Chapter 2 the EGF-TM7 Family
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Edinburgh Research Explorer
Edinburgh Research Explorer International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list Citation for published version: Davenport, AP, Alexander, SPH, Sharman, JL, Pawson, AJ, Benson, HE, Monaghan, AE, Liew, WC, Mpamhanga, CP, Bonner, TI, Neubig, RR, Pin, JP, Spedding, M & Harmar, AJ 2013, 'International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands', Pharmacological reviews, vol. 65, no. 3, pp. 967-86. https://doi.org/10.1124/pr.112.007179 Digital Object Identifier (DOI): 10.1124/pr.112.007179 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Pharmacological reviews Publisher Rights Statement: U.S. Government work not protected by U.S. copyright General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 1521-0081/65/3/967–986$25.00 http://dx.doi.org/10.1124/pr.112.007179 PHARMACOLOGICAL REVIEWS Pharmacol Rev 65:967–986, July 2013 U.S. -

General Discussion
UvA-DARE (Digital Academic Repository) EGF-TM7 receptors: A diverse and still evolving family of receptors on the leukocyte surface Matmati, M. Publication date 2008 Link to publication Citation for published version (APA): Matmati, M. (2008). EGF-TM7 receptors: A diverse and still evolving family of receptors on the leukocyte surface. General rights It is not permitted to download or to forward/distribute the text or part of it without the consent of the author(s) and/or copyright holder(s), other than for strictly personal, individual use, unless the work is under an open content license (like Creative Commons). Disclaimer/Complaints regulations If you believe that digital publication of certain material infringes any of your rights or (privacy) interests, please let the Library know, stating your reasons. In case of a legitimate complaint, the Library will make the material inaccessible and/or remove it from the website. Please Ask the Library: https://uba.uva.nl/en/contact, or a letter to: Library of the University of Amsterdam, Secretariat, Singel 425, 1012 WP Amsterdam, The Netherlands. You will be contacted as soon as possible. UvA-DARE is a service provided by the library of the University of Amsterdam (https://dare.uva.nl) Download date:23 Sep 2021 Chapter 6 General Discussion GENERAL DISCUSSION Since their identification, starting about 15 years ago, a growing amount of data has accumulated about the structure, the expression, the ligands and, more recently, also the functional implications of EGF-TM7 receptors. Studies with antibody treatment and gene targeting in mice and antibody treatment of human cells in vitro, led to the implication of EGF-TM7 receptors in the trafficking of granulocytes, the generation of efferent antigen specific regulatory T cells and the potentiation of different granulocyte effector functions [1-4]. -

Flow Reagents Single Color Antibodies CD Chart
CD CHART CD N° Alternative Name CD N° Alternative Name CD N° Alternative Name Beckman Coulter Clone Beckman Coulter Clone Beckman Coulter Clone T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells CD1a T6, R4, HTA1 Act p n n p n n S l CD99 MIC2 gene product, E2 p p p CD223 LAG-3 (Lymphocyte activation gene 3) Act n Act p n CD1b R1 Act p n n p n n S CD99R restricted CD99 p p CD224 GGT (γ-glutamyl transferase) p p p p p p CD1c R7, M241 Act S n n p n n S l CD100 SEMA4D (semaphorin 4D) p Low p p p n n CD225 Leu13, interferon induced transmembrane protein 1 (IFITM1). p p p p p CD1d R3 Act S n n Low n n S Intest CD101 V7, P126 Act n p n p n n p CD226 DNAM-1, PTA-1 Act n Act Act Act n p n CD1e R2 n n n n S CD102 ICAM-2 (intercellular adhesion molecule-2) p p n p Folli p CD227 MUC1, mucin 1, episialin, PUM, PEM, EMA, DF3, H23 Act p CD2 T11; Tp50; sheep red blood cell (SRBC) receptor; LFA-2 p S n p n n l CD103 HML-1 (human mucosal lymphocytes antigen 1), integrin aE chain S n n n n n n n l CD228 Melanotransferrin (MT), p97 p p CD3 T3, CD3 complex p n n n n n n n n n l CD104 integrin b4 chain; TSP-1180 n n n n n n n p p CD229 Ly9, T-lymphocyte surface antigen p p n p n -
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
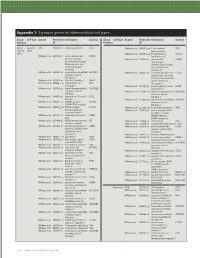
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al. -

ADGRE2-NTF Is Regulated by Site- Specific N-Glycosylation
www.nature.com/scientificreports OPEN Membrane-association of EMR2/ ADGRE2-NTF is regulated by site- specifc N-glycosylation Received: 19 December 2017 Yi-Shu Huang1,4, Nien-Yi Chiang1, Gin-Wen Chang1 & Hsi-Hsien Lin1,2,3 Accepted: 27 February 2018 The evolutionarily conserved adhesion G protein-coupled receptors (aGPCRs) play critical roles in Published: xx xx xxxx biological processes as diverse as brain development, cell polarity and innate immune functions. A defning feature of aGPCRs is the GPCR autoproteolysis inducing (GAIN) domain capable of self- catalytic cleavage, resulting in the generation of an extracellular N-terminal fragment (NTF) and a seven-transmembrane C-terminal fragment (CTF) involved in the cellular adhesion and signaling functions, respectively. Interestingly, two diferent NTF subtypes have previously been identifed, namely an NTF that couples non-covalently with the CTF and a membrane-associated NTF that tethers on cell surface independently. The two NTF subtypes are expected to regulate aGPCR signaling via distinct mechanisms however their molecular characteristics are largely unknown. Herein, the membrane-associated NTF of EMR2/ADGRE2 is investigated and found to be modifed by diferential N-glycosylation. The membrane association of EMR2-NTF occurs in post-ER compartments and site- specifc N-glycosylation in the GAIN domain is involved in modulating its membrane-association ability. Finally, a unique amphipathic α-helix in the GAIN domain is identifed as a putative membrane anchor of EMR2-NTF. These results provide novel insights into the complex interaction and activation mechanisms of aGPCRs. Characterized by a long extracellular domain (ECD) with cell-adhesion functions and a seven-transmembrane (7TM) domain with signaling functions, the adhesion G protein-coupled receptors (aGPCRs) have been impli- cated in diverse biological activities and human diseases1. -
![EMR3 (ADGRE3) Hamster Monoclonal Antibody [Clone ID: 3D7] Product Data](https://docslib.b-cdn.net/cover/2432/emr3-adgre3-hamster-monoclonal-antibody-clone-id-3d7-product-data-802432.webp)
EMR3 (ADGRE3) Hamster Monoclonal Antibody [Clone ID: 3D7] Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for AM01316FC-N EMR3 (ADGRE3) Hamster Monoclonal Antibody [Clone ID: 3D7] Product data: Product Type: Primary Antibodies Clone Name: 3D7 Applications: FC Recommended Dilution: Flow cytometry ( Neat - 1/10): use 10 µl of the suggested working dilution to label 10e6 cells in 100 µl. Reactivity: Human Host: Hamster Isotype: IgG Clonality: Monoclonal Immunogen: ARHO-EMR3-CD97 (EGF1) transfectants. Spleen cells from immunised Armenian hamster were fused with cells of the mouse SP2/0 myeloma cell line. Specificity: This antibody recognises epidermal growth factor (EGF) module-containing mucin-like hormone receptor 3 (EMR3), which is a 56kD member of the EGF-7 transmembrane (TM7) family of adhesion receptors. Formulation: PBS, pH 7.4, containing 0.09% Sodium Azide Label: FITC State: Liquid purified IgG Label: Fluorescein Isothiocyanate Isomer I Concentration: lot specific Purification: Affinity chromatography on Protein G Conjugation: FITC Storage: Store the antibody undiluted at 2-8°C for one month or (in aliquots) at -20°C for longer. Avoid repeated freezing and thawing. This product is photosensitive and should be protected from light. Stability: Shelf life: one year from despatch. Database Link: Entrez Gene 84658 Human Q9BY15 This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 EMR3 (ADGRE3) Hamster Monoclonal Antibody [Clone ID: 3D7] – AM01316FC-N Background: EMR3 is expressed at the cell surface as a heterodimer. -

An Overview on G Protein-Coupled Receptor-Induced Signal Transduction in Acute Myeloid Leukemia
An overview on G protein-coupled receptor-induced signal transduction in Acute Myeloid Leukemia 1* 1,3 4,5,6 2* Frode Selheim , Elise Aasebø , Catalina Ribas and Anna M. Aragay 1The Proteomics Unit at the University of Bergen, Department of Biomedicine, University of Bergen, Jonas Lies vei 91, 5020 Bergen, Norway; 3 Department of Clinical Science, University of Bergen, Jonas Lies vei 87, 5021 Bergen, Norway; [email protected]. 2Departamento de Biologia Celular. Instituto de Biología Molecular de Barcelona (IBMB-CSIC), Spanish National Research Council (CSIC), Baldiri i Reixac, 15, 08028 Barcelona, Spain; [email protected]. 4Departamento de Biología Molecular and Centro de Biología Molecular “Severo Ochoa” (UAM-CSIC), 28049 Madrid, Spain; 5Instituto de Investigación Sanitaria La Princesa, 28006 Madrid, Spain; 6CIBER de Enfermedades Cardiovasculares, ISCIII (CIBERCV), 28029 Madrid, Spain, [email protected] * Corresponding authors: Frode Selheim Adr: Jonas Lies vei 91, 5020 Bergen, Norway Email: [email protected], Tel:+4755586091 Anna M. Aragay Adr: Baldiri i Reixac, 15, 08028 Barcelona. Spain. E-mail: [email protected]; Tel.: +934098671 1 Abstract Background: Acute myeloid leukemia (AML) is a genetically heterogeneous disease characterized by uncontrolled proliferation of precursor myeloid-lineage cells in the bone marrow. AML is also characterized with patients with poor long-term survival outcomes due to relapse. Many efforts have been made to understand the biological heterogeneity of AML and the challenges to develop new therapies are therefore enormous. G protein-coupled receptors (GPCRs) are a large attractive drug targeted family of transmembrane proteins, and aberrant GPCR expression and GPCR-mediated signaling have been implicated in leukemogenesis of AML. -

Biased Signaling of G Protein Coupled Receptors (Gpcrs): Molecular Determinants of GPCR/Transducer Selectivity and Therapeutic Potential
Pharmacology & Therapeutics 200 (2019) 148–178 Contents lists available at ScienceDirect Pharmacology & Therapeutics journal homepage: www.elsevier.com/locate/pharmthera Biased signaling of G protein coupled receptors (GPCRs): Molecular determinants of GPCR/transducer selectivity and therapeutic potential Mohammad Seyedabadi a,b, Mohammad Hossein Ghahremani c, Paul R. Albert d,⁎ a Department of Pharmacology, School of Medicine, Bushehr University of Medical Sciences, Iran b Education Development Center, Bushehr University of Medical Sciences, Iran c Department of Toxicology–Pharmacology, School of Pharmacy, Tehran University of Medical Sciences, Iran d Ottawa Hospital Research Institute, Neuroscience, University of Ottawa, Canada article info abstract Available online 8 May 2019 G protein coupled receptors (GPCRs) convey signals across membranes via interaction with G proteins. Origi- nally, an individual GPCR was thought to signal through one G protein family, comprising cognate G proteins Keywords: that mediate canonical receptor signaling. However, several deviations from canonical signaling pathways for GPCR GPCRs have been described. It is now clear that GPCRs can engage with multiple G proteins and the line between Gprotein cognate and non-cognate signaling is increasingly blurred. Furthermore, GPCRs couple to non-G protein trans- β-arrestin ducers, including β-arrestins or other scaffold proteins, to initiate additional signaling cascades. Selectivity Biased Signaling Receptor/transducer selectivity is dictated by agonist-induced receptor conformations as well as by collateral fac- Therapeutic Potential tors. In particular, ligands stabilize distinct receptor conformations to preferentially activate certain pathways, designated ‘biased signaling’. In this regard, receptor sequence alignment and mutagenesis have helped to iden- tify key receptor domains for receptor/transducer specificity. -

Multi-Functionality of Proteins Involved in GPCR and G Protein Signaling: Making Sense of Structure–Function Continuum with In
Cellular and Molecular Life Sciences (2019) 76:4461–4492 https://doi.org/10.1007/s00018-019-03276-1 Cellular andMolecular Life Sciences REVIEW Multi‑functionality of proteins involved in GPCR and G protein signaling: making sense of structure–function continuum with intrinsic disorder‑based proteoforms Alexander V. Fonin1 · April L. Darling2 · Irina M. Kuznetsova1 · Konstantin K. Turoverov1,3 · Vladimir N. Uversky2,4 Received: 5 August 2019 / Revised: 5 August 2019 / Accepted: 12 August 2019 / Published online: 19 August 2019 © Springer Nature Switzerland AG 2019 Abstract GPCR–G protein signaling system recognizes a multitude of extracellular ligands and triggers a variety of intracellular signal- ing cascades in response. In humans, this system includes more than 800 various GPCRs and a large set of heterotrimeric G proteins. Complexity of this system goes far beyond a multitude of pair-wise ligand–GPCR and GPCR–G protein interactions. In fact, one GPCR can recognize more than one extracellular signal and interact with more than one G protein. Furthermore, one ligand can activate more than one GPCR, and multiple GPCRs can couple to the same G protein. This defnes an intricate multifunctionality of this important signaling system. Here, we show that the multifunctionality of GPCR–G protein system represents an illustrative example of the protein structure–function continuum, where structures of the involved proteins represent a complex mosaic of diferently folded regions (foldons, non-foldons, unfoldons, semi-foldons, and inducible foldons). The functionality of resulting highly dynamic conformational ensembles is fne-tuned by various post-translational modifcations and alternative splicing, and such ensembles can undergo dramatic changes at interaction with their specifc partners. -

Development and Validation of a Protein-Based Risk Score for Cardiovascular Outcomes Among Patients with Stable Coronary Heart Disease
Supplementary Online Content Ganz P, Heidecker B, Hveem K, et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA. doi: 10.1001/jama.2016.5951 eTable 1. List of 1130 Proteins Measured by Somalogic’s Modified Aptamer-Based Proteomic Assay eTable 2. Coefficients for Weibull Recalibration Model Applied to 9-Protein Model eFigure 1. Median Protein Levels in Derivation and Validation Cohort eTable 3. Coefficients for the Recalibration Model Applied to Refit Framingham eFigure 2. Calibration Plots for the Refit Framingham Model eTable 4. List of 200 Proteins Associated With the Risk of MI, Stroke, Heart Failure, and Death eFigure 3. Hazard Ratios of Lasso Selected Proteins for Primary End Point of MI, Stroke, Heart Failure, and Death eFigure 4. 9-Protein Prognostic Model Hazard Ratios Adjusted for Framingham Variables eFigure 5. 9-Protein Risk Scores by Event Type This supplementary material has been provided by the authors to give readers additional information about their work. Downloaded From: https://jamanetwork.com/ on 10/02/2021 Supplemental Material Table of Contents 1 Study Design and Data Processing ......................................................................................................... 3 2 Table of 1130 Proteins Measured .......................................................................................................... 4 3 Variable Selection and Statistical Modeling ........................................................................................ -

Datasheet: MCA2476A647 Product Details
Datasheet: MCA2476A647 Description: HAMSTER ANTI HUMAN EMR3:Alexa Fluor® 647 Specificity: EMR3 Format: ALEXA FLUOR® 647 Product Type: Monoclonal Antibody Clone: 3D7 Isotype: IgG Quantity: 100 TESTS/1ml Product Details Applications This product has been reported to work in the following applications. This information is derived from testing within our laboratories, peer-reviewed publications or personal communications from the originators. Please refer to references indicated for further information. For general protocol recommendations, please visit www.bio-rad-antibodies.com/protocols. Yes No Not Determined Suggested Dilution Flow Cytometry Neat - 1/10 Where this antibody has not been tested for use in a particular technique this does not necessarily exclude its use in such procedures. Suggested working dilutions are given as a guide only. It is recommended that the user titrates the antibody for use in their own system using appropriate negative/positive controls. Target Species Human Product Form Purified IgG conjugated to Alexa Fluor® 647 - liquid Max Ex/Em Fluorophore Excitation Max (nm) Emission Max (nm) Alexa Fluor®647 650 665 Preparation Purified IgG prepared by affinity chromatography on Protein G from tissue culture supernatant Buffer Solution Phosphate buffered saline Preservative 0.09% Sodium Azide Stabilisers 1% Bovine Serum Albumin Approx. Protein IgG concentration 0.05 mg/ml Concentrations Immunogen ARHO-EMR3-CD97 (EGF1) transfectants. External Database UniProt: Links Q9BY15 Related reagents Entrez Gene: Page 1 of 3 84658 EMR3 Related reagents Fusion Partners Spleen cells from immunised Armenian hamster were fused with cells of the mouse SP2/0 myeloma cell line. Specificity Hamster anti Human EMR3 antibody, clone 3D7 recognizes human epidermal growth factor (EGF) module-containing mucin-like hormone receptor 3 (EMR3), a ~56 kDa member of the EGF-7 transmembrane (TM7) family of adhesion receptors. -

Human CD Marker Chart Reviewed by HLDA1 Bdbiosciences.Com/Cdmarkers
BD Biosciences Human CD Marker Chart Reviewed by HLDA1 bdbiosciences.com/cdmarkers 23-12399-01 CD Alternative Name Ligands & Associated Molecules T Cell B Cell Dendritic Cell NK Cell Stem Cell/Precursor Macrophage/Monocyte Granulocyte Platelet Erythrocyte Endothelial Cell Epithelial Cell CD Alternative Name Ligands & Associated Molecules T Cell B Cell Dendritic Cell NK Cell Stem Cell/Precursor Macrophage/Monocyte Granulocyte Platelet Erythrocyte Endothelial Cell Epithelial Cell CD Alternative Name Ligands & Associated Molecules T Cell B Cell Dendritic Cell NK Cell Stem Cell/Precursor Macrophage/Monocyte Granulocyte Platelet Erythrocyte Endothelial Cell Epithelial Cell CD1a R4, T6, Leu6, HTA1 b-2-Microglobulin, CD74 + + + – + – – – CD93 C1QR1,C1qRP, MXRA4, C1qR(P), Dj737e23.1, GR11 – – – – – + + – – + – CD220 Insulin receptor (INSR), IR Insulin, IGF-2 + + + + + + + + + Insulin-like growth factor 1 receptor (IGF1R), IGF-1R, type I IGF receptor (IGF-IR), CD1b R1, T6m Leu6 b-2-Microglobulin + + + – + – – – CD94 KLRD1, Kp43 HLA class I, NKG2-A, p39 + – + – – – – – – CD221 Insulin-like growth factor 1 (IGF-I), IGF-II, Insulin JTK13 + + + + + + + + + CD1c M241, R7, T6, Leu6, BDCA1 b-2-Microglobulin + + + – + – – – CD178, FASLG, APO-1, FAS, TNFRSF6, CD95L, APT1LG1, APT1, FAS1, FASTM, CD95 CD178 (Fas ligand) + + + + + – – IGF-II, TGF-b latency-associated peptide (LAP), Proliferin, Prorenin, Plasminogen, ALPS1A, TNFSF6, FASL Cation-independent mannose-6-phosphate receptor (M6P-R, CIM6PR, CIMPR, CI- CD1d R3G1, R3 b-2-Microglobulin, MHC II CD222 Leukemia