Structure Prediction of G-Protein Coupled Receptors
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Expression of G Protein-Coupled Receptor 19 in Human Lung Cancer Cells Is Triggered by Entry Into S-Phase and Supports
Published OnlineFirst August 21, 2012; DOI: 10.1158/1541-7786.MCR-12-0139 Molecular Cancer Cell Cycle, Cell Death, and Senescence Research Expression of G Protein-Coupled Receptor 19 in Human Lung Cancer Cells Is Triggered by Entry into S-Phase and Supports G2–M Cell-Cycle Progression Stefan Kastner1, Tilman Voss1, Simon Keuerleber2, Christina Glockel€ 2, Michael Freissmuth2, and Wolfgang Sommergruber1 Abstract It has long been known that G protein-coupled receptors (GPCR) are subject to illegitimate expression in tumor cells. Presumably, hijacking the normal physiologic functions of GPCRs contributes to all biologic capabilities acquired during tumorigenesis. Here, we searched for GPCRs that were expressed in lung cancer: the mRNA encoding orphan G protein-coupled receptor 19 (GPR19) was found frequently overexpressed in tissue samples obtained from patients with small cell lung cancer. Several observations indicate that over- expression of Gpr19 confers a specific advantage to lung cancer cells by accelerating transition through the cell- cycle. (i) Knockdown of Gpr19 mRNA by RNA interference reduced cell growth of human lung cancer cell – lines. (ii) Cell-cycle progression through G2 M-phase was impaired in cells transfected with siRNAs directed against Gpr19 and this was associated with increased protein levels of cyclin B1 and phosphorylated histone H3. (iii) The expression levels of Gpr19 mRNA varied along the cell-cycle with a peak observed in S-phase. (iv)TheputativecontrolofGpr19 expression by E2F transcription factors was verified by chromatin immuno- precipitation: antibodies directed against E2F-1 to -4 allowed for the recovery of the Gpr19 promoter. (v) Removal of E2F binding sites in the Gpr19 promoter diminished the expression of a luciferase reporter. -

Genome-Wide Prediction of Small Molecule Binding to Remote
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.04.236729; this version posted August 5, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Genome-wide Prediction of Small Molecule Binding 2 to Remote Orphan Proteins Using Distilled Sequence 3 Alignment Embedding 1 2 3 4 4 Tian Cai , Hansaim Lim , Kyra Alyssa Abbu , Yue Qiu , 5,6 1,2,3,4,7,* 5 Ruth Nussinov , and Lei Xie 1 6 Ph.D. Program in Computer Science, The Graduate Center, The City University of New York, New York, 10016, USA 2 7 Ph.D. Program in Biochemistry, The Graduate Center, The City University of New York, New York, 10016, USA 3 8 Department of Computer Science, Hunter College, The City University of New York, New York, 10065, USA 4 9 Ph.D. Program in Biology, The Graduate Center, The City University of New York, New York, 10016, USA 5 10 Computational Structural Biology Section, Basic Science Program, Frederick National Laboratory for Cancer Research, 11 Frederick, MD 21702, USA 6 12 Department of Human Molecular Genetics and Biochemistry, Sackler School of Medicine, Tel Aviv University, Tel 13 Aviv, Israel 7 14 Helen and Robert Appel Alzheimer’s Disease Research Institute, Feil Family Brain & Mind Research Institute, Weill 15 Cornell Medicine, Cornell University, New York, 10021, USA * 16 [email protected] 17 July 27, 2020 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.08.04.236729; this version posted August 5, 2020. -

Edinburgh Research Explorer
Edinburgh Research Explorer International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list Citation for published version: Davenport, AP, Alexander, SPH, Sharman, JL, Pawson, AJ, Benson, HE, Monaghan, AE, Liew, WC, Mpamhanga, CP, Bonner, TI, Neubig, RR, Pin, JP, Spedding, M & Harmar, AJ 2013, 'International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands', Pharmacological reviews, vol. 65, no. 3, pp. 967-86. https://doi.org/10.1124/pr.112.007179 Digital Object Identifier (DOI): 10.1124/pr.112.007179 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Pharmacological reviews Publisher Rights Statement: U.S. Government work not protected by U.S. copyright General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 1521-0081/65/3/967–986$25.00 http://dx.doi.org/10.1124/pr.112.007179 PHARMACOLOGICAL REVIEWS Pharmacol Rev 65:967–986, July 2013 U.S. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Targeting Lysophosphatidic Acid in Cancer: the Issues in Moving from Bench to Bedside
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by IUPUIScholarWorks cancers Review Targeting Lysophosphatidic Acid in Cancer: The Issues in Moving from Bench to Bedside Yan Xu Department of Obstetrics and Gynecology, Indiana University School of Medicine, 950 W. Walnut Street R2-E380, Indianapolis, IN 46202, USA; [email protected]; Tel.: +1-317-274-3972 Received: 28 August 2019; Accepted: 8 October 2019; Published: 10 October 2019 Abstract: Since the clear demonstration of lysophosphatidic acid (LPA)’s pathological roles in cancer in the mid-1990s, more than 1000 papers relating LPA to various types of cancer were published. Through these studies, LPA was established as a target for cancer. Although LPA-related inhibitors entered clinical trials for fibrosis, the concept of targeting LPA is yet to be moved to clinical cancer treatment. The major challenges that we are facing in moving LPA application from bench to bedside include the intrinsic and complicated metabolic, functional, and signaling properties of LPA, as well as technical issues, which are discussed in this review. Potential strategies and perspectives to improve the translational progress are suggested. Despite these challenges, we are optimistic that LPA blockage, particularly in combination with other agents, is on the horizon to be incorporated into clinical applications. Keywords: Autotaxin (ATX); ovarian cancer (OC); cancer stem cell (CSC); electrospray ionization tandem mass spectrometry (ESI-MS/MS); G-protein coupled receptor (GPCR); lipid phosphate phosphatase enzymes (LPPs); lysophosphatidic acid (LPA); phospholipase A2 enzymes (PLA2s); nuclear receptor peroxisome proliferator-activated receptor (PPAR); sphingosine-1 phosphate (S1P) 1. -

Flow Reagents Single Color Antibodies CD Chart
CD CHART CD N° Alternative Name CD N° Alternative Name CD N° Alternative Name Beckman Coulter Clone Beckman Coulter Clone Beckman Coulter Clone T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells CD1a T6, R4, HTA1 Act p n n p n n S l CD99 MIC2 gene product, E2 p p p CD223 LAG-3 (Lymphocyte activation gene 3) Act n Act p n CD1b R1 Act p n n p n n S CD99R restricted CD99 p p CD224 GGT (γ-glutamyl transferase) p p p p p p CD1c R7, M241 Act S n n p n n S l CD100 SEMA4D (semaphorin 4D) p Low p p p n n CD225 Leu13, interferon induced transmembrane protein 1 (IFITM1). p p p p p CD1d R3 Act S n n Low n n S Intest CD101 V7, P126 Act n p n p n n p CD226 DNAM-1, PTA-1 Act n Act Act Act n p n CD1e R2 n n n n S CD102 ICAM-2 (intercellular adhesion molecule-2) p p n p Folli p CD227 MUC1, mucin 1, episialin, PUM, PEM, EMA, DF3, H23 Act p CD2 T11; Tp50; sheep red blood cell (SRBC) receptor; LFA-2 p S n p n n l CD103 HML-1 (human mucosal lymphocytes antigen 1), integrin aE chain S n n n n n n n l CD228 Melanotransferrin (MT), p97 p p CD3 T3, CD3 complex p n n n n n n n n n l CD104 integrin b4 chain; TSP-1180 n n n n n n n p p CD229 Ly9, T-lymphocyte surface antigen p p n p n -

Metabolite Sensing Gpcrs: Promising Therapeutic Targets for Cancer Treatment?
cells Review Metabolite Sensing GPCRs: Promising Therapeutic Targets for Cancer Treatment? Jesús Cosín-Roger 1,*, Dolores Ortiz-Masia 2 , Maria Dolores Barrachina 3 and Sara Calatayud 3 1 Hospital Dr. Peset, Fundación para la Investigación Sanitaria y Biomédica de la Comunitat Valenciana, FISABIO, 46017 Valencia, Spain 2 Departament of Medicine, Faculty of Medicine, University of Valencia, 46010 Valencia, Spain; [email protected] 3 Departament of Pharmacology and CIBER, Faculty of Medicine, University of Valencia, 46010 Valencia, Spain; [email protected] (M.D.B.); [email protected] (S.C.) * Correspondence: [email protected]; Tel.: +34-963851234 Received: 30 September 2020; Accepted: 21 October 2020; Published: 23 October 2020 Abstract: G-protein-coupled receptors constitute the most diverse and largest receptor family in the human genome, with approximately 800 different members identified. Given the well-known metabolic alterations in cancer development, we will focus specifically in the 19 G-protein-coupled receptors (GPCRs), which can be selectively activated by metabolites. These metabolite sensing GPCRs control crucial processes, such as cell proliferation, differentiation, migration, and survival after their activation. In the present review, we will describe the main functions of these metabolite sensing GPCRs and shed light on the benefits of their potential use as possible pharmacological targets for cancer treatment. Keywords: G-protein-coupled receptor; metabolite sensing GPCR; cancer 1. Introduction G-protein-coupled receptors (GPCRs) are characterized by a seven-transmembrane configuration, constitute the largest and most ubiquitous family of plasma membrane receptors, and regulate virtually all known physiological processes in humans [1,2]. This family includes almost one thousand genes that were initially classified on the basis of sequence homology into six classes (A–F), where classes D and E were not found in vertebrates [3]. -

ADGRE1 (EMR1, F4/80) Is a Rapidly-Evolving Gene Expressed In
Edinburgh Research Explorer ADGRE1 (EMR1, F4/80) Is a Rapidly-Evolving Gene Expressed in Mammalian Monocyte-Macrophages Citation for published version: Waddell, L, Lefevre, L, Bush, S, Raper, A, Young, R, Lisowski, Z, McCulloch, M, Muriuki, C, Sauter, K, Clark, E, Irvine, KN, Pridans, C, Hope, J & Hume, D 2018, 'ADGRE1 (EMR1, F4/80) Is a Rapidly-Evolving Gene Expressed in Mammalian Monocyte-Macrophages' Frontiers in Immunology, vol. 9, 2246. DOI: 10.3389/fimmu.2018.02246 Digital Object Identifier (DOI): 10.3389/fimmu.2018.02246 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Frontiers in Immunology General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 05. Apr. 2019 ORIGINAL RESEARCH published: 01 October 2018 doi: 10.3389/fimmu.2018.02246 ADGRE1 (EMR1, F4/80) Is a Rapidly-Evolving Gene Expressed in Mammalian Monocyte-Macrophages Lindsey A. Waddell 1†, Lucas Lefevre 1†, Stephen J. Bush 2†, Anna Raper 1, Rachel Young 1, Zofia M. -

Conformational Dynamics Between Transmembrane Domains And
RESEARCH ARTICLE Conformational dynamics between transmembrane domains and allosteric modulation of a metabotropic glutamate receptor Vanessa A Gutzeit1, Jordana Thibado2, Daniel Starer Stor2, Zhou Zhou3, Scott C Blanchard2,3,4, Olaf S Andersen2,3, Joshua Levitz2,4,5* 1Neuroscience Graduate Program, Weill Cornell Graduate School of Medical Sciences, New York, United States; 2Physiology, Biophysics and Systems Biology Graduate Program, Weill Cornell Graduate School of Medical Sciences, New York, United States; 3Department of Physiology and Biophysics, Weill Cornell Medicine, New York, United States; 4Tri-Institutional PhD Program in Chemical Biology, New York, United States; 5Department of Biochemistry, Weill Cornell Medicine, New York, United States Abstract Metabotropic glutamate receptors (mGluRs) are class C, synaptic G-protein-coupled receptors (GPCRs) that contain large extracellular ligand binding domains (LBDs) and form constitutive dimers. Despite the existence of a detailed picture of inter-LBD conformational dynamics and structural snapshots of both isolated domains and full-length receptors, it remains unclear how mGluR activation proceeds at the level of the transmembrane domains (TMDs) and how TMD-targeting allosteric drugs exert their effects. Here, we use time-resolved functional and conformational assays to dissect the mechanisms by which allosteric drugs activate and modulate mGluR2. Single-molecule subunit counting and inter-TMD fluorescence resonance energy transfer *For correspondence: measurements in living cells -

G Protein-Coupled Receptors in Stem Cell Maintenance and Somatic Reprogramming to Pluripotent Or Cancer Stem Cells
BMB Reports - Manuscript Submission Manuscript Draft Manuscript Number: BMB-14-250 Title: G protein-coupled receptors in stem cell maintenance and somatic reprogramming to pluripotent or cancer stem cells Article Type: Mini Review Keywords: G protein-coupled receptors; stem cell maintenance; somatic reprogramming; cancer stem cells; pluripotent stem cell Corresponding Author: Ssang-Goo Cho Authors: Ssang-Goo Cho1,*, Hye Yeon Choi1, Subbroto Kumar Saha1, Kyeongseok Kim1, Sangsu Kim1, Gwang-Mo Yang1, BongWoo Kim1, Jin-hoi Kim1 Institution: 1Department of Animal Biotechnology, Animal Resources Research Center, and Incurable Disease Animal Model and Stem Cell Institute (IDASI), Konkuk University, 120 Neungdong-ro, Gwangjin-gu, Seoul 143-701, Republic of Korea, UNCORRECTED PROOF 1 G protein-coupled receptors in stem cell maintenance and somatic reprogramming to 2 pluripotent or cancer stem cells 3 4 Hye Yeon Choi, Subbroto Kumar Saha, Kyeongseok Kim, Sangsu Kim, Gwang-Mo 5 Yang, BongWoo Kim, Jin-hoi Kim, and Ssang-Goo Cho 6 7 Department of Animal Biotechnology, Animal Resources Research Center, and 8 Incurable Disease Animal Model and Stem Cell Institute (IDASI), Konkuk University, 9 120 Neungdong-ro, Gwangjin-gu, Seoul 143-701, Republic of Korea 10 * 11 Address correspondence to Ssang-Goo Cho, Department of Animal Biotechnology and 12 Animal Resources Research Center. Konkuk University, 120 Neungdong-ro, Gwangjin- 13 gu, Seoul 143-701, Republic of Korea. Tel: 82-2-450-4207, Fax: 82-2-450-1044, E-mail: 14 [email protected] 15 16 17 18 19 1 UNCORRECTED PROOF 20 Abstract 21 The G protein-coupled receptors (GPCRs) compose the third largest gene family in the 22 human genome, representing more than 800 distinct genes and 3–5% of the human genome. -

Rabbit Anti-FFAR3/GPR42/FITC Conjugated Antibody-SL16076R
SunLong Biotech Co.,LTD Tel: 0086-571- 56623320 Fax:0086-571- 56623318 E-mail:[email protected] www.sunlongbiotech.com Rabbit Anti-FFAR3/GPR42/FITC Conjugated antibody SL16076R-FITC Product Name: Anti-FFAR3/GPR42/FITC Chinese Name: FITC标记的G protein-coupled receptor42抗体 FFA3R; Ffar3; FFAR3_HUMAN; Free fatty acid receptor 3; G protein coupled Alias: receptor 41; G-protein coupled receptor 41; gpcr41; GPR41; gpr42. Organism Species: Rabbit Clonality: Polyclonal React Species: Human, ICC=1:50-200IF=1:50-200 Applications: not yet tested in other applications. optimal dilutions/concentrations should be determined by the end user. Molecular weight: 39kDa Form: Lyophilized or Liquid Concentration: 1mg/ml immunogen: KLH conjugated synthetic peptide derived from human FFAR3 Lsotype: IgG Purification: affinitywww.sunlongbiotech.com purified by Protein A Storage Buffer: 0.01M TBS(pH7.4) with 1% BSA, 0.03% Proclin300 and 50% Glycerol. Store at -20 °C for one year. Avoid repeated freeze/thaw cycles. The lyophilized antibody is stable at room temperature for at least one month and for greater than a year Storage: when kept at -20°C. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 °C. background: G protein-coupled receptors (GPRs), also known as seven transmembrane receptors, heptahelical receptors or 7TM receptors, comprise a superfamily of proteins that play a Product Detail: role in many different stimulus-response pathways. GPRs translate extracellular signals into intracellular signals (a process called G-protein activation) and they respond to a variety of signaling molecules, such as hormones and neurotransmitters. -
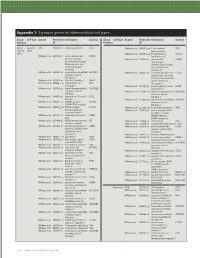
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al.