi-implicit Euler Integration. .................................................................. 5 Theory II: Numeric integration of Newton's laws Part II ............................................................ 6 Figure 3: Velocity Verlet vs. Semi-implicit Euler Integration. ..................................................... 7 Figure 3: Velocity Verlet vs. Semi-implicit Euler -

ANIMATION 4 : MODELES PHYSIQUES ET DYNAMIQUES Rémi Ronfard, Animation, M2R IICAO Plan Du Cours
ANIMATION 4 : MODELES PHYSIQUES ET DYNAMIQUES Rémi Ronfard, Animation, M2R IICAO Plan du cours Cours 1: Armatures (squelettes) et attachement a la surface du corps (skinning). TP : programmation d’un rig dans Blender Cours 2: Animation par keyframes + techniques d’interpolation TP : animation par keyframes dans Blender Cours 3: Cinématique inverse TP : animation d’une marche dans Blender Cours 4: Animation physique TP: animation d’un ragdoll dans Blender Summary of physics-based animation Motivation Newton’s Laws Point-mass models Rigid and articulated bodies Ragdolls Uncanny Valley Cartoon physics vs. Real world physics Any body suspended in space will remain in space until made aware of its situation. Any body in motion will tend to remain in motion until solid matter intervenes suddenly. Any body passing through solid matter will leave a perforation conforming to its perimeter. The time required for an object to fall twenty stories is greater than or equal to the time it takes for whoever knocked it off the ledge to spiral down twenty flights to attempt to capture it unbroken. Certain bodies can pass through solid walls painted to resemble tunnel entrances; others cannot. Motivation : useful motor skills Motivation : Animation physique dans Blender Game Engine : Sensors, Controllers and Actuators Bullet Physics Ragdolls De la cinématique à la dynamique Calcul des positions, vitesses, angles et vitesses angulaires en fonction du temps Intégration à partir des équations du mouvement Principaux concepts en dynamique Forces Centres de masse et inerties Newton’s Law and Euler integration Euler method, named after Leonhard Euler, is a first-order Start at x0 numerical procedure for solving ordinary differential Compute acceleration equations (ODEs) with a given from f = ma (Newton) initial value. -
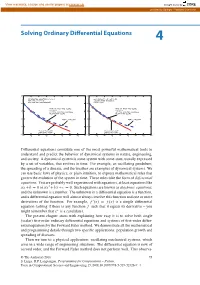
Solving Ordinary Differential Equations 4
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Springer - Publisher Connector Solving Ordinary Differential Equations 4 Differential equations constitute one of the most powerful mathematical tools to understand and predict the behavior of dynamical systems in nature, engineering, and society. A dynamical system is some system with some state, usually expressed by a set of variables, that evolves in time. For example, an oscillating pendulum, the spreading of a disease, and the weather are examples of dynamical systems. We can use basic laws of physics, or plain intuition, to express mathematical rules that govern the evolution of the system in time. These rules take the form of differential equations. You are probably well experienced with equations, at least equations like axCb D 0 or ax2 CbxCc D 0. Such equations are known as algebraic equations, and the unknown is a number. The unknown in a differential equation is a function, and a differential equation will almost always involve this function and one or more derivatives of the function. For example, f 0.x/ D f.x/ is a simple differential equation (asking if there is any function f such that it equals its derivative – you might remember that ex is a candidate). The present chapter starts with explaining how easy it is to solve both single (scalar) first-order ordinary differential equations and systems of first-order differ- ential equations by the Forward Euler method. We demonstrate all the mathematical and programming details through two specific applications: population growth and spreading of diseases. -

Simulation Methods in Physics 1
...script in development... Simulation Methods in Physics 1 Prof. Dr. Christian Holm Institute for Computational Physics University of Stuttgart WS 2012/2013 Contents 1 Introduction 3 1.1 Computer Science . 3 1.1.1 Historical developments . 3 1.1.2 TOP 500 Supercomputer list . 4 1.1.3 Moore's law (1970) . 5 1.2 Computer Simulations in Physics . 6 1.2.1 The New Trinity of Physics . 6 1.2.2 Optimization of simulation performance . 8 1.2.3 Optimization of simulation algorithms . 8 1.2.4 Computational approaches for different length scales . 9 2 Molecular Dynamics integrators 11 2.1 Principles of Molecular Dynamics . 11 2.1.1 Ergodicity . 11 2.2 Integration of equations of motion . 12 2.2.1 Algorithms . 12 2.2.2 Estimation of time step . 13 2.2.3 Verlet Algorithm . 13 2.2.4 Velocity Verlet Algorithm . 14 2.2.5 Leapfrog method . 15 2.2.6 Runge-Kutta method . 16 2.2.7 Predictor-Corrector methods . 17 2.2.8 Ljapunov-Characteristics . 17 3 Statistical mechanics - Quick & Dirty 20 3.1 Microstates . 20 3.1.1 Distribution functions . 20 3.1.2 Average values and thermodynamic limit . 22 3.2 Definition of Entropy and Temperature . 23 3.3 Boltzmann distribution . 24 3.3.1 Total energy and free energy . 25 3.4 Quantum mechanics and classcal partition functions . 26 3.5 Ensemble averages . 26 2 1 Introduction 1.1 Computer Science 1.1.1 Historical developments • The first calculating tool for performing arithmetic processes, the Abacus is used ∼ 1000 b.C.