First Profiling of Lysine Crotonylation of Myofilament Proteins and Ribosomal
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Targeted Genes and Methodology Details for Neuromuscular Genetic Panels
Targeted Genes and Methodology Details for Neuromuscular Genetic Panels Reference transcripts based on build GRCh37 (hg19) interrogated by Neuromuscular Genetic Panels Next-generation sequencing (NGS) and/or Sanger sequencing is performed Motor Neuron Disease Panel to test for the presence of a mutation in these genes. Gene GenBank Accession Number Regions of homology, high GC-rich content, and repetitive sequences may ALS2 NM_020919 not provide accurate sequence. Therefore, all reported alterations detected ANG NM_001145 by NGS are confirmed by an independent reference method based on laboratory developed criteria. However, this does not rule out the possibility CHMP2B NM_014043 of a false-negative result in these regions. ERBB4 NM_005235 Sanger sequencing is used to confirm alterations detected by NGS when FIG4 NM_014845 appropriate.(Unpublished Mayo method) FUS NM_004960 HNRNPA1 NM_031157 OPTN NM_021980 PFN1 NM_005022 SETX NM_015046 SIGMAR1 NM_005866 SOD1 NM_000454 SQSTM1 NM_003900 TARDBP NM_007375 UBQLN2 NM_013444 VAPB NM_004738 VCP NM_007126 ©2018 Mayo Foundation for Medical Education and Research Page 1 of 14 MC4091-83rev1018 Muscular Dystrophy Panel Muscular Dystrophy Panel Gene GenBank Accession Number Gene GenBank Accession Number ACTA1 NM_001100 LMNA NM_170707 ANO5 NM_213599 LPIN1 NM_145693 B3GALNT2 NM_152490 MATR3 NM_199189 B4GAT1 NM_006876 MYH2 NM_017534 BAG3 NM_004281 MYH7 NM_000257 BIN1 NM_139343 MYOT NM_006790 BVES NM_007073 NEB NM_004543 CAPN3 NM_000070 PLEC NM_000445 CAV3 NM_033337 POMGNT1 NM_017739 CAVIN1 NM_012232 POMGNT2 -

Impairments in Contractility and Cytoskeletal Organisation Cause Nuclear Defects in Nemaline Myopathy
bioRxiv preprint doi: https://doi.org/10.1101/518522; this version posted January 28, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Impairments in contractility and cytoskeletal organisation cause nuclear defects in nemaline myopathy Jacob A Ross1, Yotam Levy1, Michela Ripolone2, Justin S Kolb3, Mark Turmaine4, Mark Holt5, Maurizio Moggio2, Chiara Fiorillo6, Johan Lindqvist3, Nicolas Figeac5, Peter S Zammit5, Heinz Jungbluth5,7,8, John Vissing9, Nanna Witting9, Henk Granzier3, Edmar Zanoteli10, Edna C Hardeman11, Carina Wallgren- Pettersson12, Julien Ochala1,5. 1. Centre for Human & Applied Physiological Sciences, School of Basic & Medical Biosciences, Faculty of Life Sciences & Medicine, Guy’s Campus, King’s College London, SE1 1UL, UK 2. Neuromuscular and Rare Diseases Unit, Department of Neuroscience, Fondazione IRCCS Ca' Granda, Ospedale Maggiore Policlinico, Milan 20122, Italy 3. Department of Cellular and Molecular Medicine, University of Arizona, Tucson, Arizona, 85721, USA 4. Division of Biosciences, University College London, Gower Street, London WC1E 6BT, UK 5. Randall Centre for Cell and Molecular Biophysics, School of Basic & Medical Biosciences, Faculty of Life Sciences & Medicine, Guy’s Campus, King’s College London, SE1 1UL, UK 6. Molecular Medicine, IRCCS Fondazione Stella Maris, Pisa and Department of Neuroscience, Rehabilitation, Ophthalmology, Genetics, Maternal and Child Health, University of Genova, Genoa, Italy 7. Department of Paediatric Neurology, Neuromuscular Service, Evelina's Children Hospital, Guy's and St Thomas' Hospital National Health Service Foundation Trust, London, SE1 9RT, UK 8. Department of Basic and Clinical Neuroscience, Institute of Psychiatry, Psychology & Neuroscience, King's College, London, SE1 1UL, UK 9. -

Analysis of Gene Expression Data for Gene Ontology
ANALYSIS OF GENE EXPRESSION DATA FOR GENE ONTOLOGY BASED PROTEIN FUNCTION PREDICTION A Thesis Presented to The Graduate Faculty of The University of Akron In Partial Fulfillment of the Requirements for the Degree Master of Science Robert Daniel Macholan May 2011 ANALYSIS OF GENE EXPRESSION DATA FOR GENE ONTOLOGY BASED PROTEIN FUNCTION PREDICTION Robert Daniel Macholan Thesis Approved: Accepted: _______________________________ _______________________________ Advisor Department Chair Dr. Zhong-Hui Duan Dr. Chien-Chung Chan _______________________________ _______________________________ Committee Member Dean of the College Dr. Chien-Chung Chan Dr. Chand K. Midha _______________________________ _______________________________ Committee Member Dean of the Graduate School Dr. Yingcai Xiao Dr. George R. Newkome _______________________________ Date ii ABSTRACT A tremendous increase in genomic data has encouraged biologists to turn to bioinformatics in order to assist in its interpretation and processing. One of the present challenges that need to be overcome in order to understand this data more completely is the development of a reliable method to accurately predict the function of a protein from its genomic information. This study focuses on developing an effective algorithm for protein function prediction. The algorithm is based on proteins that have similar expression patterns. The similarity of the expression data is determined using a novel measure, the slope matrix. The slope matrix introduces a normalized method for the comparison of expression levels throughout a proteome. The algorithm is tested using real microarray gene expression data. Their functions are characterized using gene ontology annotations. The results of the case study indicate the protein function prediction algorithm developed is comparable to the prediction algorithms that are based on the annotations of homologous proteins. -

Genequery™ Human Skeletal Muscle Contraction And
GeneQuery™ Human Skeletal Muscle Contraction and Muscular Dystrophies qPCR Array Kit (GQH-SKC) Catalog #GK065 Product Description ScienCell's GeneQuery™ Human Skeletal Muscle Contraction and Muscular Dystrophies qPCR Array (GQH-SKC) profiles 88 key genes involved in the contraction of skeletal muscle. There are three major muscle types in the human body: skeletal, cardiac, and smooth. Skeletal muscle is a striated muscle that relaxes and contracts to generate movement and force in the body. Of the three main muscle types, it is the only one under voluntary control. Neuromuscular diseases involving skeletal muscle dysfunction are very diverse and range from myopathies to muscular dystrophies. Below are brief examples of how included genes may be grouped according to their function in skeletal muscle biology: • Myogenesis: MYOD1, MYOG, PAX3, UTRN, MYF6, MDTN • Contractility: MYH2, SLC2A4, LMNA, DYSF, APT2A1 • Sarcomere Assembly : NEB, TRIM63, TTN, CAPN3, ACTA1 • Muscle Atrophy: NOS2, TRIM63, FOXO1, MAPK8, AKT1 • Muscular Dystrophy Development: POMT2, FKRP, ANO5, FRG1, SGCB Note : all gene names follow their official symbols by the Human Genome Organization Gene Nomenclature Committee (HGNC). GeneQuery™ qPCR array kits are qPCR ready in a 96-well plate format, with each well containing one primer set that can specifically recognize and efficiently amplify a target gene's cDNA. The carefully designed primers ensure that: (i) the optimal annealing temperature in qPCR analysis is 65°C (with 2 mM Mg 2+ , and no DMSO); (ii) the primer set recognizes all known transcript variants of target gene, unless otherwise indicated; and (iii) only one gene is amplified. Each primer set has been validated by qPCR with melt curve analysis, and gel electrophoresis. -

Defining Functional Interactions During Biogenesis of Epithelial Junctions
ARTICLE Received 11 Dec 2015 | Accepted 13 Oct 2016 | Published 6 Dec 2016 | Updated 5 Jan 2017 DOI: 10.1038/ncomms13542 OPEN Defining functional interactions during biogenesis of epithelial junctions J.C. Erasmus1,*, S. Bruche1,*,w, L. Pizarro1,2,*, N. Maimari1,3,*, T. Poggioli1,w, C. Tomlinson4,J.Lees5, I. Zalivina1,w, A. Wheeler1,w, A. Alberts6, A. Russo2 & V.M.M. Braga1 In spite of extensive recent progress, a comprehensive understanding of how actin cytoskeleton remodelling supports stable junctions remains to be established. Here we design a platform that integrates actin functions with optimized phenotypic clustering and identify new cytoskeletal proteins, their functional hierarchy and pathways that modulate E-cadherin adhesion. Depletion of EEF1A, an actin bundling protein, increases E-cadherin levels at junctions without a corresponding reinforcement of cell–cell contacts. This unexpected result reflects a more dynamic and mobile junctional actin in EEF1A-depleted cells. A partner for EEF1A in cadherin contact maintenance is the formin DIAPH2, which interacts with EEF1A. In contrast, depletion of either the endocytic regulator TRIP10 or the Rho GTPase activator VAV2 reduces E-cadherin levels at junctions. TRIP10 binds to and requires VAV2 function for its junctional localization. Overall, we present new conceptual insights on junction stabilization, which integrate known and novel pathways with impact for epithelial morphogenesis, homeostasis and diseases. 1 National Heart and Lung Institute, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. 2 Computing Department, Imperial College London, London SW7 2AZ, UK. 3 Bioengineering Department, Faculty of Engineering, Imperial College London, London SW7 2AZ, UK. 4 Department of Surgery & Cancer, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. -

Human Periprostatic Adipose Tissue: Secretome from Patients With
CANCER GENOMICS & PROTEOMICS 16 : 29-58 (2019) doi:10.21873/cgp.20110 Human Periprostatic Adipose Tissue: Secretome from Patients With Prostate Cancer or Benign Prostate Hyperplasia PAULA ALEJANDRA SACCA 1, OSVALDO NÉSTOR MAZZA 2, CARLOS SCORTICATI 2, GONZALO VITAGLIANO 3, GABRIEL CASAS 4 and JUAN CARLOS CALVO 1,5 1Institute of Biology and Experimental Medicine (IBYME), CONICET, Buenos Aires, Argentina; 2Department of Urology, School of Medicine, University of Buenos Aires, Clínical Hospital “José de San Martín”, Buenos Aires, Argentina; 3Department of Urology, Deutsches Hospital, Buenos Aires, Argentina; 4Department of Pathology, Deutsches Hospital, Buenos Aires, Argentina; 5Department of Biological Chemistry, School of Exact and Natural Sciences, University of Buenos Aires, Buenos Aires, Argentina Abstract. Background/Aim: Periprostatic adipose tissue Prostate cancer (PCa) is the second most common cancer in (PPAT) directs tumour behaviour. Microenvironment secretome men worldwide. While most men have indolent disease, provides information related to its biology. This study was which can be treated properly, the problem consists in performed to identify secreted proteins by PPAT, from both reliably distinguishing between indolent and aggressive prostate cancer and benign prostate hyperplasia (BPH) disease. Evidence shows that the microenvironment affects patients. Patients and Methods: Liquid chromatography-mass tumour behavior. spectrometry-based proteomic analysis was performed in Adipose tissue microenvironment is now known to direct PPAT-conditioned media (CM) from patients with prostate tumour growth, invasion and metastases (1, 2). Adipose cancer (CMs-T) (stage T3: CM-T3, stage T2: CM-T2) or tissue is adjacent to the prostate gland and the site of benign disease (CM-BPH). Results: The highest number and invasion of PCa. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplementary Table 3 Complete List of RNA-Sequencing Analysis of Gene Expression Changed by ≥ Tenfold Between Xenograft and Cells Cultured in 10%O2
Supplementary Table 3 Complete list of RNA-Sequencing analysis of gene expression changed by ≥ tenfold between xenograft and cells cultured in 10%O2 Expr Log2 Ratio Symbol Entrez Gene Name (culture/xenograft) -7.182 PGM5 phosphoglucomutase 5 -6.883 GPBAR1 G protein-coupled bile acid receptor 1 -6.683 CPVL carboxypeptidase, vitellogenic like -6.398 MTMR9LP myotubularin related protein 9-like, pseudogene -6.131 SCN7A sodium voltage-gated channel alpha subunit 7 -6.115 POPDC2 popeye domain containing 2 -6.014 LGI1 leucine rich glioma inactivated 1 -5.86 SCN1A sodium voltage-gated channel alpha subunit 1 -5.713 C6 complement C6 -5.365 ANGPTL1 angiopoietin like 1 -5.327 TNN tenascin N -5.228 DHRS2 dehydrogenase/reductase 2 leucine rich repeat and fibronectin type III domain -5.115 LRFN2 containing 2 -5.076 FOXO6 forkhead box O6 -5.035 ETNPPL ethanolamine-phosphate phospho-lyase -4.993 MYO15A myosin XVA -4.972 IGF1 insulin like growth factor 1 -4.956 DLG2 discs large MAGUK scaffold protein 2 -4.86 SCML4 sex comb on midleg like 4 (Drosophila) Src homology 2 domain containing transforming -4.816 SHD protein D -4.764 PLP1 proteolipid protein 1 -4.764 TSPAN32 tetraspanin 32 -4.713 N4BP3 NEDD4 binding protein 3 -4.705 MYOC myocilin -4.646 CLEC3B C-type lectin domain family 3 member B -4.646 C7 complement C7 -4.62 TGM2 transglutaminase 2 -4.562 COL9A1 collagen type IX alpha 1 chain -4.55 SOSTDC1 sclerostin domain containing 1 -4.55 OGN osteoglycin -4.505 DAPL1 death associated protein like 1 -4.491 C10orf105 chromosome 10 open reading frame 105 -4.491 -

Transcriptional Control of Tissue-Resident Memory T Cell Generation
Transcriptional control of tissue-resident memory T cell generation Filip Cvetkovski Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2019 © 2019 Filip Cvetkovski All rights reserved ABSTRACT Transcriptional control of tissue-resident memory T cell generation Filip Cvetkovski Tissue-resident memory T cells (TRM) are a non-circulating subset of memory that are maintained at sites of pathogen entry and mediate optimal protection against reinfection. Lung TRM can be generated in response to respiratory infection or vaccination, however, the molecular pathways involved in CD4+TRM establishment have not been defined. Here, we performed transcriptional profiling of influenza-specific lung CD4+TRM following influenza infection to identify pathways implicated in CD4+TRM generation and homeostasis. Lung CD4+TRM displayed a unique transcriptional profile distinct from spleen memory, including up-regulation of a gene network induced by the transcription factor IRF4, a known regulator of effector T cell differentiation. In addition, the gene expression profile of lung CD4+TRM was enriched in gene sets previously described in tissue-resident regulatory T cells. Up-regulation of immunomodulatory molecules such as CTLA-4, PD-1, and ICOS, suggested a potential regulatory role for CD4+TRM in tissues. Using loss-of-function genetic experiments in mice, we demonstrate that IRF4 is required for the generation of lung-localized pathogen-specific effector CD4+T cells during acute influenza infection. Influenza-specific IRF4−/− T cells failed to fully express CD44, and maintained high levels of CD62L compared to wild type, suggesting a defect in complete differentiation into lung-tropic effector T cells. -

RESEARCH ARTICLE Body Weight-Dependent Troponin T Alternative Splicing Is Evolutionarily Conserved from Insects to Mammals and I
1523 The Journal of Experimental Biology 214, 1523-1532 © 2011. Published by The Company of Biologists Ltd doi:10.1242/jeb.051763 RESEARCH ARTICLE Body weight-dependent troponin T alternative splicing is evolutionarily conserved from insects to mammals and is partially impaired in skeletal muscle of obese rats Rudolf J. Schilder1,*, Scot R. Kimball1, James H. Marden2 and Leonard S. Jefferson1 1Department of Cellular and Molecular Physiology, The Pennsylvania State University College of Medicine, 500 University Drive, Hershey, PA 17033, USA and 2Department of Biology, The Pennsylvania State University, 208 Mueller Lab, University Park, PA 16802, USA *Author for correspondence ([email protected]) Accepted 19 January 2011 SUMMARY Do animals know at a physiological level how much they weigh, and, if so, do they make homeostatic adjustments in response to changes in body weight? Skeletal muscle is a likely tissue for such plasticity, as weight-bearing muscles receive mechanical feedback regarding body weight and consume ATP in order to generate forces sufficient to counteract gravity. Using rats, we examined how variation in body weight affected alternative splicing of fast skeletal muscle troponin T (Tnnt3), a component of the thin filament that regulates the actin–myosin interaction during contraction and modulates force output. In response to normal growth and experimental body weight increases, alternative splicing of Tnnt3 in rat gastrocnemius muscle was adjusted in a quantitative fashion. The response depended on weight per se, as externally attached loads had the same effect as an equal change in actual body weight. Examining the association between Tnnt3 alternative splicing and ATP consumption rate, we found that the Tnnt3 splice form profile had a significant association with nocturnal energy expenditure, independently of effects of weight. -

TNNT3 Rabbit Pab
Leader in Biomolecular Solutions for Life Science TNNT3 Rabbit pAb Catalog No.: A15323 Basic Information Background Catalog No. The binding of Ca(2+) to the trimeric troponin complex initiates the process of muscle A15323 contraction. Increased Ca(2+) concentrations produce a conformational change in the troponin complex that is transmitted to tropomyosin dimers situated along actin Observed MW filaments. The altered conformation permits increased interaction between a myosin 37kDa head and an actin filament which, ultimately, produces a muscle contraction. The troponin complex has protein subunits C, I, and T. Subunit C binds Ca(2+) and subunit I Calculated MW binds to actin and inhibits actin-myosin interaction. Subunit T binds the troponin 29kDa/30kDa/31kDa complex to the tropomyosin complex and is also required for Ca(2+)-mediated activation of actomyosin ATPase activity. There are 3 different troponin T genes that Category encode tissue-specific isoforms of subunit T for fast skeletal-, slow skeletal-, and cardiac-muscle. This gene encodes fast skeletal troponin T protein; also known as Primary antibody troponin T type 3. Alternative splicing results in multiple transcript variants encoding additional distinct troponin T type 3 isoforms. A developmentally regulated switch Applications between fetal/neonatal and adult troponin T type 3 isoforms occurs. Additional splice WB variants have been described but their biological validity has not been established. Mutations in this gene may cause distal arthrogryposis multiplex congenita type 2B Cross-Reactivity (DA2B). Mouse, Rat Recommended Dilutions Immunogen Information WB 1:200 - 1:2000 Gene ID Swiss Prot 7140 P45378 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 147-256 of human TNNT3 (NP_001036246.1). -
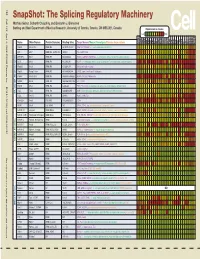
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus