New Developments in Epigenetics and Potential Clinical Applications
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
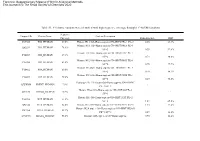
Table S1. 49 Histone Variants Were Identified with High Sequence Coverage Through LC-MS/MS Analysis Electronic Supplementary
Electronic Supplementary Material (ESI) for Analytical Methods. This journal is © The Royal Society of Chemistry 2020 Table S1. 49 histone variants were identified with high sequence coverage through LC-MS/MS analysis Sequence Uniprot IDs Protein Name Protein Description Coverage Ratio E2+/E2- RSD P07305 H10_HUMAN 67.5% Histone H1.0 OS=Homo sapiens GN=H1F0 PE=1 SV=3 4.85 23.3% Histone H1.1 OS=Homo sapiens GN=HIST1H1A PE=1 Q02539 H11_HUMAN 74.4% SV=3 0.35 92.6% Histone H1.2 OS=Homo sapiens GN=HIST1H1C PE=1 P16403 H12_HUMAN 67.1% SV=2 0.73 80.6% Histone H1.3 OS=Homo sapiens GN=HIST1H1D PE=1 P16402 H13_HUMAN 63.8% SV=2 0.75 77.7% Histone H1.4 OS=Homo sapiens GN=HIST1H1E PE=1 P10412 H14_HUMAN 69.0% SV=2 0.70 80.3% Histone H1.5 OS=Homo sapiens GN=HIST1H1B PE=1 P16401 H15_HUMAN 79.6% SV=3 0.29 98.3% Testis-specific H1 histone OS=Homo sapiens GN=H1FNT Q75WM6 H1FNT_HUMAN 7.8% \ \ PE=2 SV=3 Histone H1oo OS=Homo sapiens GN=H1FOO PE=2 Q8IZA3 H1FOO_HUMAN 5.2% \ \ SV=1 Histone H1t OS=Homo sapiens GN=HIST1H1T PE=2 P22492 H1T_HUMAN 31.4% SV=4 1.42 65.0% Q92522 H1X_HUMAN 82.6% Histone H1x OS=Homo sapiens GN=H1FX PE=1 SV=1 1.15 33.2% Histone H2A type 1 OS=Homo sapiens GN=HIST1H2AG P0C0S8 H2A1_HUMAN 99.2% PE=1 SV=2 0.57 26.8% Q96QV6 H2A1A_HUMAN 58.0% Histone H2A type 1-A OS=Homo sapiens 0.90 11.2% GN=HIST1H2AA PE=1 SV=3 Histone H2A type 1-B/E OS=Homo sapiens P04908 H2A1B_HUMAN 99.2% GN=HIST1H2AB PE=1 SV=2 0.92 30.2% Histone H2A type 1-C OS=Homo sapiens Q93077 H2A1C_HUMAN 100.0% GN=HIST1H2AC PE=1 SV=3 0.76 27.6% Histone H2A type 1-D OS=Homo sapiens P20671 -

UNIVERSITY of CALIFORNIA, SAN DIEGO Functional Analysis of Sall4
UNIVERSITY OF CALIFORNIA, SAN DIEGO Functional analysis of Sall4 in modulating embryonic stem cell fate A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Molecular Pathology by Pei Jen A. Lee Committee in charge: Professor Steven Briggs, Chair Professor Geoff Rosenfeld, Co-Chair Professor Alexander Hoffmann Professor Randall Johnson Professor Mark Mercola 2009 Copyright Pei Jen A. Lee, 2009 All rights reserved. The dissertation of Pei Jen A. Lee is approved, and it is acceptable in quality and form for publication on microfilm and electronically: ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ Co-Chair ______________________________________________________________ Chair University of California, San Diego 2009 iii Dedicated to my parents, my brother ,and my husband for their love and support iv Table of Contents Signature Page……………………………………………………………………….…iii Dedication…...…………………………………………………………………………..iv Table of Contents……………………………………………………………………….v List of Figures…………………………………………………………………………...vi List of Tables………………………………………………….………………………...ix Curriculum vitae…………………………………………………………………………x Acknowledgement………………………………………………….……….……..…...xi Abstract………………………………………………………………..…………….....xiii Chapter 1 Introduction ..…………………………………………………………………………….1 Chapter 2 Materials and Methods……………………………………………………………..…12 -

Quantitative Proteomics Methods for the Analysis of Histone Post-Translational Modifications
Université de Montréal Quantitative Proteomics Methods for the Analysis of Histone Post-translational Modifications par Nebiyu Ali Abshiru Département de Chimie Faculté des arts et des sciences Thèse présentée à la Faculté des études supérieures et postdoctorales en vue de l’obtention du grade de philosophiae doctor (Ph.D.) en chimie Septembre 2015 ©Nebiyu Ali Abshiru, 2015 i Résumé Les histones sont des protéines nucléaires hautement conservées chez les cellules des eucaryotes. Elles permettent d’organiser et de compacter l’ADN sous la forme de nucléosomes, ceux-ci representant les sous unités de base de la chromatine. Les histones peuvent être modifiées par de nombreuses modifications post-traductionnelles (PTMs) telles que l’acétylation, la méthylation et la phosphorylation. Ces modifications jouent un rôle essentiel dans la réplication de l’ADN, la transcription et l’assemblage de la chromatine. L’abondance de ces modifications peut varier de facon significative lors du developpement des maladies incluant plusieurs types de cancer. Par exemple, la perte totale de la triméthylation sur H4K20 ainsi que l’acétylation sur H4K16 sont des marqueurs tumoraux spécifiques a certains types de cancer chez l’humain. Par conséquent, l’étude de ces modifications et des événements determinant la dynamique des leurs changements d’abondance sont des atouts importants pour mieux comprendre les fonctions cellulaires et moléculaires lors du développement de la maladie. De manière générale, les modifications des histones sont étudiées par des approches biochimiques telles que les immuno-buvardage de type Western ou les méthodes d’immunoprécipitation de la chromatine (ChIP). Cependant, ces approches présentent plusieurs inconvénients telles que le manque de spécificité ou la disponibilité des anticorps, leur coût ou encore la difficulté de les produire et de les valider. -

SETD2 Mutations in Primary Central Nervous System Tumors Angela N
Viaene et al. Acta Neuropathologica Communications (2018) 6:123 https://doi.org/10.1186/s40478-018-0623-0 RESEARCH Open Access SETD2 mutations in primary central nervous system tumors Angela N. Viaene1, Mariarita Santi1, Jason Rosenbaum2, Marilyn M. Li1, Lea F. Surrey1 and MacLean P. Nasrallah2,3* Abstract Mutations in SETD2 are found in many tumors, including central nervous system (CNS) tumors. Previous work has shown these mutations occur specifically in high grade gliomas of the cerebral hemispheres in pediatric and young adult patients. We investigated SETD2 mutations in a cohort of approximately 640 CNS tumors via next generation sequencing; 23 mutations were detected across 19 primary CNS tumors. Mutations were found in a wide variety of tumors and locations at a broad range of allele frequencies. SETD2 mutations were seen in both low and high grade gliomas as well as non-glial tumors, and occurred in patients greater than 55 years of age, in addition to pediatric and young adult patients. High grade gliomas at first occurrence demonstrated either frameshift/truncating mutations or point mutations at high allele frequencies, whereas recurrent high grade gliomas frequently harbored subclones with point mutations in SETD2 at lower allele frequencies in the setting of higher mutational burdens. Comparison with the TCGA dataset demonstrated consistent findings. Finally, immunohistochemistry showed decreased staining for H3K36me3 in our cohort of SETD2 mutant tumors compared to wildtype controls. Our data further describe the spectrum of tumors in which SETD2 mutations are found and provide a context for interpretation of these mutations in the clinical setting. Keywords: SETD2, Histone, Brain tumor, Glioma, Epigenetics, H3K36me3 Introduction (H3K36me3) in humans. -

Active Motif Technical Data Sheet (TDS)
Histone H4K8ac antibody (pAb) Catalog Nos: 61103, 61104 Quantities: 100 µg, 10 µg RRID: AB_2793506 Purification: Protein A Chromatography Isotype: IgG Host: Rabbit Application(s): ChIP, ChIP-Seq, DB, ICC, IF, WB Concentration: 1 µg/µl Reactivity: Human, Mouse, Wide Range Predicted Molecular Weight: 8 kDa Background: Histone H4 is one of the core components of the nucleosome. The nucleosome is the smallest subunit of chromatin and consists of 147 base pairs of DNA wrapped around an octamer of core histone proteins (two each of Histone H2A, Histone H2B, Histone H3 and Histone H4). Histone H1 is a linker histone, present at the interface between the nucleosome core and DNA entry/exit points; it is responsible for establishing higher-order chromatin structure. Chromatin is subject to a variety of chemical modifications, including post-translational modifications of the histone proteins and the methylation of cytosine residues in the DNA. Reported histone modifications include acetylation, methylation, phosphorylation, ubiquitylation, glycosylation, ADP-ribosylation, carbonylation and SUMOylation; they play a major role in regulating gene expression. Lysine N-ε-acetylation is a dynamic, reversible and tightly regulated protein and histone modification that plays a major role in chromatin remodeling and in the regulation of gene expression in various cellular functions. The chromatin-remodeling complex SWI/SNF is recruited to promoters through the interaction of the bromodomain of the protein BRG1, belonging to the SWI/SNF complex, and CBP-acetylated histone H4 Lysine 8, leading to a chromatin remodeling. Immunogen: This Histone H4 acetyl Lys8 antibody was raised against a peptide containing acetyl Lys8 of human Histone H4. -

Active Motif Technical Data Sheet (TDS)
Histone H4K8ac antibody (mAb) Catalog No: 61525 Quantity: 100 µg RRID: AB_2793669 Purification: Protein G Chromatography Clone: MABI 0408 Host: Mouse Isotype: IgG1 Concentration: 0.66 µg/µl Application(s): ChIP, DB, WB Molecular Weight: 8 kDa Reactivity: Human, Wide Range Predicted Background: Histone H4 is one of the core components of the nucleosome. The nucleosome is the smallest subunit of chromatin and consists of 147 base pairs of DNA wrapped around an octamer of core histone proteins (two each of Histone H2A, Histone H2B, Histone H3 and Histone H4). Histone H1 is a linker histone, present at the interface between the nucleosome core and DNA entry/exit points; it is responsible for establishing higher-order chromatin structure. Chromatin is subject to a variety of chemical modifications, including post-translational modifications of the histone proteins and the methylation of cytosine residues in the DNA. Reported histone modifications include acetylation, methylation, phosphorylation, ubiquitylation, glycosylation, ADP-ribosylation, carbonylation and SUMOylation; they play a major role in regulating gene expression. Lysine N-ε-acetylation is a dynamic, reversible and tightly regulated protein and histone modification that plays a major role in chromatin remodeling and in the regulation of gene expression in various cellular functions. The chromatin-remodeling complex SWI/SNF is recruited to promoters through the interaction of the bromodomain of the protein BRG1, belonging to the SWI/SNF complex, and CBP-acetylated histone H4 Lysine 8, leading to a chromatin remodeling. Immunogen: This antibody was raised against a synthetic peptide containing acetyl-lysine 8 of human Histone H4. Buffer: Purified IgG in PBS with 30% glycerol and 0.035% sodium azide. -

EIN2 Mediates Direct Regulation of Histone Acetylation in the Ethylene Response
EIN2 mediates direct regulation of histone acetylation in the ethylene response Fan Zhanga,b,1, Likai Wanga,b,1, Bin Qic, Bo Zhaoa,b, Eun Esther Koa,b, Nathaniel D. Riggana,b, Kevin China,b, and Hong Qiaoa,b,2 aInstitute for Cellular and Molecular Biology, The University of Texas at Austin, Austin, TX 78712; bDepartment of Molecular Biosciences, The University of Texas at Austin, Austin, TX 78712; and cDepartment of Molecular, Cellular, and Developmental Biology, University of Colorado Boulder, Boulder, CO 80309 Edited by Steven E. Jacobsen, University of California, Los Angeles, CA, and approved August 10, 2017 (received for review May 15, 2017) Ethylene gas is essential for developmental processes and stress of EBF1 and EBF2 (19, 20). In the nucleus, the EIN2-C transduces responses in plants. Although the membrane-bound protein EIN2 is signals to the transcription factors EIN3 and EIL1, which are key for critical for ethylene signaling, the mechanism by which the ethylene activation of expression of all ethylene-response genes (21, 22). We signal is transduced remains largely unknown. Here we show the recently discovered that acetylation at H3K23 and H3K14Ac is in- levels of H3K14Ac and H3K23Ac are correlated with the levels of volved in ethylene-regulated gene activation in a manner that de- EIN2 protein and demonstrate EIN2 C terminus (EIN2-C) is sufficient to pends on both EIN2 and EIN3 (23, 24). rescue the levels of H3K14/23Ac of ein2-5 at the target loci, using Here we show that the levels of H3K14/23Ac are positively CRISPR/dCas9-EIN2-C. -

Glycolytic Metabolism Influences Global Chromatin Structure
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 6, No.6 Glycolytic metabolism influences global chromatin structure Xue-Song Liu, John B. Little, Zhi-Min Yuan Department of Genetics and Complex Diseases, Harvard School of Public Health, Boston, MA 02115, USA Correspondence to: Zhi-Min Yuan, e-mail: [email protected] Keywords: glycolysis, acetylation, chromatin structure, chemosensitivity Received: September 29, 2014 Accepted: December 15, 2014 Published: January 13, 2015 ABSTRACT Metabolic rewiring, specifically elevated glycolytic metabolism is a hallmark of cancer. Global chromatin structure regulates gene expression, DNA repair, and also affects cancer progression. But the interrelationship between tumor metabolism and chromatin architecture remain unclear. Here we show that increased glycolysis in cancer cells promotes an open chromatin configuration. Using complementary methods including Micrococcal nuclease (MNase) digestion assay, electron microscope and immunofluorescence staining, we demonstrate that glycolysis inhibition by pharmacological and genetic approaches was associated with induction of compacted chromatin structure. This condensed chromatin status appeared to result chiefly from histone hypoacetylation as restoration of histone acetylation with an HDAC inhibitor reversed the compacted chromatin state. Interestingly, glycolysis inhibition-induced chromatin condensation impeded DNA repair efficiency leading to increased sensitivity of cancer cells to DNA damage drugs, which may represent a novel molecular mechanism that can be exploited for cancer therapy. INTRODUCTION Tumor cells metabolize most glucose into lactate and thus generate abundant glycolytic intermediates as Pathologist have observed for long time that the precursors for macromolecular biosynthesis, which nucleus of cancer cells show distinct morphological enables tumor cells to meet their increased anabolic and alterations compared to the nucleus of normal cells, and energetic demands due to rapid tumor growth [8]. -

H4k8ac Antibody
08/14 For research use only BioVision H4K8ac Antibody ALTERNATE NAMES: Histone H4 CATALOG #: 6807-50 AMOUNT: 50 µl HOST/ISOTYPE: Rabbit To determine the titer, an ELISA was performed using a IMMUNOGEN: KLH-conjugated synthetic peptide of Histone H4 containing the HeLa cells (15 µg) were analysed by serial dilution of the antibody. The antigen used was a acetylated lysine 8 WB blot using the H4K8ac antibody. peptide containing the histone modification of interest. By plotting the absorbance against the antibody dilution the FORM: Liquid titer of the antibody was estimated to be 1:17,500. FORMULATION: In PBS with 0.05% (W/V) sodium azide. PURIFICATION: Whole antiserum from rabbit SPECIES REACTIVITY: Human. STORAGE CONDITIONS: Store at -20°C; for long storage, store at -80°C. Avoid multiple freeze-thaw cycles. DESCRIPTION: Histones are the main constituents of the protein part of chromosomes of eukaryotic cells. They are rich in the amino acids arginine and lysine and have been greatly conserved during evolution. Histones pack the DNA into tight masses of chromatin. Histone tails undergo numerous post-translational modifications, which either directly or indirectly alter chromatin structure to facilitate transcriptional activation or repression or other nuclear processes. In addition to the genetic code, combinations of the different histone modifications ChIP assays were performed using human reveal the so-called “histone code”. Histone methylation and demethylation is dynamically osteosarcoma (U2OS) cells and the antibody and regulated by respectively histone methyl transferases and histone demethylases. Acetylation optimized PCR primer sets for qPCR. A titration of the A Dot Blot analysis was performed to test the of histone H4 is associated with active gene transcription. -

H1 Linker Histones Silence Repetitive Elements by Promoting Both Histone H3K9 Methylation and Chromatin Compaction
H1 linker histones silence repetitive elements by promoting both histone H3K9 methylation and chromatin compaction Sean E. Healtona,1,2, Hugo D. Pintoa,1, Laxmi N. Mishraa, Gregory A. Hamiltona,b, Justin C. Wheata, Kalina Swist-Rosowskac, Nicholas Shukeirc, Yali Doud, Ulrich Steidla, Thomas Jenuweinc, Matthew J. Gamblea,b, and Arthur I. Skoultchia,2 aDepartment of Cell Biology, Albert Einstein College of Medicine, Bronx, NY 10461; bDepartment of Molecular Pharmacology, Albert Einstein College of Medicine, Bronx, NY 10461; cMax Planck Institute of Immunobiology and Epigenetics, Stübeweg 51, Freiburg D-79108, Germany; and dDepartment of Pathology, University of Michigan, Ann Arbor, MI 48109 Edited by Robert G. Roeder, Rockefeller University, New York, NY, and approved May 1, 2020 (received for review December 15, 2019) Nearly 50% of mouse and human genomes are composed of repetitive mechanisms of this regulation have not been fully explored. To sequences. Transcription of these sequences is tightly controlled during further investigate the roles of H1 in epigenetic regulation, we have development to prevent genomic instability, inappropriate gene used CRISPR-Cas9–mediated genome editing to inactivate addi- activation and other maladaptive processes. Here, we demonstrate tional H1 genes in the H1 TKO ES cells and thereby deplete the H1 an integral role for H1 linker histones in silencing repetitive elements in content to even lower levels. mouse embryonic stem cells. Strong H1 depletion causes a profound Nearly 50% of the mouse and human genomes consist of re- de-repression of several classes of repetitive sequences, including major petitive sequences, including tandem repeats, such as satellite se- satellite, LINE-1, and ERV. -

Le G´Enome En Action
LEGENOME´ EN ACTION SEQUENC´ ¸ AGE HAUT DEBIT´ ET EPIG´ ENOMIQUE´ Epig´enomique´ ? IFT6299 H2014 ? UdeM ? Mikl´osCs}ur¨os Regulation´ d’expression la transcription d’une region´ de l’ADN necessite´ ? liaisons proteine-ADN´ (facteur de transcription et son site reconnu) ? accessibilite´ de la chromatine REVIEWS Identification of regions that control transcription An initial step in the analysis of any gene is the identifi- cation of larger regions that might harbour regulatory control elements. Several advances have facilitated the prediction of such regions in the absence of knowl- edge about the specific characteristics of individual cis- Chromatin regulatory elements. These tools broadly fall into two categories: promoter (transcription start site; TSS) and enhancer detection. The methods are influenced Distal TFBS by sequence conservation between ORTHOLOGOUS genes (PHYLOGENETIC FOOTPRINTING), nucleotide composition and the assessment of available transcript data. Functional regulatory regions that control transcrip- tion rates tend to be proximal to the initiation site(s) of transcription. Although there is some circularity in the Co-activator complex data-collection process (regulatory sequences are sought near TSSs and are therefore found most often in these regions), the current set of laboratory-annotated regula- tory sequences indicates that sequences near a TSS are Transcription more likely to contain functionally important regulatory initiation complex Transcription controls than those that are more distal. However, specifi- initiation cation of the position of a TSS can be difficult. This is fur- ther complicated by the growing number of genes that CRM Proximal TFBS selectively use alternative start sites in certain contexts. Underlying most algorithms for promoter prediction is a Figure 1 | Components of transcriptional regulation. -

RNF8 Regulates Active Epigenetic Modifications and Escape Gene Activation from Inactive Sex Chromosomes in Post-Meiotic Spermatids
Downloaded from genesdev.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press RNF8 regulates active epigenetic modifications and escape gene activation from inactive sex chromosomes in post-meiotic spermatids Ho-Su Sin,1,2,3 Artem Barski,3,4,5 Fan Zhang,3,6 Andrey V. Kartashov,3,4,5 Andre Nussenzweig,7 Junjie Chen,8 Paul R. Andreassen,4,5,6 and Satoshi H. Namekawa1,2,3,9 1Division of Reproductive Sciences, 2Division of Developmental Biology, Perinatal Institute, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio 45229, USA; 3Department of Pediatrics, University of Cincinnati College of Medicine, Cincinnati, Ohio 49229, USA; 4Division of Allergy and Immunology, 5Division of Human Genetics, 6Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio 45229, USA; 7Laboratory of Genome Integrity, National Cancer Institute, National Institutes of Health, Bethesda, Maryland 20892, USA; 8Department of Experimental Radiation Oncology, University of Texas M.D. Anderson Cancer Center, Houston, Texas 77030, USA Sex chromosomes are uniquely subject to chromosome-wide silencing during male meiosis, and silencing persists into post-meiotic spermatids. Against this background, a select set of sex chromosome-linked genes escapes silencing and is activated in post-meiotic spermatids. Here, we identify a novel mechanism that regulates escape gene activation in an environment of chromosome-wide silencing in murine germ cells. We show that RNF8- dependent ubiquitination of histone H2A during meiosis establishes active epigenetic modifications, including dimethylation of H3K4 on the sex chromosomes. RNF8-dependent active epigenetic memory, defined by dimethylation of H3K4, persists throughout meiotic division.