Integrating Multi-Omics Data to Identify Cancer Driver Modules Supplementary Information
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Connections of Wnt Pathway Components with Cell Cycle and Centrosome: Side Effects Or a Hidden Logic?
Critical Reviews in Biochemistry and Molecular Biology ISSN: 1040-9238 (Print) 1549-7798 (Online) Journal homepage: http://www.tandfonline.com/loi/ibmg20 The connections of Wnt pathway components with cell cycle and centrosome: side effects or a hidden logic? Vítězslav Bryja , Igor Červenka & Lukáš Čajánek To cite this article: Vítězslav Bryja , Igor Červenka & Lukáš Čajánek (2017): The connections of Wnt pathway components with cell cycle and centrosome: side effects or a hidden logic?, Critical Reviews in Biochemistry and Molecular Biology, DOI: 10.1080/10409238.2017.1350135 To link to this article: http://dx.doi.org/10.1080/10409238.2017.1350135 Published online: 25 Jul 2017. Submit your article to this journal Article views: 72 View related articles View Crossmark data Full Terms & Conditions of access and use can be found at http://www.tandfonline.com/action/journalInformation?journalCode=ibmg20 Download by: [Masarykova Univerzita v Brne], [Lukas Cajanek] Date: 08 August 2017, At: 01:58 CRITICAL REVIEWS IN BIOCHEMISTRY AND MOLECULAR BIOLOGY, 2017 https://doi.org/10.1080/10409238.2017.1350135 REVIEW ARTICLE The connections of Wnt pathway components with cell cycle and centrosome: side effects or a hidden logic? Vıtezslav Bryjaa , Igor Cervenka b and Lukas Caj anekc aDepartment of Experimental Biology, Faculty of Science, Masaryk University, Brno, Czech Republic; bMolecular and Cellular Exercise Physiology, Department of Physiology and Pharmacology, Karolinska Institutet, Stockholm, Sweden; cDepartment of Histology and Embryology, Faculty of Medicine, Masaryk University, Brno, Czech Republic ABSTRACT ARTICLE HISTORY Wnt signaling cascade has developed together with multicellularity to orchestrate the develop- Received 10 April 2017 ment and homeostasis of complex structures. -

Next Generation Sequencing Panels for Disorders of Sex Development
Next Generation Sequencing Panels for Disorders of Sex Development Disorders of Sex Development – Overview Disorders of sex development (DSDs) occur when sex development does not follow the course of typical male or female patterning. Types of DSDs include congenital development of ambiguous genitalia, disjunction between the internal and external sex anatomy, incomplete development of the sex anatomy, and abnormalities of the development of gonads (such as ovotestes or streak ovaries) (1). Sex chromosome anomalies including Turner syndrome and Klinefelter syndrome as well as sex chromosome mosaicism are also considered to be DSDs. DSDs can be caused by a wide range of genetic abnormalities (2). Determining the etiology of a patient’s DSD can assist in deciding gender assignment, provide recurrence risk information for future pregnancies, and can identify potential health problems such as adrenal crisis or gonadoblastoma (1, 3). Sex chromosome aneuploidy and copy number variation are common genetic causes of DSDs. For this reason, chromosome analysis and/or microarray analysis typically should be the first genetic analysis in the case of a patient with ambiguous genitalia or other suspected disorder of sex development. Identifying whether a patient has a 46,XY or 46,XX karyotype can also be helpful in determining appropriate additional genetic testing. Abnormal/Ambiguous Genitalia Panel Our Abnormal/Ambiguous Genitalia Panel includes mutation analysis of 72 genes associated with both syndromic and non-syndromic DSDs. This comprehensive panel evaluates a broad range of genetic causes of ambiguous or abnormal genitalia, including conditions in which abnormal genitalia are the primary physical finding as well as syndromic conditions that involve abnormal genitalia in addition to other congenital anomalies. -

Defining Functional Interactions During Biogenesis of Epithelial Junctions
ARTICLE Received 11 Dec 2015 | Accepted 13 Oct 2016 | Published 6 Dec 2016 | Updated 5 Jan 2017 DOI: 10.1038/ncomms13542 OPEN Defining functional interactions during biogenesis of epithelial junctions J.C. Erasmus1,*, S. Bruche1,*,w, L. Pizarro1,2,*, N. Maimari1,3,*, T. Poggioli1,w, C. Tomlinson4,J.Lees5, I. Zalivina1,w, A. Wheeler1,w, A. Alberts6, A. Russo2 & V.M.M. Braga1 In spite of extensive recent progress, a comprehensive understanding of how actin cytoskeleton remodelling supports stable junctions remains to be established. Here we design a platform that integrates actin functions with optimized phenotypic clustering and identify new cytoskeletal proteins, their functional hierarchy and pathways that modulate E-cadherin adhesion. Depletion of EEF1A, an actin bundling protein, increases E-cadherin levels at junctions without a corresponding reinforcement of cell–cell contacts. This unexpected result reflects a more dynamic and mobile junctional actin in EEF1A-depleted cells. A partner for EEF1A in cadherin contact maintenance is the formin DIAPH2, which interacts with EEF1A. In contrast, depletion of either the endocytic regulator TRIP10 or the Rho GTPase activator VAV2 reduces E-cadherin levels at junctions. TRIP10 binds to and requires VAV2 function for its junctional localization. Overall, we present new conceptual insights on junction stabilization, which integrate known and novel pathways with impact for epithelial morphogenesis, homeostasis and diseases. 1 National Heart and Lung Institute, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. 2 Computing Department, Imperial College London, London SW7 2AZ, UK. 3 Bioengineering Department, Faculty of Engineering, Imperial College London, London SW7 2AZ, UK. 4 Department of Surgery & Cancer, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Atherosclerosis-Susceptible and Atherosclerosis-Resistant Pigeon Aortic Cells Express Different Genes in Vivo
University of New Hampshire University of New Hampshire Scholars' Repository New Hampshire Agricultural Experiment Station Publications New Hampshire Agricultural Experiment Station 7-1-2013 Atherosclerosis-susceptible and atherosclerosis-resistant pigeon aortic cells express different genes in vivo Janet L. Anderson University of New Hampshire, [email protected] C. M. Ashwell University of New Hampshire - Main Campus S. C. Smith University of New Hampshire - Main Campus R. Shine University of New Hampshire - Main Campus E. C. Smith University of New Hampshire - Main Campus See next page for additional authors Follow this and additional works at: https://scholars.unh.edu/nhaes Part of the Poultry or Avian Science Commons Recommended Citation J. L. Anderson, C. M. Ashwell, S. C. Smith, R. Shine, E. C. Smith and R. L. Taylor, Jr. Atherosclerosis- susceptible and atherosclerosis-resistant pigeon aortic cells express different genes in vivo Poultry Science (2013) 92 (10): 2668-2680 doi:10.3382/ps.2013-03306 This Article is brought to you for free and open access by the New Hampshire Agricultural Experiment Station at University of New Hampshire Scholars' Repository. It has been accepted for inclusion in New Hampshire Agricultural Experiment Station Publications by an authorized administrator of University of New Hampshire Scholars' Repository. For more information, please contact [email protected]. Authors Janet L. Anderson, C. M. Ashwell, S. C. Smith, R. Shine, E. C. Smith, and Robert L. Taylor Jr. This article is available at University of New Hampshire Scholars' Repository: https://scholars.unh.edu/nhaes/207 Atherosclerosis-susceptible and atherosclerosis-resistant pigeon aortic cells express different genes in vivo J. -

Spectrin Binding Motifs Control Scribble Cortical Dynamics And
1 2 3 Spectrin binding motifs control Scribble cortical 4 dynamics and polarity function 5 6 Batiste Boëda and Sandrine Etienne Manneville 7 8 Institut Pasteur (CNRS URA 3691-INSERM), France 9 Correspondance to Sandrine Etienne-Manneville. Cell Polarity, Migration and Cancer Unit, 10 Institut Pasteur, 25 rue du Dr Roux, 75724 Paris cedex 15, France; Phone: +33 1 4438 9591; 11 FAX: +33 1 4568 8548. 12 e-mail: [email protected] 13 14 Competing interests statement: 15 The authors declare that no competing interests exist. 16 17 1 18 Abstract 19 The tumor suppressor protein Scribble (SCRIB) plays an evolutionary conserved role in 20 cell polarity. Despite being central for its function, the molecular basis of SCRIB 21 recruitment and stabilization at the cell cortex is poorly understood. Here we show that 22 SCRIB binds directly to the CH1 domain of spectrins, a molecular scaffold that 23 contributes to the cortical actin cytoskeleton and connects it to the plasma membrane. 24 We have identified a short evolutionary conserved peptide motif named SADH motif 25 (SCRIB ABLIMs DMTN Homology) which is necessary and sufficient to mediate protein 26 interaction with spectrins. The SADH domains contribute to SCRIB dynamics at the 27 cell cortex and SCRIB polarity function. Furthermore, mutations in SCRIB SADH 28 domains associated with spina bifida and cancer impact the stability of SCRIB at the 29 plasma membrane, suggesting that SADH domain alterations may participate in human 30 pathology. 31 32 33 34 35 36 37 2 38 Introduction 39 The protein SCRIB has been implicated in a staggering array of cellular processes 40 including polarity, migration, proliferation, differentiation, apoptosis, stemcell 41 maintenance, and vesicle trafficking [1]. -

Myopathy Genes (HGNC) Neuropathy (HGNC) Neuromuscular Disease
Myopathy Genes Neuropathy Neuromuscular Disease (HGNC) (HGNC) (HGNC) ABHD5 ABCA1 ADCK3 ACTG2 ACO2 AGRN AGK AGXT ALS2 ALDOA AIFM1 ANG AMER1 ALAD AP4B1 ANO5 AMACR AP4E1 AR AP1S1 AP4M1 AUH APTX AP4S1 B4GALT1 AR AP5Z1 CACNA1S ATL3 ATM CASQ1 B4GALNT1 ATXN10 CCDC78 BAG3 ATXN7 CHCHD10 BRP44L BEAN1 CHRNA1 C12orf65 C9orf72 CHRNB1 C19orf12 CACNB4 CHRND C1NH CAPN3 CHRNE CECR1 CHAT CLPB CISD2 CHKB COL6A1 CLCF1 CHMP2B COL6A2 CLCN2 CHRNG COL6A3 CLP1 CLCN1 COLQ CMT2G COL9A3 CTNS CMT2H COQ2 DGUOK CMTDIA COQ6 DNA2 CMTX2 COQ9 DNAJB6 CMTX3 COX15 DNAJC19 COASY CPT1A DNM2 COX6A1 CYP7B1 DPM2 CPOX DAG1 DYSF CYP27A1 DDHD2 EMD CYP2U1 DOK7 EPG5 DARS2 DPAGT1 FAM111B DCAF8 DPM3 FBXL4 DDHD1 DUX4 FKBP14 DFNX5 ECEL1 FKRP DHTKD1 ERBB3 FLH1 DIAPH3 ERLIN2 FLNC DNAJB2 FA2H HNRNPA1 DNAJC3 FKTN HNRNPDL ELOVL5 FUS HNRPA2B1 ERCC8 G6PC KLHL40 FAH GFPT1 KLHL41 FAM126A GLE1 LAMA2 FBN1 GYS2 LDB3 FMR1 HSPD1 LMOD3 FXN IFRD1 MEGF10 GALC INF2 MGME1 GBE1 ISPD MTAP GJC2 ITGA7 MTMR14 GP1BA ITPR1 MYF6 HADHA KCNA1 MYH14 HADHB KCNC3 MYLK2 HFE KCNE3 NARS2 HINT1 KCNJ18 NEB HK1 KCNJ2 ORAI1 HMBS KIAA0196 PRKAG2 HSD17B4 KIF21A PTEN HSN1B L1CAM RBCK1 IARS2 LAMB2 RET IGHMBP2 LARGE RMND1 KCNJ10 MCCC2 SCN4A KIF5A MRE11A SERAC1 LRSAM1 MRPL3 SGCA LYST MTO1 SIL1 MANBA MTPAP SPEG MARS MTTP STAC3 MTATP6 MUSK STIM1 MYH14 MYBPC3 SYNE1 MYOT MYH3 SYNE2 NAMSD MYH8 TAZ NF2 NF1 TIA1 NGLY1 NIPA1 TMEM43 NMSR NOP56 TNPO3 NOTCH3 OPTN TNXB OPA1 PDSS2 TPM2 OPA3 PDYN TRPV4 OTOF PFN1 UBA1 PDK3 PHKA2 VCP PDSS1 PHKG2 XDH PEX10 PHOX2A ACADS PEX2 PIP5K1C ACADVL PMM2 PLEC ACTA1 PNPLA6 PLP1 AGL PPOX POMGNT1 AMPD1 PRICKLE1 -

Cldn19 Clic2 Clmp Cln3
NewbornDx™ Advanced Sequencing Evaluation When time to diagnosis matters, the NewbornDx™ Advanced Sequencing Evaluation from Athena Diagnostics delivers rapid, 5- to 7-day results on a targeted 1,722-genes. A2ML1 ALAD ATM CAV1 CLDN19 CTNS DOCK7 ETFB FOXC2 GLUL HOXC13 JAK3 AAAS ALAS2 ATP1A2 CBL CLIC2 CTRC DOCK8 ETFDH FOXE1 GLYCTK HOXD13 JUP AARS2 ALDH18A1 ATP1A3 CBS CLMP CTSA DOK7 ETHE1 FOXE3 GM2A HPD KANK1 AASS ALDH1A2 ATP2B3 CC2D2A CLN3 CTSD DOLK EVC FOXF1 GMPPA HPGD K ANSL1 ABAT ALDH3A2 ATP5A1 CCDC103 CLN5 CTSK DPAGT1 EVC2 FOXG1 GMPPB HPRT1 KAT6B ABCA12 ALDH4A1 ATP5E CCDC114 CLN6 CUBN DPM1 EXOC4 FOXH1 GNA11 HPSE2 KCNA2 ABCA3 ALDH5A1 ATP6AP2 CCDC151 CLN8 CUL4B DPM2 EXOSC3 FOXI1 GNAI3 HRAS KCNB1 ABCA4 ALDH7A1 ATP6V0A2 CCDC22 CLP1 CUL7 DPM3 EXPH5 FOXL2 GNAO1 HSD17B10 KCND2 ABCB11 ALDOA ATP6V1B1 CCDC39 CLPB CXCR4 DPP6 EYA1 FOXP1 GNAS HSD17B4 KCNE1 ABCB4 ALDOB ATP7A CCDC40 CLPP CYB5R3 DPYD EZH2 FOXP2 GNE HSD3B2 KCNE2 ABCB6 ALG1 ATP8A2 CCDC65 CNNM2 CYC1 DPYS F10 FOXP3 GNMT HSD3B7 KCNH2 ABCB7 ALG11 ATP8B1 CCDC78 CNTN1 CYP11B1 DRC1 F11 FOXRED1 GNPAT HSPD1 KCNH5 ABCC2 ALG12 ATPAF2 CCDC8 CNTNAP1 CYP11B2 DSC2 F13A1 FRAS1 GNPTAB HSPG2 KCNJ10 ABCC8 ALG13 ATR CCDC88C CNTNAP2 CYP17A1 DSG1 F13B FREM1 GNPTG HUWE1 KCNJ11 ABCC9 ALG14 ATRX CCND2 COA5 CYP1B1 DSP F2 FREM2 GNS HYDIN KCNJ13 ABCD3 ALG2 AUH CCNO COG1 CYP24A1 DST F5 FRMD7 GORAB HYLS1 KCNJ2 ABCD4 ALG3 B3GALNT2 CCS COG4 CYP26C1 DSTYK F7 FTCD GP1BA IBA57 KCNJ5 ABHD5 ALG6 B3GAT3 CCT5 COG5 CYP27A1 DTNA F8 FTO GP1BB ICK KCNJ8 ACAD8 ALG8 B3GLCT CD151 COG6 CYP27B1 DUOX2 F9 FUCA1 GP6 ICOS KCNK3 ACAD9 ALG9 -

High Diagnostic Yield in Skeletal Ciliopathies Using Massively Parallel Genome Sequencing, Structural Variant Screening and RNA Analyses
Journal of Human Genetics (2021) 66:995–1008 https://doi.org/10.1038/s10038-021-00925-x ARTICLE High diagnostic yield in skeletal ciliopathies using massively parallel genome sequencing, structural variant screening and RNA analyses 1 1 2,3 4 1 Anna Hammarsjö ● Maria Pettersson ● David Chitayat ● Atsuhiko Handa ● Britt-Marie Anderlid ● 5 6 7 8 9 Marco Bartocci ● Donald Basel ● Dominyka Batkovskyte ● Ana Beleza-Meireles ● Peter Conner ● 10 11 12,13 7,14 15 Jesper Eisfeldt ● Katta M. Girisha ● Brian Hon-Yin Chung ● Eva Horemuzova ● Hironobu Hyodo ● 16 1 17 18,19 20 Liene Korņejeva ● Kristina Lagerstedt-Robinson ● Angela E. Lin ● Måns Magnusson ● Shahida Moosa ● 11 10 21 15 18,22 Shalini S. Nayak ● Daniel Nilsson ● Hirofumi Ohashi ● Naoko Ohashi-Fukuda ● Henrik Stranneheim ● 1 23 24 19,22 1 7,25 Fulya Taylan ● Rasa Traberg ● Ulrika Voss ● Valtteri Wirta ● Ann Nordgren ● Gen Nishimura ● 1 1 Anna Lindstrand ● Giedre Grigelioniene Received: 4 December 2020 / Revised: 31 March 2021 / Accepted: 31 March 2021 / Published online: 20 April 2021 © The Author(s) 2021. This article is published with open access Abstract Skeletal ciliopathies are a heterogenous group of disorders with overlapping clinical and radiographic features including 1234567890();,: 1234567890();,: bone dysplasia and internal abnormalities. To date, pathogenic variants in at least 30 genes, coding for different structural cilia proteins, are reported to cause skeletal ciliopathies. Here, we summarize genetic and phenotypic features of 34 affected individuals from 29 families with skeletal ciliopathies. Molecular diagnostic testing was performed using massively parallel sequencing (MPS) in combination with copy number variant (CNV) analyses and in silico filtering for variants in known skeletal ciliopathy genes. -

Supplementary Material Contents
Supplementary Material Contents Immune modulating proteins identified from exosomal samples.....................................................................2 Figure S1: Overlap between exosomal and soluble proteomes.................................................................................... 4 Bacterial strains:..............................................................................................................................................4 Figure S2: Variability between subjects of effects of exosomes on BL21-lux growth.................................................... 5 Figure S3: Early effects of exosomes on growth of BL21 E. coli .................................................................................... 5 Figure S4: Exosomal Lysis............................................................................................................................................ 6 Figure S5: Effect of pH on exosomal action.................................................................................................................. 7 Figure S6: Effect of exosomes on growth of UPEC (pH = 6.5) suspended in exosome-depleted urine supernatant ....... 8 Effective exosomal concentration....................................................................................................................8 Figure S7: Sample constitution for luminometry experiments..................................................................................... 8 Figure S8: Determining effective concentration ......................................................................................................... -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -
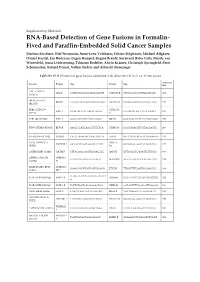
RNA-Based Detection of Gene Fusions in Formalin- Fixed And
Supplementary Materials RNA-Based Detection of Gene Fusions in Formalin- Fixed and Paraffin-Embedded Solid Cancer Samples Martina Kirchner, Olaf Neumann, Anna-Lena Volckmar, Fabian Stögbauer, Michael Allgäuer, Daniel Kazdal, Jan Budczies, Eugen Rempel, Regine Brandt, Suranand Babu Talla, Moritz von Winterfeld, Jonas Leichsenring, Tilmann Bochtler, Alwin Krämer, Christoph Springfeld, Peter Schirmacher, Roland Penzel, Volker Endris and Albrecht Stenzinger Table S1. PCR Primers for gene fusions identified with either the OCAv3- or Archer-panel. Amplicon Fusion Primer Seq Primer Seq [bp] AXL::CAPN15 AXL F CATGGATGAGGGTGGAGGTT CAPN15 R CTGGGCACACGTGAATCAC 178 (A19C2) BRD3::NUTM1 BRD3 F AAGAAACAGGCAGCCAAGTC NUTM1 R CTGGTGGGTCAGAAGTTGGT 217 (B11N2) ESR1-CCDC170 CCDC170 ESR1 F GGAGACTCGCTACTGTGCA CCCAGACTCCTTTCCCAACT 167 (E2C7) R ESR1-QKI (E2Q5) ESR1 F GGAGACTCGCTACTGTGCA QKI R GGCTGGTGATTTAATGTTGGC 197 ETV6::NTRK3 (E5N15) ETV6 F AAGCCCATCAACCTCTCTCA NTRK3 R GGGCTGAGGTTGTAGCACTC 206 FGFR2::INA (F17I2) FGFR2 F CTCCCAGAGACCAACGTTCA INA R GTCCTGGTATTCCCGAAAGGT 148 FNDC3B-PIK3CA PIK3CA FNDC3B F GCAGCTCAGCAGGTTATTCT GTCGTGGAGGCATTGTTCTG 177 (F3P2) R1 GATM::RAF1 (G2R8) GATM F CTTACAACGAATGGGACCCC RAF1 R GTTGGGCTCAGATTGTTGGG 160 GPBP1L1::MAST2 GPBP1L1 CGTAGTGGAGGTGGCACA MAST2 R1 AGGTGATGTGCTAGAGGTCA 178 (G6M4) F1 HNRNPA2B1::ETV1 HNRNPA GGAGGATATGGTGGTGGAGG ETV1 R TTGATTTTCAGTGGCAGGCC 164 (H9E6) 2B1 F TGATGAATCTGGAATTGTTGCT MYB::NFIB (M12N9) MYB12 F NFIB9 R CGTAATTTTGGACATTGGCCG 150 G MYB::NFIB (M13N9) MYB13 F TCTTCTGCTCACACCACTGG NFIB9 R CGTAATTTTGGACATTGGCCG 160 SND1::BRAF (S9B9) SND1 F CGATTCACCTGTCCAGCATC BRAF R CGCTGAGGTCCTGGAGATTT 184 TBL1XR1::PIK3CA TBL1XR1 F TTTCCTTGTGCCTCCATTCC PIK3CA R GTCGTGGAGGCATTGTTCTG 195 (T1P2) TMPRSS2 TMPRSS2::ERG (T2E4) CGCGGCAGGTCATATTGAA ERG R CCTTCCCATCGATGTTCTGG 190 F WHSC1L1::FGFR1 WHSC1L1 TGATCGCACTGACACGGC FGFR1 R ACAAGGCTCCACATCTCCAT 108 (W1F2) F Table S2. Clinical and diagnostic implications of the detected gene fusions. Reclassification Entities Where Entity Where Fusion Is Based on Molecular Fusion Is Fusion Drug Ref.