Chromosomes 8, 12, and 17 Copy Number in Astler-Coller Stage C Colon Cancer in Relation to Proliferative Activity and DNA Ploidy1
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

12Q Deletions FTNW
12q deletions rarechromo.org What is a 12q deletion? A deletion from chromosome 12q is a rare genetic condition in which a part of one of the body’s 46 chromosomes is missing. When material is missing from a chromosome, it is called a deletion. What are chromosomes? Chromosomes are the structures in each of the body’s cells that carry genetic information telling the body how to develop and function. They come in pairs, one from each parent, and are numbered 1 to 22 approximately from largest to smallest. Additionally there is a pair of sex chromosomes, two named X in females, and one X and another named Y in males. Each chromosome has a short (p) arm and a long (q) arm. Looking at chromosome 12 Chromosome analysis You can’t see chromosomes with the naked eye, but if you stain and magnify them many hundreds of times under a microscope, you can see that each one has a distinctive pattern of light and dark bands. In the diagram of the long arm of chromosome 12 on page 3 you can see the bands are numbered outwards starting from the point at the top of the diagram where the short and long arms meet (the centromere). Molecular techniques If you magnify chromosome 12 about 850 times, a small piece may be visibly missing. But sometimes the missing piece is so tiny that the chromosome looks normal through a microscope. The missing section can then only be found using more sensitive molecular techniques such as FISH (fluorescence in situ hybridisation, a technique that reveals the chromosomes in fluorescent colour), MLPA (multiplex ligation-dependent probe amplification) and/or array-CGH (microarrays), a technique that shows gains and losses of tiny amounts of DNA throughout all the chromosomes. -

Investigation of Differentially Expressed Genes in Nasopharyngeal Carcinoma by Integrated Bioinformatics Analysis
916 ONCOLOGY LETTERS 18: 916-926, 2019 Investigation of differentially expressed genes in nasopharyngeal carcinoma by integrated bioinformatics analysis ZhENNING ZOU1*, SIYUAN GAN1*, ShUGUANG LIU2, RUjIA LI1 and jIAN hUANG1 1Department of Pathology, Guangdong Medical University, Zhanjiang, Guangdong 524023; 2Department of Pathology, The Eighth Affiliated hospital of Sun Yat‑sen University, Shenzhen, Guangdong 518033, P.R. China Received October 9, 2018; Accepted April 10, 2019 DOI: 10.3892/ol.2019.10382 Abstract. Nasopharyngeal carcinoma (NPC) is a common topoisomerase 2α and TPX2 microtubule nucleation factor), malignancy of the head and neck. The aim of the present study 8 modules, and 14 TFs were identified. Modules analysis was to conduct an integrated bioinformatics analysis of differ- revealed that cyclin-dependent kinase 1 and exportin 1 were entially expressed genes (DEGs) and to explore the molecular involved in the pathway of Epstein‑Barr virus infection. In mechanisms of NPC. Two profiling datasets, GSE12452 and summary, the hub genes, key modules and TFs identified in GSE34573, were downloaded from the Gene Expression this study may promote our understanding of the pathogenesis Omnibus database and included 44 NPC specimens and of NPC and require further in-depth investigation. 13 normal nasopharyngeal tissues. R software was used to identify the DEGs between NPC and normal nasopharyngeal Introduction tissues. Distributions of DEGs in chromosomes were explored based on the annotation file and the CYTOBAND database Nasopharyngeal carcinoma (NPC) is a common malignancy of DAVID. Gene ontology (GO) and Kyoto Encyclopedia of occurring in the head and neck. It is prevalent in the eastern Genes and Genomes (KEGG) pathway enrichment analysis and southeastern parts of Asia, especially in southern China, were applied. -

Pallister-Killian Syndrome Groups Worldwide Who Are Studying the Different Aspects of This Syndrome
Inform Network Support Rare Chromosome Disorder Support Group G1, The Stables, Station Road West, Oxted, Surrey RH8 9EE, United Kingdom Tel: +44(0)1883 723356 [email protected] I www.rarechromo.org Join Unique for family links, information and support. Unique is a charity without government funding, existing entirely on donations and grants. If you can, please make a donation via our website at Pallister-Killian www.rarechromo.org Please help us to help you! Websites, Facebook groups http://pkskids.net syndrome http://pkskids.ning.com www.pks.org.au www.pksitalia.org www.facebook.com/pages/Pallister-Killian-Syndrome-PKS www.facebook.com/pages/Sindrome-Pallister-Killian (in Spanish) Unique mentions other organisations’ message boards and websites to help families looking for information. This does not imply that we endorse their content or have any responsibility for it. This information guide is not a substitute for personal medical advice. Families should consult a medically qualified clinician in all matters relating to genetic diagnosis, management and health. Information on genetic changes is a very fast-moving field and while the information in this guide is believed to be the best available at the time of publication, some facts may later change. Unique does its best to keep abreast of changing information and to review its published guides as needed. This booklet was compiled by Unique and reviewed by Dr Michel Vekemans, Department of Genetics, Hopital Necker Enfants Malades, Paris, France 2004 and by Unique’s chief medical advisor, Professor Maj Hultén, Professor of Reproductive Genetics, University of Warwick, UK. 2005 (PM). -

Centromere Repositioning Causes Inversion of Meiosis and Generates a Reproductive Barrier
Centromere repositioning causes inversion of meiosis and generates a reproductive barrier Min Lua and Xiangwei Hea,1 aMinistry of Education Key Laboratory of Biosystems Homeostasis & Protection and Innovation Center for Cell Signaling Network, Life Sciences Institute, Zhejiang University, 310058 Hangzhou, Zhejiang, China Edited by J. Richard McIntosh, University of Colorado, Boulder, CO, and approved September 20, 2019 (received for review July 10, 2019) The chromosomal position of each centromere is determined postulated that a neocentromere may seed the formation of an epigenetically and is highly stable, whereas incremental cases ENC at a site devoid of satellite DNA, which is then matured have supported the occurrence of centromere repositioning on an through acquisition of repetitive DNA. ENCs and neocentromeres evolutionary time scale (evolutionary new centromeres, ENCs), are considered as two sides of the same coin, manifestations of the which is thought to be important in speciation. The mechanisms same biological phenomenon at drastically different time scales underlying the high stability of centromeres and its functional and population sizes (7). Hence, understanding centromere repo- significance largely remain an enigma. Here, in the fission yeast sitioning may provide mechanistic insights into ENC emergence Schizosaccharomyces pombe, we identify a feedback mechanism: and progression. The kinetochore, whose assembly is guided by the centromere, in CENP-A–containing chromatin directly recruits specific com- turn, enforces centromere stability. Upon going through meiosis, ponents of the kinetochore, called the constitutive centromere- specific inner kinetochore mutations induce centromere reposi- associated network. The kinetochore is a proteinaceous ma- tioning—inactivation of the original centromere and formation chinery comprised of inner and outer parts, each compassing of a new centromere elsewhere—in 1 of the 3 chromosomes at several subcomplexes. -

Receptor Signaling Through Osteoclast-Associated Monocyte
Downloaded from http://www.jimmunol.org/ by guest on September 29, 2021 is online at: average * The Journal of Immunology The Journal of Immunology , 20 of which you can access for free at: 2015; 194:3169-3179; Prepublished online 27 from submission to initial decision 4 weeks from acceptance to publication February 2015; doi: 10.4049/jimmunol.1402800 http://www.jimmunol.org/content/194/7/3169 Collagen Induces Maturation of Human Monocyte-Derived Dendritic Cells by Signaling through Osteoclast-Associated Receptor Heidi S. Schultz, Louise M. Nitze, Louise H. Zeuthen, Pernille Keller, Albrecht Gruhler, Jesper Pass, Jianhe Chen, Li Guo, Andrew J. Fleetwood, John A. Hamilton, Martin W. Berchtold and Svetlana Panina J Immunol cites 43 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Author Choice option Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Freely available online through http://www.jimmunol.org/content/suppl/2015/02/27/jimmunol.140280 0.DCSupplemental This article http://www.jimmunol.org/content/194/7/3169.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Author Choice Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2015 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. -

Chicken Skeletal Muscle-Associated Macroarray for Gene Discovery
Chicken skeletal muscle-associated macroarray for gene discovery E.C. Jorge1, C.M.R. Melo2, M.F. Rosário1, J.R.S. Rossi1, M.C. Ledur3, A.S.A.M.T. Moura4 and L.L. Coutinho1 1Departamento de Zootecnia, Escola Superior de Agricultura “Luiz de Queiroz”, Universidade de São Paulo, Piracicaba, SP, Brasil 2Departamento de Aquicultura, Universidade Federal de Santa Catarina, Florianópolis, SC, Brasil 3Embrapa Suínos e Aves, Genética e Melhoramento Animal, Vila Tamanduá, Concórdia, SC, Brasil 4Departamento de Produção Animal, Faculdade de Medicina Veterinária e Zootecnia de Botucatu, Universidade Estadual Paulista Júlio de Mesquita Filho, Botucatu, SP, Brasil Corresponding author: L.L. Coutinho E-mail: [email protected] Genet. Mol. Res. 9 (1): 188-207 (2010) Received November 4, 2009 Accepted December 2, 2009 Published February 2, 2010 ABSTRACT. Macro- and microarrays are well-established technologies to determine gene functions through repeated measurements of transcript abundance. We constructed a chicken skeletal muscle-associated array based on a muscle-specific EST database, which was used to generate a tissue expression dataset of ~4500 chicken genes across 5 adult tissues (skeletal muscle, heart, liver, brain, and skin). Only a small number of ESTs were sufficiently well characterized by BLAST searches to determine their probable cellular functions. Evidence of a particular tissue-characteristic expression can be considered an indication that the transcript is likely to be functionally significant. The skeletal muscle macroarray platform was first used to search for evidence of tissue- specific expression, focusing on the biological function of genes/ transcripts, since gene expression profiles generated across tissues were found to be reliable and consistent. -
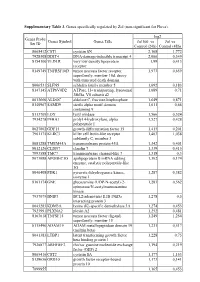
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis -

Chromosome 12 Introduction the Genetic Length of Chromosome 12 Is ~132 Mb. It Is ~4–4.5% of the Total Human Genome, Approximat
Chromosome 12 ©Chromosome Disorder Outreach Inc. (CDO) Technical genetic content provided by Dr. Iosif Lurie, M.D. Ph.D Medical Geneticist and CDO Medical Consultant/Advisor. Ideogram courtesy of the University of Washington Department of Pathology: ©1994 David Adler.hum_12.gif Introduction The genetic length of chromosome 12 is ~132 Mb. It is ~4–4.5% of the total human genome, approximately the same size as chromosomes 10 and 11. The length of the short arm is ~35 Mb; the length of the long arm is 97 Mb. Chromosome 12 contains ~1,300 genes. Some of these genes control the development of body organs or participate in maintaining body functions. Structural abnormalities of chromosome 12 are relatively rare. There are only ~150 reports on patients with deletions of the short arm and approximately the same number of persons with deletions of the long arm (in both cases, including patients having an additional imbalance for another chromosome). Surprisingly enough, there are no clear–cut syndromes caused by deletions of chromosome 12 (except, maybe, patients with tiny deletions 12q12 having Buschke–Ollendorf syndrome). More precise determination of breakpoints and a clinical comparison of patients having similar well– characterized deletions may lead to a description of a new syndrome caused by deletions of chromosome 12. Deletions of Chromosome 12 Deletions of 12p The genetic size of the chromosome 12 is ~132 Mb, where the short arm is ~35 Mb. Deletions of this area are relatively rare: there are only ~50 reports of patients with “pure” deletions of the short arm. Until now, there are no syndromes caused by such deletions. -

A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression
Supplementary Materials A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression Table S1. Statistically significant DEGs (Adj. p-value < 0.01) derived from meta-analysis for samples irradiated with high doses of HZE particles, collected 6-24 h post-IR not common with any other meta- analysis group. This meta-analysis group consists of 3 DEG lists obtained from DGEA, using a total of 11 control and 11 irradiated samples [Data Series: E-MTAB-5761 and E-MTAB-5754]. Ensembl ID Gene Symbol Gene Description Up-Regulated Genes ↑ (2425) ENSG00000000938 FGR FGR proto-oncogene, Src family tyrosine kinase ENSG00000001036 FUCA2 alpha-L-fucosidase 2 ENSG00000001084 GCLC glutamate-cysteine ligase catalytic subunit ENSG00000001631 KRIT1 KRIT1 ankyrin repeat containing ENSG00000002079 MYH16 myosin heavy chain 16 pseudogene ENSG00000002587 HS3ST1 heparan sulfate-glucosamine 3-sulfotransferase 1 ENSG00000003056 M6PR mannose-6-phosphate receptor, cation dependent ENSG00000004059 ARF5 ADP ribosylation factor 5 ENSG00000004777 ARHGAP33 Rho GTPase activating protein 33 ENSG00000004799 PDK4 pyruvate dehydrogenase kinase 4 ENSG00000004848 ARX aristaless related homeobox ENSG00000005022 SLC25A5 solute carrier family 25 member 5 ENSG00000005108 THSD7A thrombospondin type 1 domain containing 7A ENSG00000005194 CIAPIN1 cytokine induced apoptosis inhibitor 1 ENSG00000005381 MPO myeloperoxidase ENSG00000005486 RHBDD2 rhomboid domain containing 2 ENSG00000005884 ITGA3 integrin subunit alpha 3 ENSG00000006016 CRLF1 cytokine receptor like -

Is Mitochondrial DNA Depletion Involved in Alzheimer's Disease?
European Journal of Human Genetics (2001) 9, 279 ± 285 ã 2001 Nature Publishing Group All rights reserved 1018-4813/01 $15.00 www.nature.com/ejhg ARTICLE Is mitochondrial DNA depletion involved in Alzheimer's disease? BenjamõÂn RodrõÂguez-Santiago1, Jordi Casademont2 and Virginia Nunes*,1 1Medical and Molecular Genetics Center, Institut de Recerca OncoloÁgica,Barcelona,Spain;2Muscle Research Group, Hospital ClõÂnic, IDIBAPS, Universitat de Barcelona, Barcelona, Spain Several studies have suggested that mitochondrial metabolism disturbances and mitochondrial DNA (mtDNA) abnormalities may contribute to the progression of the pathology of Alzheimer's disease (AD). In this study we have investigated whether the amount of mtDNA is modified in different brain regions (cerebellum, hippocampus and frontal cortex) of confirmed AD necropsies and in blood of living AD patients. We used a real-time PCR method to analyse the mtDNA relative abundance in brain regions from 12 AD and seven controls and from a group of blood samples (17 living AD patients and 11 controls). MtDNA from blood samples together with hippocampus and cerebellum brain areas did not show differences between controls and AD. However, AD patients showed a 28% decrease in the amount of mtDNA in the frontal cortex when compared to controls for this specific area. Since frontal cortex is a severely affected region in AD, our results support the hypothesis that mitochondrial defects may play a role in the pathogenesis of AD. European Journal of Human Genetics (2001) 9, 279 ± 285. Keywords: mtDNA; depletion; mitochondria; Alzheimer's disease; brain Introduction of developing the disease.3 Many genes associated with the Alzheimer's disease (AD) is one of the major causes of disease have been identified. -

Novel and Highly Recurrent Chromosomal Alterations in Se´Zary Syndrome
Research Article Novel and Highly Recurrent Chromosomal Alterations in Se´zary Syndrome Maarten H. Vermeer,1 Remco van Doorn,1 Remco Dijkman,1 Xin Mao,3 Sean Whittaker,3 Pieter C. van Voorst Vader,4 Marie-Jeanne P. Gerritsen,5 Marie-Louise Geerts,6 Sylke Gellrich,7 Ola So¨derberg,8 Karl-Johan Leuchowius,8 Ulf Landegren,8 Jacoba J. Out-Luiting,1 Jeroen Knijnenburg,2 Marije IJszenga,2 Karoly Szuhai,2 Rein Willemze,1 and Cornelis P. Tensen1 Departments of 1Dermatology and 2Molecular Cell Biology, Leiden University Medical Center, Leiden, the Netherlands; 3Department of Dermatology, St Thomas’ Hospital, King’s College, London, United Kingdom; 4Department of Dermatology, University Medical Center Groningen, Groningen, the Netherlands; 5Department of Dermatology, Radboud University Nijmegen Medical Center, Nijmegen, the Netherlands; 6Department of Dermatology, Gent University Hospital, Gent, Belgium; 7Department of Dermatology, Charite, Berlin, Germany; and 8Department of Genetics and Pathology, Rudbeck Laboratory, University of Uppsala, Uppsala, Sweden Abstract Introduction This study was designed to identify highly recurrent genetic Se´zary syndrome (Sz) is an aggressive type of cutaneous T-cell alterations typical of Se´zary syndrome (Sz), an aggressive lymphoma/leukemia of skin-homing, CD4+ memory T cells and is cutaneous T-cell lymphoma/leukemia, possibly revealing characterized by erythroderma, generalized lymphadenopathy, and pathogenetic mechanisms and novel therapeutic targets. the presence of neoplastic T cells (Se´zary cells) in the skin, lymph High-resolution array-based comparative genomic hybridiza- nodes, and peripheral blood (1). Sz has a poor prognosis, with a tion was done on malignant T cells from 20 patients. disease-specific 5-year survival of f24% (1). -

Delineation of C12orf65-Related Phenotypes: a Genotype&Ndash
European Journal of Human Genetics (2014) 22, 1019–1025 & 2014 Macmillan Publishers Limited All rights reserved 1018-4813/14 www.nature.com/ejhg ARTICLE Delineation of C12orf65-related phenotypes: a genotype–phenotype relationship Ronen Spiegel1,2,10, Hanna Mandel2,3,10, Ann Saada4, Issy Lerer4, Ayala Burger4, Avraham Shaag5, Stavit A Shalev1,2, Haneen Jabaly-Habib6, Dorit Goldsher7, John M Gomori8, Alex Lossos9, Orly Elpeleg5 and Vardiella Meiner*,4 C12orf65 participates in the process of mitochondrial translation and has been shown to be associated with a spectrum of phenotypes, including early onset optic atrophy, progressive encephalomyopathy, peripheral neuropathy, and spastic paraparesis.We used whole-genome homozygosity mapping as well as exome sequencing and targeted gene sequencing to identify novel C12orf65 disease-causing mutations in seven affected individuals originating from two consanguineous families. In four family members affected with childhood-onset optic atrophy accompanied by slowly progressive peripheral neuropathy and spastic paraparesis, we identified a homozygous frame shift mutation c.413_417 delAACAA, which predicts a truncated protein lacking the C-terminal portion. In the second family, we studied three affected individuals who presented with early onset optic atrophy, peripheral neuropathy, and spastic gait in addition to moderate intellectual disability. Muscle biopsy in two of the patients revealed decreased activities of the mitochondrial respiratory chain complexes I and IV. In these patients, we identified a homozygous splice mutation, g.21043 T4A (c.282 þ 2T4A) which leads to skipping of exon 2. Our study broadens the phenotypic spectrum of C12orf65 defects and highlights the triad of optic atrophy, axonal neuropathy and spastic paraparesis as its key clinical features.