CD24, CD27, CD36 and CD302 Gene Expression for Outcome Prediction in Patients with Multiple Myeloma
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

Role of C5ar1 and C5L2 Receptors in Ischemia-Reperfusion Injury
Journal of Clinical Medicine Article Role of C5aR1 and C5L2 Receptors in Ischemia-Reperfusion Injury Carlos Arias-Cabrales 1,* , Eva Rodriguez-Garcia 1, Javier Gimeno 2, David Benito 3, María José Pérez-Sáez 1, Dolores Redondo-Pachón 1, Anna Buxeda 1, Carla Burballa 1, Marta Crespo 1, Marta Riera 3,* and Julio Pascual 1,* 1 Department of Nephrology, Parc de Salut Mar, 08003 Barcelona, Spain; [email protected] (E.R.-G.); [email protected] (M.J.P.-S.); [email protected] (D.R.-P.); [email protected] (A.B.); [email protected] (C.B.); [email protected] (M.C.) 2 Department of Pathology, Parc de Salut Mar, 08003 Barcelona, Spain; [email protected] 3 Kidney Research Group, Hospital del Mar Medical Research Institute, IMIM, 08003 Barcelona, Spain; [email protected] * Correspondence: [email protected] (C.A.-C.); [email protected] (M.R.); [email protected] (J.P.) Abstract: The role of C5a receptors (C5aR1 and C5L2) in renal ischemia-reperfusion injury (IRI) is uncertain. We generated an in vitro model of hypoxia/reoxygenation with human proximal tubule epithelial cells to mimic some IRI events. C5aR1, membrane attack complex (MAC) and factor H (FH) deposits were evaluated with immunofluorescence. Quantitative polymerase chain reaction evaluated the expression of C5aR1, C5L2 genes as well as genes related to tubular injury, inflammation, and profibrotic pathways. Additionally, C5aR1 and C5L2 deposits were evaluated in kidney graft biopsies (KB) from transplant patients with delayed graft function (DGF, n = 12) and compared with Citation: Arias-Cabrales, C.; a control group (n = 8). We observed higher immunofluorescence expression of C5aR1, MAC and FH Rodriguez-Garcia, E.; Gimeno, J.; as higher expression of genes related to tubular injury, inflammatory and profibrotic pathways and Benito, D.; Pérez-Sáez, M.J.; of C5aR1 in the hypoxic cells; whereas, C5L2 gene expression was unaffected by the hypoxic stimulus. -

The Membrane Complement Regulatory Protein CD59 and Its Association with Rheumatoid Arthritis and Systemic Lupus Erythematosus
Current Medicine Research and Practice 9 (2019) 182e188 Contents lists available at ScienceDirect Current Medicine Research and Practice journal homepage: www.elsevier.com/locate/cmrp Review Article The membrane complement regulatory protein CD59 and its association with rheumatoid arthritis and systemic lupus erythematosus * Nibhriti Das a, Devyani Anand a, Bintili Biswas b, Deepa Kumari c, Monika Gandhi c, a Department of Biochemistry, All India Institute of Medical Sciences, New Delhi 110029, India b Department of Zoology, Ramjas College, University of Delhi, India c University School of Biotechnology, Guru Gobind Singh Indraprastha University, India article info abstract Article history: The complement cascade consisting of about 50 soluble and cell surface proteins is activated in auto- Received 8 May 2019 immune inflammatory disorders. This contributes to the pathological manifestations in these diseases. In Accepted 30 July 2019 normal health, the soluble and membrane complement regulatory proteins protect the host against Available online 5 August 2019 complement-mediated self-tissue injury by controlling the extent of complement activation within the desired limits for the host's benefit. CD59 is a membrane complement regulatory protein that inhibits the Keywords: formation of the terminal complement complex or membrane attack complex (C5b6789n) which is CD59 generated on complement activation by any of the three pathways, namely, the classical, alternative, and RA SLE the mannose-binding lectin pathway. Animal experiments and human studies have suggested impor- Pathophysiology tance of membrane complement proteins including CD59 in the pathophysiology of rheumatoid arthritis Disease marker (RA) and systemic lupus erythematosus (SLE). Here is a brief review on CD59 and its distribution, structure, functions, and association with RA and SLE starting with a brief introduction on the com- plement system, its activation, the biological functions, and relations of membrane complement regu- latory proteins, especially CD59, with RA and SLE. -
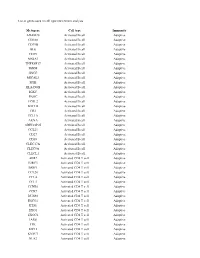
List of Genes Used in Cell Type Enrichment Analysis
List of genes used in cell type enrichment analysis Metagene Cell type Immunity ADAM28 Activated B cell Adaptive CD180 Activated B cell Adaptive CD79B Activated B cell Adaptive BLK Activated B cell Adaptive CD19 Activated B cell Adaptive MS4A1 Activated B cell Adaptive TNFRSF17 Activated B cell Adaptive IGHM Activated B cell Adaptive GNG7 Activated B cell Adaptive MICAL3 Activated B cell Adaptive SPIB Activated B cell Adaptive HLA-DOB Activated B cell Adaptive IGKC Activated B cell Adaptive PNOC Activated B cell Adaptive FCRL2 Activated B cell Adaptive BACH2 Activated B cell Adaptive CR2 Activated B cell Adaptive TCL1A Activated B cell Adaptive AKNA Activated B cell Adaptive ARHGAP25 Activated B cell Adaptive CCL21 Activated B cell Adaptive CD27 Activated B cell Adaptive CD38 Activated B cell Adaptive CLEC17A Activated B cell Adaptive CLEC9A Activated B cell Adaptive CLECL1 Activated B cell Adaptive AIM2 Activated CD4 T cell Adaptive BIRC3 Activated CD4 T cell Adaptive BRIP1 Activated CD4 T cell Adaptive CCL20 Activated CD4 T cell Adaptive CCL4 Activated CD4 T cell Adaptive CCL5 Activated CD4 T cell Adaptive CCNB1 Activated CD4 T cell Adaptive CCR7 Activated CD4 T cell Adaptive DUSP2 Activated CD4 T cell Adaptive ESCO2 Activated CD4 T cell Adaptive ETS1 Activated CD4 T cell Adaptive EXO1 Activated CD4 T cell Adaptive EXOC6 Activated CD4 T cell Adaptive IARS Activated CD4 T cell Adaptive ITK Activated CD4 T cell Adaptive KIF11 Activated CD4 T cell Adaptive KNTC1 Activated CD4 T cell Adaptive NUF2 Activated CD4 T cell Adaptive PRC1 Activated -

CD109 Released from Human Bone Marrow Mesenchymal Stem Cells Attenuates TGF-Β-Induced Epithelial to Mesenchymal Transition and Stemness of Squamous Cell Carcinoma
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 56), pp: 95632-95647 Research Paper CD109 released from human bone marrow mesenchymal stem cells attenuates TGF-β-induced epithelial to mesenchymal transition and stemness of squamous cell carcinoma Shufeng Zhou1, Renzo Cecere2 and Anie Philip1 1Division of Plastic Surgery, Department of Surgery, McGill University, Montreal, QC, Canada 2Division of Cardiac Surgery, Department of Surgery, McGill University, Montreal, QC, Canada Correspondence to: Anie Philip, email: [email protected] Keywords: human bone marrow mesenchymal stem cells; squamous cell carcinoma; CD109; TGF-β; epithelial to mesenchymal transition Abbreviations: hBM-MSC-CM: human bone marrow mesenchymal stem cells conditioned medium; hFibro-CM: human fibroblast cells conditioned medium; EMT: epithelial to mesenchymal transition; SCC: squamous cell carcinoma Received: March 31, 2017 Accepted: July 06, 2017 Published: September 16, 2017 Copyright: Zhou et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Although there is increasing evidence that human bone marrow mesenchymal stem cells (hBM-MSCs) play an important role in cancer progression, the underlying mechanisms are poorly understood. Transforming growth factor β (TGF-β) is an important pro-metastatic cytokine. We have previously shown that CD109, a glycosylphosphatidylinositol-anchored protein, is a TGF-β co-receptor and a strong inhibitor of TGF-β signalling. Moreover, CD109 can be released from the cell surface. In the current study, we examined whether hBM-MSCs regulate the malignant properties of squamous cell carcinoma cells, and whether CD109 plays a role in mediating the effect of hBM-MSCs on cancer cells. -

4-6 Weeks Old Female C57BL/6 Mice Obtained from Jackson Labs Were Used for Cell Isolation
Methods Mice: 4-6 weeks old female C57BL/6 mice obtained from Jackson labs were used for cell isolation. Female Foxp3-IRES-GFP reporter mice (1), backcrossed to B6/C57 background for 10 generations, were used for the isolation of naïve CD4 and naïve CD8 cells for the RNAseq experiments. The mice were housed in pathogen-free animal facility in the La Jolla Institute for Allergy and Immunology and were used according to protocols approved by the Institutional Animal Care and use Committee. Preparation of cells: Subsets of thymocytes were isolated by cell sorting as previously described (2), after cell surface staining using CD4 (GK1.5), CD8 (53-6.7), CD3ε (145- 2C11), CD24 (M1/69) (all from Biolegend). DP cells: CD4+CD8 int/hi; CD4 SP cells: CD4CD3 hi, CD24 int/lo; CD8 SP cells: CD8 int/hi CD4 CD3 hi, CD24 int/lo (Fig S2). Peripheral subsets were isolated after pooling spleen and lymph nodes. T cells were enriched by negative isolation using Dynabeads (Dynabeads untouched mouse T cells, 11413D, Invitrogen). After surface staining for CD4 (GK1.5), CD8 (53-6.7), CD62L (MEL-14), CD25 (PC61) and CD44 (IM7), naïve CD4+CD62L hiCD25-CD44lo and naïve CD8+CD62L hiCD25-CD44lo were obtained by sorting (BD FACS Aria). Additionally, for the RNAseq experiments, CD4 and CD8 naïve cells were isolated by sorting T cells from the Foxp3- IRES-GFP mice: CD4+CD62LhiCD25–CD44lo GFP(FOXP3)– and CD8+CD62LhiCD25– CD44lo GFP(FOXP3)– (antibodies were from Biolegend). In some cases, naïve CD4 cells were cultured in vitro under Th1 or Th2 polarizing conditions (3, 4). -

An Anticomplement Agent That Homes to the Damaged Brain and Promotes Recovery After Traumatic Brain Injury in Mice
An anticomplement agent that homes to the damaged brain and promotes recovery after traumatic brain injury in mice Marieta M. Rusevaa,1,2, Valeria Ramagliab,1, B. Paul Morgana, and Claire L. Harrisa,3 aInstitute of Infection and Immunity, School of Medicine, Cardiff University, Cardiff CF14 4XN, United Kingdom; and bDepartment of Genome Analysis, Academic Medical Center, Amsterdam 1105 AZ, The Netherlands Edited by Douglas T. Fearon, Cornell University, Cambridge, United Kingdom, and approved September 29, 2015 (received for review July 15, 2015) Activation of complement is a key determinant of neuropathology to rapidly and specifically inhibit MAC at sites of complement and disability after traumatic brain injury (TBI), and inhibition is activation, and test its therapeutic potential in experimental TBI. neuroprotective. However, systemic complement is essential to The construct, termed CD59-2a-CRIg, comprises CD59a linked fight infections, a critical complication of TBI. We describe a to CRIg via the murine IgG2a hinge. CD59a prevents assembly targeted complement inhibitor, comprising complement receptor of MAC in cell membranes (16), whereas CRIg binds C3b/iC3b of the Ig superfamily (CRIg) fused with complement regulator CD59a, deposited at sites of complement activation (17). The IgG2a designed to inhibit membrane attack complex (MAC) assembly at hinge promotes dimerization to increase ligand avidity. CD59- sites of C3b/iC3b deposition. CRIg and CD59a were linked via the 2a-CRIg protected in the TBI model, demonstrating that site- IgG2a hinge, yielding CD59-2a-CRIg dimer with increased iC3b/C3b targeted anti-MAC therapeutics may be effective in prevention binding avidity and MAC inhibitory activity. CD59-2a-CRIg inhibited of secondary neuropathology and improve neurologic recovery MAC formation and prevented complement-mediated lysis in vitro. -

Flow Reagents Single Color Antibodies CD Chart
CD CHART CD N° Alternative Name CD N° Alternative Name CD N° Alternative Name Beckman Coulter Clone Beckman Coulter Clone Beckman Coulter Clone T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells CD1a T6, R4, HTA1 Act p n n p n n S l CD99 MIC2 gene product, E2 p p p CD223 LAG-3 (Lymphocyte activation gene 3) Act n Act p n CD1b R1 Act p n n p n n S CD99R restricted CD99 p p CD224 GGT (γ-glutamyl transferase) p p p p p p CD1c R7, M241 Act S n n p n n S l CD100 SEMA4D (semaphorin 4D) p Low p p p n n CD225 Leu13, interferon induced transmembrane protein 1 (IFITM1). p p p p p CD1d R3 Act S n n Low n n S Intest CD101 V7, P126 Act n p n p n n p CD226 DNAM-1, PTA-1 Act n Act Act Act n p n CD1e R2 n n n n S CD102 ICAM-2 (intercellular adhesion molecule-2) p p n p Folli p CD227 MUC1, mucin 1, episialin, PUM, PEM, EMA, DF3, H23 Act p CD2 T11; Tp50; sheep red blood cell (SRBC) receptor; LFA-2 p S n p n n l CD103 HML-1 (human mucosal lymphocytes antigen 1), integrin aE chain S n n n n n n n l CD228 Melanotransferrin (MT), p97 p p CD3 T3, CD3 complex p n n n n n n n n n l CD104 integrin b4 chain; TSP-1180 n n n n n n n p p CD229 Ly9, T-lymphocyte surface antigen p p n p n -

Mouse CD302/CLEC13A Antibody Antigen Affinity-Purified Polyclonal Sheep Igg Catalog Number: AF6424
Mouse CD302/CLEC13A Antibody Antigen Affinity-purified Polyclonal Sheep IgG Catalog Number: AF6424 DESCRIPTION Species Reactivity Mouse Specificity Detects mouse CD302/CLEC13A in direct ELISAs and Western blots. In direct ELISAs, approximately 30% crossreactivity with recombinant human CD302 is observed. Source Polyclonal Sheep IgG Purification Antigen Affinitypurified Immunogen Mouse myeloma cell line NS0derived recombinant mouse CD302/CLEC13A Asp21His165 Accession # Q9DCG2 Formulation Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. See Certificate of Analysis for details. *Small pack size (SP) is supplied either lyophilized or as a 0.2 μm filtered solution in PBS. APPLICATIONS Please Note: Optimal dilutions should be determined by each laboratory for each application. General Protocols are available in the Technical Information section on our website. Recommended Sample Concentration Western Blot 1 µg/mL See Below DATA Western Blot Detection of Mouse CD302/CLEC13A by Western Blot. Western blot shows lysates of mouse spleen tissue. PVDF Membrane was probed with 1 µg/mL of Mouse CD302/CLEC13A Antigen Affinitypurified Polyclonal Antibody (Catalog # AF6424) followed by HRPconjugated AntiSheep IgG Secondary Antibody (Catalog # HAF016). A specific band was detected for CD302/CLEC13A at approximately 32 kDa (as indicated). This experiment was conducted under reducing conditions and using Immunoblot Buffer Group 8. PREPARATION AND STORAGE Reconstitution Sterile PBS to a final concentration of 0.2 mg/mL. Shipping The product is shipped at ambient temperature. Upon receipt, store it immediately at the temperature recommended below. *Small pack size (SP) is shipped with polar packs. Upon receipt, store it immediately at 20 to 70 °C Stability & Storage Use a manual defrost freezer and avoid repeated freezethaw cycles. -

Quantitation of the Number of Molecules of Glycophorins C and D on Normal Red Blood Cells Using Radioiodinatedfab Fragments of Monoclonal Antibodies
Quantitation of the Number of Molecules of Glycophorins C and D on Normal Red Blood Cells Using RadioiodinatedFab Fragments of Monoclonal Antibodies By Jon Smythe, Brigitte Gardner, andDavid J. Anstee Two rat monoclonal antibodies (BRAC 1 and BRAC 1 1 ) cytes. Fabfragments of BRAC 1 1 and ERIC 10 gave values have been produced. BRAC 1 recognizes an epitope com- of 143,000 molecules GPC per red blood cell (RBC). Fab mon to the human erythrocyte membrane glycoproteins fragments of BRAC1 gave 225,000 molecules of GPC and glycophorin C (GPC) and glycophorin D (GPD). BRAC 11 GPD per RBC. These results indicate that GPC and GPD is specific for GPC. Fabfragments of these antibodies and together are sufficiently abundantto provide membrane at- BRlC 10, a murine monoclonal anti-GPC,were radioiodin- tachment sites for all ofthe protein 4.1 in normal RBCs. ated and used in quantitative binding assays to measure 0 1994 by The American Societyof Hematology. the number of GPC and GPD molecules on normal erythro- HE SHAPE AND deformability of the mature human (200,000)" and those reported for GPC (50,000).7 This nu- Downloaded from http://ashpublications.org/blood/article-pdf/83/6/1668/612763/1668.pdf by guest on 24 September 2021 T erythrocyte is controlled by a flexible two-dimensional merical differencehas led to the suggestion that a significant lattice of proteins, which together comprise the membrane proportion of protein 4.1 in normal erythrocyte membranes skeleton.' The major components of the skeleton are spec- must be bound to sites other than GPC and GPD.3 The trin, actin, ankyrin, and protein 4.1. -

Anti-GPRC5D/CD3 Bispecific T Cell-Redirecting Antibody for the Treatment of Multiple Myeloma
Author Manuscript Published OnlineFirst on July 3, 2019; DOI: 10.1158/1535-7163.MCT-18-1216 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. MCT-18-1216R1, Molecular Cancer Therapeutics, T. Kodama et al. Anti-GPRC5D/CD3 bispecific T cell-redirecting antibody for the treatment of multiple myeloma Tatsushi Kodama1, 2, Yu Kochi3, Waka Nakai2, Hideaki Mizuno2, Takeshi Baba2, Kiyoshi Habu2, Noriaki Sawada2, Hiroyuki Tsunoda2, Takahiro Shima3, Kohta Miyawaki3, Yoshikane Kikushige3, Yasuo Mori3, Toshihiro Miyamoto3, Takahiro Maeda4, and Koichi Akashi3, 4 Authors' Affiliations: 1Chugai Pharmabody Research Pte. Ltd., Singapore, 2Research Division, Chugai Pharmaceutical Co., Ltd., Kamakura, Kanagawa, Japan, 3Department of Medicine and Biosystemic Science, Kyushu University Graduate School of Medical Sciences, Fukuoka, Japan, 4Center for Cellular and Molecular Medicine, Kyushu University Hospital, Fukuoka, Japan Corresponding Authors: Tatsushi Kodama, Chugai Pharmabody Research Pte. Ltd., 3 Biopolis Drive, #07-11 to 16, Synapse, 138623, Singapore; Phone: + 65-6933-4860; Fax: + 65-6684-2257; E-mail: [email protected] Running Title: Anti-GPRC5D/CD3 bispecific T cell-redirecting antibody Keywords: GPRC5D, bispecific T cell-redirecting antibody, multiple myeloma Financial information: This study was funded by Chugai Pharmaceutical Co., Ltd. Conflict of interest statement: Tatsushi Kodama, Waka Nakai, Hideaki Mizuno, Takeshi Baba, Kiyoshi Habu, Noriaki Sawada, and Hiroyuki Tsunoda are employees of Chugai 1 Downloaded from mct.aacrjournals.org on September 24, 2021. © 2019 American Association for Cancer Research. Author Manuscript Published OnlineFirst on July 3, 2019; DOI: 10.1158/1535-7163.MCT-18-1216 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2