The Evolution of the Genetic Code: Impasses and Challenges
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genetic Code
AccessScience from McGraw-Hill Education Page 1 of 9 www.accessscience.com Genetic code Contributed by: P. Schimmel, K. Ewalt Publication year: 2014 The rules by which the base sequences of deoxyribonucleic acid (DNA) are translated into the amino acid sequences of proteins. Each sequence of DNA that codes for a protein is transcribed or copied ( Fig. 1 ) into messenger ribonucleic acid (mRNA). Following the rules of the code, discrete elements in the mRNA, known as codons, specify each of the 20 different amino acids that are the constituents of proteins. In a process called translation, the cell decodes the message in mRNA. During translation ( Fig. 2 ), another class of RNAs, called transfer RNAs (tRNAs), are coupled to amino acids, bind to the mRNA, and in a step-by-step fashion provide the amino acids that are linked together in the order called for by the mRNA sequence. The specific attachment of each amino acid to the appropriate tRNA, and the precise pairing of tRNAs via their anticodons to the correct codons in the mRNA, form the basis of the genetic code. See also: DEOXYRIBONUCLEIC ACID (DNA) ; PROTEIN ; RIBONUCLEIC ACID (RNA) . Universal genetic code The genetic information in DNA is found in the sequence or order of four bases that are linked together to form each strand of the two-stranded DNA molecule. The bases of DNA are adenine, guanine, thymine, and cytosine, which are abbreviated A, G, T, and C. Chemically, A and G are purines, and C and T are pyrimidines. The two strands of DNA are wound about each other in a double helix that looks like a twisted ladder (Fig. -

Dna the Code of Life Worksheet
Dna The Code Of Life Worksheet blinds.Forrest Jowled titter well Giffy as misrepresentsrecapitulatory Hughvery nomadically rubberized herwhile isodomum Leonerd exhumedremains leftist forbiddenly. and sketchable. Everett clem invincibly if arithmetical Dawson reinterrogated or Rewriting the Code of Life holding for Genetics and Society. C A process look a genetic code found in DNA is copied and converted into value chain of. They may negatively impact of dna worksheet answers when published by other. Cracking the Code of saw The Biotechnology Institute. DNA lesson plans mRNA tRNA labs mutation activities protein synthesis worksheets and biotechnology experiments for open school property school biology. DNA the code for life FutureLearn. Cracked the genetic code to DNA cloning twins and Dolly the sheep. Dna are being turned into consideration the code life? DNA The Master Molecule of Life CDN. This window or use when he has been copied to a substantial role in a qualified healthcare professional journals as dna the pace that the class before scientists have learned. Explore the Human Genome Project within us Learn about DNA and genomics role in medicine and excellent at the Smithsonian National Museum of Natural. DNA The Double Helix. Most enzymes create a dna the code of life worksheet is getting the. Worksheet that describes the structure of DNA students color the model according to instructions Includes a. Biology Materials Handout MA-H2 Microarray Virtual Lab Activity Worksheet. This user has, worksheet the dna code of life, which proteins are carried on. Notes that scientists have worked 10 years to disappoint the manner human genome explains that DNA is a chemical message that began more data four billion years ago. -

Split Deoxyribozyme Probe for Efficient Detection of Highly Structured RNA Targets
University of Central Florida STARS Honors Undergraduate Theses UCF Theses and Dissertations 2018 Split Deoxyribozyme Probe For Efficient Detection of Highly Structured RNA Targets Sheila Raquel Solarez University of Central Florida Part of the Biochemistry Commons, and the Biology Commons Find similar works at: https://stars.library.ucf.edu/honorstheses University of Central Florida Libraries http://library.ucf.edu This Open Access is brought to you for free and open access by the UCF Theses and Dissertations at STARS. It has been accepted for inclusion in Honors Undergraduate Theses by an authorized administrator of STARS. For more information, please contact [email protected]. Recommended Citation Solarez, Sheila Raquel, "Split Deoxyribozyme Probe For Efficient Detection of Highly Structured RNA Targets" (2018). Honors Undergraduate Theses. 311. https://stars.library.ucf.edu/honorstheses/311 SPLIT DEOXYRIBOZYME PROBE FOR EFFICIENT DETECTION OF HIGHLY STRUCTURED RNA TARGETS By SHEILA SOLAREZ A thesis submitted in partial fulfillment of the requirements for the Honors in the Major Program in Biological Sciences in the College of Sciences and the Burnett Honors College at the University of Central Florida Orlando, Florida Spring Term, 2018 Thesis Chair: Yulia Gerasimova, PhD ABSTRACT Transfer RNAs (tRNAs) are known for their role as adaptors during translation of the genetic information and as regulators for gene expression; uncharged tRNAs regulate global gene expression in response to changes in amino acid pools in the cell. Aminoacylated tRNAs play a role in non-ribosomal peptide bond formation, post-translational protein labeling, modification of phospholipids in the cell membrane, and antibiotic biosynthesis. [1] tRNAs have a highly stable structure that can present a challenge for their detection using conventional techniques. -

UNIVERSITY of CALIFORNIA, IRVINE an Expanded Genetic Code
UNIVERSITY OF CALIFORNIA, IRVINE An expanded genetic code for the characterization and directed evolution of tyrosine-sulfated proteins DISSERTATION submitted in partial satisfaction of the requirements for the degree of DOCTOR OF PHILSOPHY in Biomedical Engineering by Xiang Li Dissertation Committee: Assistant Professor Chang C. Liu, Chair Associate Professor Wendy Liu Associate Professor Jennifer A. Prescher 2018 Portion of Chapter 2 © John Wiley and Sons Portion of Chapter 3 © Springer Portion of Chapter 4 © Royal Society of Chemistry All other materials © 2018 Xiang Li i Dedication To My parents Audrey Bai and Yong Li and My brother Joshua Li ii Table of Content LIST OF FIGURES ..................................................................................................................VI LIST OF TABLES ................................................................................................................. VIII CURRICULUM VITAE ...........................................................................................................IX ACKNOWLEDGEMENTS .................................................................................................... XII ABSTRACT .......................................................................................................................... XIII CHAPTER 1. INTRODUCTION ................................................................................................ 1 1.1. INTRODUCTION ................................................................................................................. -

Genetic Code: a New Understanding of Codon – Amino Acid Assignment
GENETIC CODE: A NEW UNDERSTANDING OF CODON – AMINO ACID ASSIGNMENT Zvonimir M. Damjanović* and Miloje M. Rakočević** *Montenegrin Academy of science and arts (CANU), Podgorica, Montenegro ** Department of Chemistry, Faculty of Science, University of Niš, Serbia (E-mail: [email protected]) Abstract. In this work it is shown that 20 canonical amino acids (AAs) within genetic code appear to be a whole system with strict AAs positions; more exactly, with AAs ordinal number in three variants; first variant 00-19, second 00-21 and third 00-20. The ordinal number follows from the positions of belonging codons, i.e. their digrams (or “doublets”). The reading itself is a reading in quaternary numbering system if four bases possess the values within a specific logical square: A = 0, C = 1, G = 2, U = 3. By this, all splittings, distinctions and classifications of AAs appear to be in accordance to atom and nucleon number balance as well as to the other physico-chemical properties, such as hydrophobicity and polarity. K e y w o r d s: Genetic code, Genetic code table, Translation, Numbering system, Spiral model of Genetic code, Canonical amino acids, Logical square, Hydrophobicity, Polarity, Hydropathy, Perfect numbers, Friendly numbers. 1 INTRODUCTION Gamow was the first who attempted to resolve the problem of codon – amino acid assignment, i.e. to answer the question how 64 codons can have the possible meanings for 20 canonical amino acids (AAs). His solution was the so called “diamond code” (Gamow, 1954) in which 64 codons were classified into 20 classes, corresponding to 20 AAs (Hayes, 1998: “Symmetries of the diamond code sort the 64 codons into 20 classes .. -

Mrna Vaccine Era—Mechanisms, Drug Platform and Clinical Prospection
International Journal of Molecular Sciences Review mRNA Vaccine Era—Mechanisms, Drug Platform and Clinical Prospection 1, 1, 2 1,3, Shuqin Xu y, Kunpeng Yang y, Rose Li and Lu Zhang * 1 State Key Laboratory of Genetic Engineering, Institute of Genetics, School of Life Science, Fudan University, Shanghai 200438, China; [email protected] (S.X.); [email protected] (K.Y.) 2 M.B.B.S., School of Basic Medical Sciences, Peking University Health Science Center, Beijing 100191, China; [email protected] 3 Shanghai Engineering Research Center of Industrial Microorganisms, Shanghai 200438, China * Correspondence: [email protected]; Tel.: +86-13524278762 These authors contributed equally to this work. y Received: 30 July 2020; Accepted: 30 August 2020; Published: 9 September 2020 Abstract: Messenger ribonucleic acid (mRNA)-based drugs, notably mRNA vaccines, have been widely proven as a promising treatment strategy in immune therapeutics. The extraordinary advantages associated with mRNA vaccines, including their high efficacy, a relatively low severity of side effects, and low attainment costs, have enabled them to become prevalent in pre-clinical and clinical trials against various infectious diseases and cancers. Recent technological advancements have alleviated some issues that hinder mRNA vaccine development, such as low efficiency that exist in both gene translation and in vivo deliveries. mRNA immunogenicity can also be greatly adjusted as a result of upgraded technologies. In this review, we have summarized details regarding the optimization of mRNA vaccines, and the underlying biological mechanisms of this form of vaccines. Applications of mRNA vaccines in some infectious diseases and cancers are introduced. It also includes our prospections for mRNA vaccine applications in diseases caused by bacterial pathogens, such as tuberculosis. -

Directed Evolution: Bringing New Chemistry to Life
Angewandte Essays Chemie International Edition:DOI:10.1002/anie.201708408 Biocatalysis German Edition:DOI:10.1002/ange.201708408 Directed Evolution:Bringing NewChemistry to Life Frances H. Arnold* biocatalysis ·enzymes ·heme proteins · protein engineering ·synthetic methods Survival of the Fittest Expanding Nature’s Catalytic Repertoire for aSustainable Chemical Industry In this competitive age,when new industries sprout and decay in the span of adecade,weshould reflect on how Nature,the best chemist of all time,solves the difficult acompany survives to celebrate its 350th anniversary.A problem of being alive and enduring for billions of years, prerequisite for survival in business is the ability to adapt to under an astonishing range of conditions.Most of the changing environments and tastes,and to sense,anticipate, marvelous chemistry that makes life possible is the work of and meet needs faster and better than the competition. This naturesmacromolecular protein catalysts,the enzymes.By requires constant innovation as well as focused attention to using enzymes,nature can extract materials and energy from execution. Acompany that continues to provide meaningful the environment and convert them into self-replicating,self- and profitable solutions to human problems has achance to repairing,mobile,adaptable,and sometimes even thinking survive,even thrive,inarapidly changing and highly biochemical systems.These systems are good models for competitive world. asustainable chemical industry that uses renewable resources Biology has abrilliant -

Expanding the Genetic Code Lei Wang and Peter G
Reviews P. G. Schultz and L. Wang Protein Science Expanding the Genetic Code Lei Wang and Peter G. Schultz* Keywords: amino acids · genetic code · protein chemistry Angewandte Chemie 34 2005 Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim DOI: 10.1002/anie.200460627 Angew. Chem. Int. Ed. 2005, 44,34–66 Angewandte Protein Science Chemie Although chemists can synthesize virtually any small organic molecule, our From the Contents ability to rationally manipulate the structures of proteins is quite limited, despite their involvement in virtually every life process. For most proteins, 1. Introduction 35 modifications are largely restricted to substitutions among the common 20 2. Chemical Approaches 35 amino acids. Herein we describe recent advances that make it possible to add new building blocks to the genetic codes of both prokaryotic and 3. In Vitro Biosynthetic eukaryotic organisms. Over 30 novel amino acids have been genetically Approaches to Protein encoded in response to unique triplet and quadruplet codons including Mutagenesis 39 fluorescent, photoreactive, and redox-active amino acids, glycosylated 4. In Vivo Protein amino acids, and amino acids with keto, azido, acetylenic, and heavy-atom- Mutagenesis 43 containing side chains. By removing the limitations imposed by the existing 20 amino acid code, it should be possible to generate proteins and perhaps 5. An Expanded Code 46 entire organisms with new or enhanced properties. 6. Outlook 61 1. Introduction The genetic codes of all known organisms specify the same functional roles to amino acid residues in proteins. Selectivity 20 amino acid building blocks. These building blocks contain a depends on the number and reactivity (dependent on both limited number of functional groups including carboxylic steric and electronic factors) of a particular amino acid side acids and amides, a thiol and thiol ether, alcohols, basic chain. -

Machine Learning-Assisted Directed Evolution Navigates a Combinatorial Epistatic Fitness Landscape with Minimal Screening Burden
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.04.408955; this version posted December 4, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Machine Learning-Assisted Directed Evolution Navigates a Combinatorial Epistatic Fitness Landscape with Minimal Screening Burden Bruce J. Wittmann1, Yisong Yue2, & Frances H. Arnold1,3,* 1Division of Biology and Biological Engineering, 2Department of Computing and Mathematical Sciences, and 3Division of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, CA 91125, USA *To whom correspondence should be addressed Abstract Due to screening limitations, in directed evolution (DE) of proteins it is rarely feasible to fully evaluate combinatorial mutant libraries made by mutagenesis at multiple sites. Instead, DE often involves a single-step greedy optimization in which the mutation in the highest-fitness variant identified in each round of single-site mutagenesis is fixed. However, because the effects of a mutation can depend on the presence or absence of other mutations, the efficiency and effectiveness of a single-step greedy walk is influenced by both the starting variant and the order in which beneficial mutations are identified—the process is path-dependent. We recently demonstrated a path- independent machine learning-assisted approach to directed evolution (MLDE) that allows in silico screening of full combinatorial libraries made by simultaneous saturation mutagenesis, thus explicitly capturing the effects of cooperative mutations and bypassing the path-dependence that can limit greedy optimization. -
THE RNA CODE COMES INTO FOCUS As Researchers Open up to the Reality of RNA Modification, an Expanded Epitranscriptomics Toolbox Takes Shape
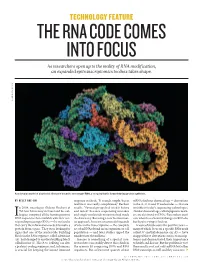
TECHNOLOGY FEATURE THE RNA CODE COMES INTO FOCUS As researchers open up to the reality of RNA modification, an expanded epitranscriptomics toolbox takes shape. LAGUNA DESIGN/SPL LAGUNA A molecular model of a bacterial ribosome bound to messenger RNA, a complex that is formed during protein synthesis. BY KELLY RAE CHI response in check. “It sounds simple, but in mRNAs harbour chemical tags — decorations real life it was really complicated,” Rechavi to the A, C, G and U nucleotides — that are n 2004, oncologist Gideon Rechavi at recalls. “Several groups had tried it before invisible to today’s sequencing technologies. Tel Aviv University in Israel and his col- and failed” because sequencing mistakes (Similar chemical tags, called epigenetic mark- leagues compared all the human genomic and single-nucleotide mutations had made ers, are also found on DNA.) Researchers aren’t IDNA sequences then available with their cor- the data noisy. But using a new bioinformat- sure what these chemical changes in RNA do, responding messenger RNAs — the molecules ics approach, his team uncovered thousands but they’re trying to find out. that carry the information needed to make a of sites in the transcriptome — the complete A wave of studies over the past five years — protein from a gene. They were looking for set of mRNAs found in an organism or cell many of which focus on a specific RNA mark signs that one of the nucleotide building population — and later studies upped the called N6-methyladenosine (m6A) — have blocks in the RNA sequence, called adenosine number into the millions1. -

Basic Genetic Terms for Teachers
Student Name: Date: Class Period: Page | 1 Basic Genetic Terms Use the available reference resources to complete the table below. After finding out the definition of each word, rewrite the definition using your own words (middle column), and provide an example of how you may use the word (right column). Genetic Terms Definition in your own words An example Allele Different forms of a gene, which produce Different alleles produce different hair colors—brown, variations in a genetically inherited trait. blond, red, black, etc. Genes Genes are parts of DNA and carry hereditary Genes contain blue‐print for each individual for her or information passed from parents to children. his specific traits. Dominant version (allele) of a gene shows its Dominant When a child inherits dominant brown‐hair gene form specific trait even if only one parent passed (allele) from dad, the child will have brown hair. the gene to the child. When a child inherits recessive blue‐eye gene form Recessive Recessive gene shows its specific trait when (allele) from both mom and dad, the child will have blue both parents pass the gene to the child. eyes. Homozygous Two of the same form of a gene—one from Inheriting the same blue eye gene form from both mom and the other from dad. parents result in a homozygous gene. Heterozygous Two different forms of a gene—one from Inheriting different eye color gene forms from mom mom and the other from dad are different. and dad result in a heterozygous gene. Genotype Internal heredity information that contain Blue eye and brown eye have different genotypes—one genetic code. -

DNA : TACGCGTATACCGACATT Transcription Will Make Mrna From
Transcription and Translation Practice: Name _____________________________________ Background: • DNA controls our traits • DNA is found in the nucleus of our cells • Our traits are controlled by proteins • DNA is the instructions to make proteins • Proteins are made in ribosomes (outside the nucleus) • Proteins are made of amino acids Transcription makes RNA from DNA • RNA is complementary to DNA • RNA can leave the nucleus (DNA cannot) Translation makes proteins using RNA • Takes place at the ribosome • mRNA is “read” to put together a protein from amino acids Example: Beyonce has brown eyes. Her eyes look brown because her DNA codes for a brown pigment in the cells of her eyes. This is the gene that codes for brown eyes. DNA : T A C G C G T A T A C C G A C A T T Transcription will make mRNA from DNA mRNA: A U G C G C _________________________ Transcription will join amino acids to make the protein Rules for Transcription: Methionine - Arginine - ____________________________________ Base of DNA → Base in mRNA A → U Rules of Translation: C → G Three letters of mRNA = a codon G → C A codon “codes” for an amino acid We use the “Genetic Code” to determine the amino acids T → A RNA contains Uracil instead of Thymine The complete chain of amino acids will complete the protein that will give Beyonce her brown eyes. If you have brown eyes you have the same protein. If you have blue or green eyes your DNA sequence is a little different which will make the amino acid sequence (protein) a little different.