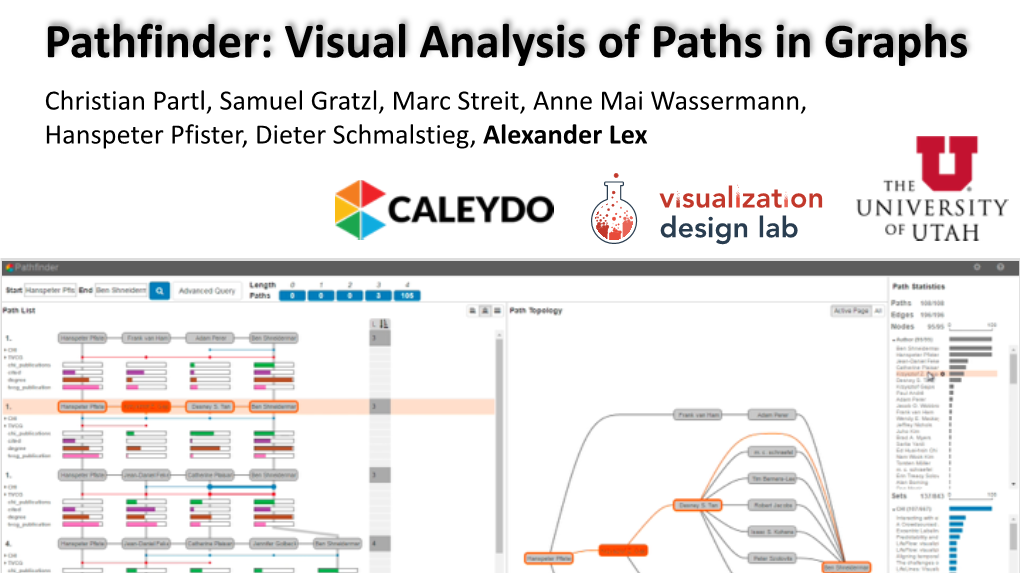
Pathfinder: Visual Analysis of Paths in Graphs
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Ways of Seeing Data: a Survey of Fields of Visualization
• Ways of seeing data: • A survey of fields of visualization • Gordon Kindlmann • [email protected] Part of the talk series “Show and Tell: Visualizing the Life of the Mind” • Nov 19, 2012 http://rcc.uchicago.edu/news/show_and_tell_abstracts.html 2012 Presidential Election REPUBLICAN DEMOCRAT http://www.npr.org/blogs/itsallpolitics/2012/11/01/163632378/a-campaign-map-morphed-by-money 2012 Presidential Election http://gizmodo.com/5960290/this-is-the-real-political-map-of-america-hint-we-are-not-that-divided 2012 Presidential Election http://www.npr.org/blogs/itsallpolitics/2012/11/01/163632378/a-campaign-map-morphed-by-money 2012 Presidential Election http://www.npr.org/blogs/itsallpolitics/2012/11/01/163632378/a-campaign-map-morphed-by-money 2012 Presidential Election http://www.npr.org/blogs/itsallpolitics/2012/11/01/163632378/a-campaign-map-morphed-by-money Clarifying distortions Tube map from 1908 http://www.20thcenturylondon.org.uk/beck-henry-harry http://briankerr.wordpress.com/2009/06/08/connections/ http://en.wikipedia.org/wiki/Harry_Beck Clarifying distortions http://www.20thcenturylondon.org.uk/beck-henry-harry Harry Beck 1933 http://briankerr.wordpress.com/2009/06/08/connections/ http://en.wikipedia.org/wiki/Harry_Beck Clarifying distortions Clarifying distortions Joachim Böttger, Ulrik Brandes, Oliver Deussen, Hendrik Ziezold, “Map Warping for the Annotation of Metro Maps” IEEE Computer Graphics and Applications, 28(5):56-65, 2008 Maps reflect conventions, choices, and priorities “A single map is but one of an indefinitely large -

Point-Based Computer Graphics
Point-Based Computer Graphics Eurographics 2002 Tutorial T6 Organizers Markus Gross ETH Zürich Hanspeter Pfister MERL, Cambridge Presenters Marc Alexa TU Darmstadt Markus Gross ETH Zürich Mark Pauly ETH Zürich Hanspeter Pfister MERL, Cambridge Marc Stamminger Bauhaus-Universität Weimar Matthias Zwicker ETH Zürich Contents Tutorial Schedule................................................................................................2 Presenters Biographies........................................................................................3 Presenters Contact Information ..........................................................................4 References...........................................................................................................5 Project Pages.......................................................................................................6 Tutorial Schedule 8:30-8:45 Introduction (M. Gross) 8:45-9:45 Point Rendering (M. Zwicker) 9:45-10:00 Acquisition of Point-Sampled Geometry and Appearance I (H. Pfister) 10:00-10:30 Coffee Break 10:30-11:15 Acquisition of Point-Sampled Geometry and Appearance II (H. Pfister) 11:15-12:00 Dynamic Point Sampling (M. Stamminger) 12:00-14:00 Lunch 14:00-15:00 Point-Based Surface Representations (M. Alexa) 15:00-15:30 Spectral Processing of Point-Sampled Geometry (M. Gross) 15:30-16:00 Coffee Break 16:00-16:30 Efficient Simplification of Point-Sampled Geometry (M. Pauly) 16:30-17:15 Pointshop3D: An Interactive System for Point-Based Surface Editing (M. Pauly) 17:15-17:30 Discussion (all) 2 Presenters Biographies Dr. Markus Gross is a professor of computer science and the director of the computer graphics laboratory of the Swiss Federal Institute of Technology (ETH) in Zürich. He received a degree in electrical and computer engineering and a Ph.D. on computer graphics and image analysis, both from the University of Saarbrucken, Germany. From 1990 to 1994 Dr. Gross was with the Computer Graphics Center in Darmstadt, where he established and directed the Visual Computing Group. -

Mike Roberts Curriculum Vitae
Mike Roberts +1 (650) 924–7168 [email protected] mikeroberts3000.github.io Research Using photorealistic synthetic data for computer vision; motion planning, trajectory optimization, Interests and control methods for robotics; reconstructing 3D scenes from images; continuous and discrete optimization; submodular optimization; software tools and algorithms for creativity support Education Stanford University Stanford, California Ph.D. Computer Science 2012–2019 Advisor: Pat Hanrahan Dissertation: Trajectory Optimization Methods for Drone Cameras Harvard University Cambridge, Massachusetts Visiting Research Fellow Summer 2013 Advisor: Hanspeter Pfister University of Calgary Calgary, Canada M.S. Computer Science 2010 University of Calgary Calgary, Canada B.S. Computer Science 2007 Employment Intel Labs Seattle, Washington Research Scientist 2021– Mentor: Vladlen Koltun Apple Seattle, Washington Research Scientist 2018–2021 Microsoft Research Redmond, Washington Research Intern Summer 2016, 2017 Mentors: Neel Joshi, Sudipta Sinha Skydio Redwood City, California Research Intern Spring 2016 Mentors: Adam Bry, Frank Dellaert Udacity Mountain View, California Course Developer, Introduction to Parallel Computing 2012–2013 Instructors: John Owens, David Luebke Harvard University Cambridge, Massachusetts Research Fellow 2010–2012 Advisor: Hanspeter Pfister NVIDIA Austin, Texas Developer Tools Programmer Intern Summer 2009 Radical Entertainment Vancouver, Canada Graphics Programmer Intern 2005–2006 Honors And Featured in the Highlights of -

Mike Roberts Curriculum Vitae
Mike Roberts Union Street, Suite , Seattle, Washington, + () / [email protected] http://mikeroberts.github.io Research Using photorealistic synthetic data for computer vision; motion planning, trajectory optimization, Interests and control methods for robotics; reconstructing D scenes from images; continuous and discrete optimization; submodular optimization; software tools and algorithms for creativity support. Education Stanford University Stanford, California Ph.D. Computer Science – Advisor: Pat Hanrahan Dissertation: Trajectory Optimization Methods for Drone Cameras Harvard University Cambridge, Massachusetts Visiting Research Fellow Summer John A. Paulson School of Engineering and Applied Sciences Advisor: Hanspeter Pfister University of Calgary Calgary, Canada M.S. Computer Science University of Calgary Calgary, Canada B.S. Computer Science Employment Apple Seattle, Washington Research Scientist – Microsoft Research Redmond, Washington Research Intern Summer , Advisors: Neel Joshi, Sudipta Sinha Skydio Redwood City, California Research Intern Spring Mentors: Adam Bry, Frank Dellaert Udacity Mountain View, California Course Developer, Introduction to Parallel Computing – Instructors: John Owens, David Luebke Harvard University Cambridge, Massachusetts Research Fellow, John A. Paulson School of Engineering and Applied Sciences – Advisor: Hanspeter Pfister NVIDIA Austin, Texas Developer Tools Programmer Intern Summer Radical Entertainment Vancouver, Canada Graphics Programmer Intern – Honors And Featured in the Highlights -

Monocular Neural Image Based Rendering with Continuous View Control
Monocular Neural Image Based Rendering with Continuous View Control Xu Chen∗ Jie Song∗ Otmar Hilliges AIT Lab, ETH Zurich {xuchen, jsong, otmar.hilliges}@inf.ethz.ch ... ... Figure 1: Interactive novel view synthesis: Given a single source view our approach can generate a continuous sequence of geometrically accurate novel views under fine-grained control. Top: Given a single street-view like input, a user may specify a continuous camera trajectory and our system generates the corresponding views in real-time. Bottom: An unseen hi-res internet image is used to synthesize novel views, while the camera is controlled interactively. Please refer to our project homepagey. Abstract like to view products interactively in 3D rather than from We propose a method to produce a continuous stream discrete view angles. Likewise in map applications it is of novel views under fine-grained (e.g., 1◦ step-size) cam- desirable to explore the vicinity of street-view like images era control at interactive rates. A novel learning pipeline beyond the position at which the photograph was taken. This determines the output pixels directly from the source color. is often not possible because either only 2D imagery exists, Injecting geometric transformations, including perspective or because storing and rendering of full 3D information does projection, 3D rotation and translation into the network not scale. To overcome this limitation we study the problem forces implicit reasoning about the underlying geometry. of interactive view synthesis with 6-DoF view control, taking The latent 3D geometry representation is compact and mean- only a single image as input. We propose a method that ingful under 3D transformation, being able to produce geo- can produce a continuous stream of novel views under fine- grained (e.g., 1◦ step-size) camera control (see Fig.1). -

Thouis R. Jones [email protected]
Thouis R. Jones [email protected] Summary Computational biologist with 20+ years' experience, including software engineering, algorithm development, image analysis, and solving challenging biomedical research problems. Research Experience Connectomics and Dense Reconstruction of Neural Connectivity Harvard University School of Engineering and Applied Sciences (2012-Present) My research is on software for automatic dense reconstruction of neural connectivity. Using using high-resolution serial sectioning and electron microscopy, biologists are able to image nanometer-scale details of neural structure. This data is extremely large (1 petabyte / cubic mm). The tools I develop in collaboration with the members of the Connectomics project allow biologists to analyze and explore this data. Image analysis, high-throughput screening, & large-scale data analysis Curie Institute, Department of Translational Research (2010 - 2012) Broad Institute of MIT and Harvard, Imaging Platform (2007 - 2010) My research focused on image and data analysis for high-content screening (HCS) and large-scale machine learning applied to HCS data. My work emphasized making powerful analysis methods available to the biological community in user-friendly tools. As part of this effort, I cofounded the open-source CellProfiler project with Dr. Anne Carpenter (director of the Imaging Platform at the Broad Institute). PhD Research: High-throughput microscopy and RNAi to reveal gene function. My doctoral research focused on methods for determining gene function using high- throughput microscopy, living cell microarrays, and RNA interference. To accomplish this goal, my collaborators and I designed and released the first open-source high- throughput cell image analysis software, CellProfiler (www.cellprofiler.org). We also developed advanced data mining methods for high-content screens. -

Human-AI Collaboration for Natural Language Generation with Interpretable Neural Networks
Human-AI Collaboration for Natural Language Generation with Interpretable Neural Networks a dissertation presented by Sebastian Gehrmann to The John A. Paulson School of Engineering and Applied Sciences in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the subject of Computer Science Harvard University Cambridge, Massachusetts May 2020 ©2019 – Sebastian Gehrmann Creative Commons Attribution License 4.0. You are free to share and adapt these materials for any purpose if you give appropriate credit and indicate changes. Thesis advisors: Barbara J. Grosz, Alexander M. Rush Sebastian Gehrmann Human-AI Collaboration for Natural Language Generation with Interpretable Neural Networks Abstract Using computers to generate natural language from information (NLG) requires approaches that plan the content and structure of the text and actualize it in fuent and error-free language. The typical approaches to NLG are data-driven, which means that they aim to solve the problem by learning from annotated data. Deep learning, a class of machine learning models based on neural networks, has become the standard data-driven NLG approach. While deep learning approaches lead to increased performance, they replicate undesired biases from the training data and make inexplicable mistakes. As a result, the outputs of deep learning NLG models cannot be trusted. Wethus need to develop ways in which humans can provide oversight over model outputs and retain their agency over an otherwise automated writing task. This dissertation argues that to retain agency over deep learning NLG models, we need to design them as team members instead of autonomous agents. We can achieve these team member models by considering the interaction design as an integral part of the machine learning model development. -

F I N a L P R O G R a M 1998
1998 FINAL PROGRAM October 18 • October 23, 1998 Sheraton Imperial Hotel Research Triangle Park, NC THE INSTITUTE OF ELECTRICAL IEEE IEEE VISUALIZATION 1998 & ELECTRONICS ENGINEERS, INC. COMPUTER IEEE SOCIETY Sponsored by IEEE Computer Society Technical Committee on Computer Graphics In Cooperation with ACM/SIGGRAPH Sessions include Real-time Volume Rendering, Terrain Visualization, Flow Visualization, Surfaces & Level-of-Detail Techniques, Feature Detection & Visualization, Medical Visualization, Multi-Dimensional Visualization, Flow & Streamlines, Isosurface Extraction, Information Visualization, 3D Modeling & Visualization, Multi-Source Data Analysis Challenges, Interactive Visualization/VR/Animation, Terrain & Large Data Visualization, Isosurface & Volume Rendering, Simplification, Marine Data Visualization, Tensor/Flow, Key Problems & Thorny Issues, Image-based Techniques and Volume Analysis, Engineering & Design, Texturing and Rendering, Art & Visualization Get complete, up-to-date listings of program information from URL: http://www.erc.msstate.edu/vis98 http://davinci.informatik.uni-kl.de/Vis98 Volvis URL: http://www.erc.msstate.edu/volvis98 InfoVis URL: http://www.erc.msstate.edu/infovis98 or contact: Theresa-Marie Rhyne, Lockheed Martin/U.S. EPA Sci Vis Center, 919-541-0207, [email protected] Robert Moorhead, Mississippi State University, 601-325-2850, [email protected] Direct Vehicle Loading Access S Dock Salon Salon Sheraton Imperial VII VI Imperial Convention Center HOTEL & CONVENTION CENTER Convention Center Phones -

Compositional Gan: Learning Conditional Image Composition
Under review as a conference paper at ICLR 2019 COMPOSITIONAL GAN: LEARNING CONDITIONAL IMAGE COMPOSITION Anonymous authors Paper under double-blind review ABSTRACT Generative Adversarial Networks (GANs) can produce images of surprising com- plexity and realism, but are generally structured to sample from a single latent source ignoring the explicit spatial interaction between multiple entities that could be present in a scene. Capturing such complex interactions between different objects in the world, including their relative scaling, spatial layout, occlusion, or viewpoint transformation is a challenging problem. In this work, we propose to model object composition in a GAN framework as a self-consistent composition- decomposition network. Our model is conditioned on the object images from their marginal distributions and can generate a realistic image from their joint distribu- tion. We evaluate our model through qualitative experiments and user evaluations in scenarios when either paired or unpaired examples for the individual object images and the joint scenes are given during training. Our results reveal that the learned model captures potential interactions between the two object domains given as input to output new instances of composed scene at test time in a reasonable fashion. 1I NTRODUCTION Generative Adversarial Networks (GANs) have emerged as a powerful method for generating images conditioned on a given input. The input cue could be in the form of an image (Isola et al., 2017; Zhu et al., 2017; Liu et al., 2017; Azadi et al., 2017; Wang et al., 2017; Pathak et al., 2016), a text phrase (Zhang et al., 2017; Reed et al., 2016b;a; Johnson et al., 2018) or a class label layout (Mirza & Osindero, 2014; Odena et al., 2016; Antoniou et al., 2017). -

Let's Gamble: Uncovering the Impact of Visualization on Risk Perception and Decision-Making
Let’s Gamble: Uncovering the Impact of Visualization on Risk Perception and Decision-Making Melanie Bancilhon Zhengliang Liu Alvitta Ottley [email protected] [email protected] [email protected] Washington Univ. in St. Louis ABSTRACT decision-making. However, it is reasonable to assume that Data visualizations are standard tools for assessing and com- encoding choices can influence the decisions people make. Re- municating risks. However, it is not always clear which de- searchers have shown that visualizations can impact speed [8, signs are optimal or how encoding choices might influence 37], accuracy [37], memorability [3,5], statistical reason- risk perception and decision-making. In this paper, we report ing [27, 30], and judgement [8, 11, 47]. These effects can the findings of a large-scale gambling game that immersed par- have significant repercussions, especially when decisions are ticipants in an environment where their actions impacted their life-altering. Still, a designer can represent the same data bonuses. Participants chose to either enter a draw or receive using different, yet equally theoretically valid visualization guaranteed monetary gains based on five common visualiza- designs [26], and it is sometimes difficult for designers to iden- tion designs. By measuring risk perception and observing tify a poor fit for the data [31]. Therefore, what we need is a decision-making, we showed that icon arrays tended to elicit thorough understanding of which designs lead to (in)accurate economically sound behavior. We also found that people were judgment and how probability distortions influence behavior. more likely to gamble when presented area proportioned tri- At the heart of decision-making with visualization is graphical angle and circle designs. -

Monocular Neural Image Based Rendering with Continuous View Control
Monocular Neural Image Based Rendering with Continuous View Control Xu Chen∗ Jie Song∗ Otmar Hilliges AIT Lab, ETH Zurich {xuchen, jsong, otmar.hilliges}@inf.ethz.ch ... ... Figure 1: Interactive novel view synthesis: Given a single source view our approach can generate a continuous sequence of geometrically accurate novel views under fine-grained control. Top: Given a single street-view like input, a user may specify a continuous camera trajectory and our system generates the corresponding views in real-time. Bottom: An unseen hi-res internet image is used to synthesize novel views, while the camera is controlled interactively. Please refer to our project homepage†. Abstract like to view products interactively in 3D rather than from We propose a method to produce a continuous stream discrete view angles. Likewise in map applications it is of novel views under fine-grained (e.g., 1◦ step-size) cam- desirable to explore the vicinity of street-view like images era control at interactive rates. A novel learning pipeline beyond the position at which the photograph was taken. This determines the output pixels directly from the source color. is often not possible because either only 2D imagery exists, Injecting geometric transformations, including perspective or because storing and rendering of full 3D information does projection, 3D rotation and translation into the network not scale. To overcome this limitation we study the problem forces implicit reasoning about the underlying geometry. of interactive view synthesis with 6-DoF view control, taking The latent 3D geometry representation is compact and mean- only a single image as input. We propose a method that ingful under 3D transformation, being able to produce geo- can produce a continuous stream of novel views under fine- ◦ metrically accurate views for both single objects and natural grained (e.g., 1 step-size) camera control (see Fig. -

GPU-Based Multi-Volume Rendering of Complex Data in Neuroscience and Neurosurgery
DISSERTATION GPU-based Multi-Volume Rendering of Complex Data in Neuroscience and Neurosurgery ausgeführt zum Zwecke der Erlangung des akademischen Grades eines Doktors der technischen Wissenschaften unter der Leitung von Ao.Univ.Prof. Dipl.-Ing. Dr.techn. Eduard Gröller, Institut E186 für Computergraphik und Algorithmen, und Dipl.-Ing. Dr.techn. Markus Hadwiger, VRVis Zentrum für Virtual Reality und Visualisierung, und Dipl.-Math. Dr.techn. Katja Bühler, VRVis Zentrum für Virtual Reality und Visualisierung, eingereicht an der Technischen Universität Wien, bei der Fakultät für Informatik, von Dipl.-Ing. (FH) Johanna Beyer, Matrikelnummer 0426197, Gablenzgasse 99/24 1150 Wien Wien, im Oktober 2009 DISSERTATION GPU-based Multi-Volume Rendering of Complex Data in Neuroscience and Neurosurgery Abstract Recent advances in image acquisition technology and its availability in the medical and bio-medical fields have lead to an unprecedented amount of high-resolution imaging data. However, the inherent complexity of this data, caused by its tremendous size, complex structure or multi-modality poses several challenges for current visualization tools. Recent developments in graphics hardware archi- tecture have increased the versatility and processing power of today’s GPUs to the point where GPUs can be considered parallel scientific computing devices. The work in this thesis builds on the current progress in image acquisition techniques and graphics hardware architecture to develop novel 3D visualization methods for the fields of neurosurgery and neuroscience. The first part of this thesis presents an application and framework for plan- ning of neurosurgical interventions. Concurrent GPU-based multi-volume ren- dering is used to visualize multiple radiological imaging modalities, delineating the patient’s anatomy, neurological function, and metabolic processes.