Dlgap1 Knockout Mice Exhibit Alterations of the Postsynaptic Density and Selective Reductions in Sociability
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Association Analyses of Known Genetic Variants with Gene
ASSOCIATION ANALYSES OF KNOWN GENETIC VARIANTS WITH GENE EXPRESSION IN BRAIN by Viktoriya Strumba A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Bioinformatics) in The University of Michigan 2009 Doctoral Committee: Professor Margit Burmeister, Chair Professor Huda Akil Professor Brian D. Athey Assistant Professor Zhaohui S. Qin Research Statistician Thomas Blackwell To Sam and Valentina Dmitriy and Elizabeth ii ACKNOWLEDGEMENTS I would like to thank my advisor Professor Margit Burmeister, who tirelessly guided me though seemingly impassable corridors of graduate work. Throughout my thesis writing period she provided sound advice, encouragement and inspiration. Leading by example, her enthusiasm and dedication have been instrumental in my path to becoming a better scientist. I also would like to thank my co-advisor Tom Blackwell. His careful prodding always kept me on my toes and looking for answers, which taught me the depth of careful statistical analysis. His diligence and dedication have been irreplaceable in most difficult of projects. I also would like to thank my other committee members: Huda Akil, Brian Athey and Steve Qin as well as David States. You did not make it easy for me, but I thank you for believing and not giving up. Huda’s eloquence in every subject matter she explained have been particularly inspiring, while both Huda’s and Brian’s valuable advice made the completion of this dissertation possible. I would also like to thank all the members of the Burmeister lab, both past and present: Sandra Villafuerte, Kristine Ito, Cindy Schoen, Karen Majczenko, Ellen Schmidt, Randi Burns, Gang Su, Nan Xiang and Ana Progovac. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Rare Variants and Loci for Age-Related Macular Degeneration in The
Human Genetics (2019) 138:1171–1182 https://doi.org/10.1007/s00439-019-02050-4 ORIGINAL INVESTIGATION Rare variants and loci for age‑related macular degeneration in the Ohio and Indiana Amish Andrea R. Waksmunski1,2,3 · Robert P. Igo Jr.3 · Yeunjoo E. Song3 · Jessica N. Cooke Bailey2,3 · Renee Laux3 · Denise Fuzzell3 · Sarada Fuzzell3 · Larry D. Adams4 · Laura Caywood4 · Michael Prough4 · Dwight Stambolian5 · William K. Scott4 · Margaret A. Pericak‑Vance4 · Jonathan L. Haines1,2,3 Received: 27 April 2019 / Accepted: 21 July 2019 / Published online: 31 July 2019 © The Author(s) 2019 Abstract Age-related macular degeneration (AMD) is a leading cause of blindness in the world. While dozens of independent genomic variants are associated with AMD, about one-third of AMD heritability is still unexplained. To identify novel variants and loci for AMD, we analyzed Illumina HumanExome chip data from 87 Amish individuals with early or late AMD, 79 unafected Amish individuals, and 15 related Amish individuals with unknown AMD afection status. We retained 37,428 polymorphic autosomal variants across 175 samples for association and linkage analyses. After correcting for multiple testing (n = 37,428), we identifed four variants signifcantly associated with AMD: rs200437673 (LCN9, p = 1.50 × 10−11), rs151214675 (RTEL1, p = 3.18 × 10−8), rs140250387 (DLGAP1, p = 4.49 × 10−7), and rs115333865 (CGRRF1, p = 1.05 × 10−6). These variants have not been previously associated with AMD and are not in linkage disequilibrium with the 52 known AMD-associated variants reported by the International AMD Genomics Consortium based on physical distance. Genome-wide signifcant linkage peaks were observed on chromosomes 8q21.11–q21.13 (maximum recessive HLOD = 4.03) and 18q21.2–21.32 (maximum dominant HLOD = 3.87; maximum recessive HLOD = 4.27). -

8P23.2-Pter Microdeletions: Seven New Cases Narrowing the Candidate Region and Review of the Literature
G C A T T A C G G C A T genes Article 8p23.2-pter Microdeletions: Seven New Cases Narrowing the Candidate Region and Review of the Literature Ilaria Catusi 1,* , Maria Garzo 1 , Anna Paola Capra 2 , Silvana Briuglia 2 , Chiara Baldo 3 , Maria Paola Canevini 4 , Rachele Cantone 5, Flaviana Elia 6, Francesca Forzano 7, Ornella Galesi 8, Enrico Grosso 5, Michela Malacarne 3, Angela Peron 4,9,10 , Corrado Romano 11 , Monica Saccani 4 , Lidia Larizza 1 and Maria Paola Recalcati 1 1 Istituto Auxologico Italiano, IRCCS, Laboratory of Medical Cytogenetics and Molecular Genetics, 20145 Milan, Italy; [email protected] (M.G.); [email protected] (L.L.); [email protected] (M.P.R.) 2 Department of Biomedical, Dental, Morphological and Functional Imaging Sciences, University of Messina, 98100 Messina, Italy; [email protected] (A.P.C.); [email protected] (S.B.) 3 UOC Laboratorio di Genetica Umana, IRCCS Istituto Giannina Gaslini, 16147 Genova, Italy; [email protected] (C.B.); [email protected] (M.M.) 4 Child Neuropsychiatry Unit—Epilepsy Center, Department of Health Sciences, ASST Santi Paolo e Carlo, San Paolo Hospital, Università Degli Studi di Milano, 20142 Milan, Italy; [email protected] (M.P.C.); [email protected] (A.P.); [email protected] (M.S.) 5 Medical Genetics Unit, Città della Salute e della Scienza University Hospital, 10126 Turin, Italy; [email protected] (R.C.); [email protected] (E.G.) 6 Unit of Psychology, Oasi Research Institute-IRCCS, -

ADHD) Gene Networks in Children of Both African American and European American Ancestry
G C A T T A C G G C A T genes Article Rare Recurrent Variants in Noncoding Regions Impact Attention-Deficit Hyperactivity Disorder (ADHD) Gene Networks in Children of both African American and European American Ancestry Yichuan Liu 1 , Xiao Chang 1, Hui-Qi Qu 1 , Lifeng Tian 1 , Joseph Glessner 1, Jingchun Qu 1, Dong Li 1, Haijun Qiu 1, Patrick Sleiman 1,2 and Hakon Hakonarson 1,2,3,* 1 Center for Applied Genomics, Children’s Hospital of Philadelphia, Philadelphia, PA 19104, USA; [email protected] (Y.L.); [email protected] (X.C.); [email protected] (H.-Q.Q.); [email protected] (L.T.); [email protected] (J.G.); [email protected] (J.Q.); [email protected] (D.L.); [email protected] (H.Q.); [email protected] (P.S.) 2 Division of Human Genetics, Department of Pediatrics, The Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA 19104, USA 3 Department of Human Genetics, Children’s Hospital of Philadelphia, Philadelphia, PA 19104, USA * Correspondence: [email protected]; Tel.: +1-267-426-0088 Abstract: Attention-deficit hyperactivity disorder (ADHD) is a neurodevelopmental disorder with poorly understood molecular mechanisms that results in significant impairment in children. In this study, we sought to assess the role of rare recurrent variants in non-European populations and outside of coding regions. We generated whole genome sequence (WGS) data on 875 individuals, Citation: Liu, Y.; Chang, X.; Qu, including 205 ADHD cases and 670 non-ADHD controls. The cases included 116 African Americans H.-Q.; Tian, L.; Glessner, J.; Qu, J.; Li, (AA) and 89 European Americans (EA), and the controls included 408 AA and 262 EA. -

Mirna Regulons Associated with Synaptic Function
miRNA Regulons Associated with Synaptic Function Maria Paschou1, Maria D. Paraskevopoulou2, Ioannis S. Vlachos2, Pelagia Koukouraki1, Artemis G. Hatzigeorgiou2,3, Epaminondas Doxakis1* 1 Basic Neurosciences Division, Biomedical Research Foundation of the Academy of Athens, Athens, Greece, 2 Institute of Molecular Oncology, Biomedical Sciences Research Center ‘‘Alexander Fleming’’ Vari, Greece, 3 Department of Computer and Communication Engineering, University of Thessaly, Volos, Greece Abstract Differential RNA localization and local protein synthesis regulate synapse function and plasticity in neurons. MicroRNAs are a conserved class of regulatory RNAs that control mRNA stability and translation in tissues. They are abundant in the brain but the extent into which they are involved in synaptic mRNA regulation is poorly known. Herein, a computational analysis of the coding and 39UTR regions of 242 presynaptic and 304 postsynaptic proteins revealed that 91% of them are predicted to be microRNA targets. Analysis of the longest 39UTR isoform of synaptic transcripts showed that presynaptic mRNAs have significantly longer 39UTR than control and postsynaptic mRNAs. In contrast, the shortest 39UTR isoform of postsynaptic mRNAs is significantly shorter than control and presynaptic mRNAs, indicating they avert microRNA regulation under specific conditions. Examination of microRNA binding site density of synaptic 39UTRs revealed that they are twice as dense as the rest of protein-coding transcripts and that approximately 50% of synaptic transcripts are predicted to have more than five different microRNA sites. An interaction map exploring the association of microRNAs and their targets revealed that a small set of ten microRNAs is predicted to regulate 77% and 80% of presynaptic and postsynaptic transcripts, respectively. -

Dlgap1 Knockout Mice Exhibit Alterations of the Postsynaptic
www.nature.com/scientificreports OPEN Dlgap1 knockout mice exhibit alterations of the postsynaptic density and selective reductions in Received: 11 October 2017 Accepted: 16 January 2018 sociability Published: xx xx xxxx M. P. Coba 3, M. J. Ramaker1, E. V. Ho1, S. L. Thompson1,2, N. H. Komiyama4, S. G. N. Grant 4, J. A. Knowles3 & S. C. Dulawa1 The scafold protein DLGAP1 is localized at the post-synaptic density (PSD) of glutamatergic neurons and is a component of supramolecular protein complexes organized by PSD95. Gain-of-function variants of DLGAP1 have been associated with obsessive-compulsive disorder (OCD), while haploinsufcient variants have been linked to autism spectrum disorder (ASD) and schizophrenia in human genetic studies. We tested male and female Dlgap1 wild type (WT), heterozygous (HT), and knockout (KO) mice in a battery of behavioral tests: open feld, dig, splash, prepulse inhibition, forced swim, nest building, social approach, and sucrose preference. We also used biochemical approaches to examine the role of DLGAP1 in the organization of PSD protein complexes. Dlgap1 KO mice were most notable for disruption of protein interactions in the PSD, and defcits in sociability. Other behavioral measures were largely unafected. Our data suggest that Dlgap1 knockout leads to PSD disruption and reduced sociability, consistent with reports of DLGAP1 haploinsufcient variants in schizophrenia and ASD. Te Disks Large Associated Protein 1 gene encodes the protein DLGAP1 (also known as GKAP or SAPAP1), which localizes at the postsynaptic density (PSD)1–7. Genetic variants of DLGAP1 and abnormalities of the PSD have been associated with neuropsychiatric disorders including schizophrenia8, autism spectrum disorder (ASD)9, and obsessive-compulsive disorder (OCD)10. -

The DLGAP Family: Neuronal Expression, Function and Role in Brain Disorders Andreas H
Rasmussen et al. Molecular Brain (2017) 10:43 DOI 10.1186/s13041-017-0324-9 REVIEW Open Access The DLGAP family: neuronal expression, function and role in brain disorders Andreas H. Rasmussen1, Hanne B. Rasmussen2 and Asli Silahtaroglu1* Abstract The neurotransmitter glutamate facilitates neuronal signalling at excitatory synapses. Glutamate is released from the presynaptic membrane into the synaptic cleft. Across the synaptic cleft glutamate binds to both ion channels and metabotropic glutamate receptors at the postsynapse, which expedite downstream signalling in the neuron. The postsynaptic density, a highly specialized matrix, which is attached to the postsynaptic membrane, controls this downstream signalling. The postsynaptic density also resets the synapse after each synaptic firing. It is composed of numerous proteins including a family of Discs large associated protein 1, 2, 3 and 4 (DLGAP1-4) that act as scaffold proteins in the postsynaptic density. They link the glutamate receptors in the postsynaptic membrane to other glutamate receptors, to signalling proteins and to components of the cytoskeleton. With the central localisation in the postsynapse, the DLGAP family seems to play a vital role in synaptic scaling by regulating the turnover of both ionotropic and metabotropic glutamate receptors in response to synaptic activity. DLGAP family has been directly linked to a variety of psychological and neurological disorders. In this review we focus on the direct and indirect role of DLGAP family on schizophrenia as well as other brain diseases. Keywords: DLGAP1, DLGAP2, DLGAP3, DLGAP4, SAPAP, PSD, GKAP, Schizophrenia, Scaffold proteins, Synaptic scaling Introduction interaction partners, DLGAP1–4 proteins are likely to play The postsynaptic density (PSD) is a highly specialized a role in multiple processes of the PSD. -

Genotyping of Families with Obsessive-Compulsive Disorder
University of Zagreb Faculty of Science Department of Biology Martina Ĉonda Genotyping of families with obsessive-compulsive disorder Graduation thesis Zagreb, 2014 This Graduation thesis, produced at the Neurobiochemistry laboratory of the University Clinics of Child and Adolescent Psychiatry, University of Zürich, under mentoring of Dr. Edna Grünblatt, Assoc. Prof. Department of Child and Adolescent Psychiatry at University of Zürich, Zürich (Switzerland), is subjected for evaluation to Department of Biology, Faculty of Science, University of Zagreb for acquisition of the title master of molecular biology. ACKNOWLEDGEMENTS To PD Dr. Edna Grünblatt from University of Zürich, for warm welcoming to her research group and including me to their research project for the purpose of this Master thesis. To Dr. Jasmin Bartl, Dr. Zoya Marinova and Miryame Hofmann for their selfless guidance during my work for this Master thesis, numerous beneficial advices and patience as well as delightful company. To dr. sc. Domagoj Đikić, for his kindness and cooperativeness as well as useful advices during writing of this Master thesis. To my family and friends for their endless help and support during my studies. BASIC DOCUMENTATION CARD University of Zagreb Faculty of Science Department of Biology Graduation thesis GENOTYPING OF FAMILIES WITH OBSESSIVE-COMPULSIVE DISORDER Martina Ĉonda Roosveltov trg 6, 10000 Zagreb, Croatia Obsessive-compulsive disorder (OCD), a neuropsychiatric disorder, is characterized by the presence of obsessions and/or compulsions. -

A Genome-Wide Association Study Identi Es a Novel Susceptibility
A Genome-Wide Association Study Identies a Novel Susceptibility Locus at The DLGAP1 Gene For Resistant Hypertension in The Japanese Population Yasuo Takahashi ( [email protected] ) Nihon University School of Medicine Keiko Yamazaki RIKEN Center for Integrative Medical Sciences Yoichiro Kamatani University of Tokyo Michiaki Kubo RIKEN Center for Integrative Medical Sciences Koichi Matsuda University of Tokyo Satoshi Asai Nihon University School of Medicine Research Article Keywords: resistant hypertension, genome-wide association study, DLGAP1, aldosterone synthase gene, CYP11B2 Posted Date: January 18th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-143274/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/15 Abstract Numerous genetic variants associated with hypertension and blood pressure are known, but there is a paucity of evidence from genetic studies of resistant hypertension, especially in the Asian populations. To identify novel genetic loci associated with resistant hypertension in the Japanese population, we conducted a genome-wide association study with 2,705 resistant hypertension cases and 21,296 mild hypertension controls, all from BioBank Japan. We identied one novel susceptibility locus, rs1442386 on chromosome 18p11.3 (DLGAP1), achieving genome-wide signicance (odds ratio (95% CI) = 0.85 (0.81–0.90), P = 3.75 × 10−8) and 17 loci showing suggestive association, including rs62525059 of 8q24.3 (CYP11B2). We further detected biological processes associated with resistant hypertension, including chemical synaptic transmission, regulation of transmembrane transport, neuron development and neurological system processes, highlighting the importance of the nervous system. This study provides insights into the etiology of resistant hypertension in the Japanese population. -
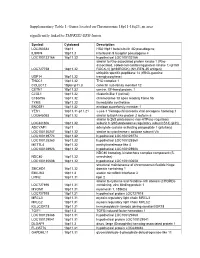
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

Rare Disruptive Variants in the DISC1 Interactome and Regulome: Association with Cognitive Ability and Schizophrenia
OPEN Molecular Psychiatry (2018) 23, 1270–1277 www.nature.com/mp ORIGINAL ARTICLE Rare disruptive variants in the DISC1 Interactome and Regulome: association with cognitive ability and schizophrenia S Teng1,2,10, PA Thomson3,4,10, S McCarthy1,10, M Kramer1, S Muller1, J Lihm1, S Morris3, DC Soares3, W Hennah5, S Harris3,4, LM Camargo6, V Malkov7, AM McIntosh8, JK Millar3, DH Blackwood8, KL Evans4, IJ Deary4,9, DJ Porteous3,4 and WR McCombie1 Schizophrenia (SCZ), bipolar disorder (BD) and recurrent major depressive disorder (rMDD) are common psychiatric illnesses. All have been associated with lower cognitive ability, and show evidence of genetic overlap and substantial evidence of pleiotropy with cognitive function and neuroticism. Disrupted in schizophrenia 1 (DISC1) protein directly interacts with a large set of proteins (DISC1 Interactome) that are involved in brain development and signaling. Modulation of DISC1 expression alters the expression of a circumscribed set of genes (DISC1 Regulome) that are also implicated in brain biology and disorder. Here we report targeted sequencing of 59 DISC1 Interactome genes and 154 Regulome genes in 654 psychiatric patients and 889 cognitively-phenotyped control subjects, on whom we previously reported evidence for trait association from complete sequencing of the DISC1 locus. Burden analyses of rare and singleton variants predicted to be damaging were performed for psychiatric disorders, cognitive variables and personality traits. The DISC1 Interactome and Regulome showed differential association across the phenotypes tested. After family-wise error correction across all traits (FWERacross), an increased burden of singleton disruptive variants in the Regulome was associated with SCZ (FWERacross P=0.0339).