Large-Scale Discovery of Mouse Transgenic Integration Sites Reveals Frequent Structural Variation and Insertional Mutagenesis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Department of Neuroscience Faculty Research Interests at the Albert Einstein
Dominick P. Purpura Department of Neuroscience Faculty Research Interests at the Albert Einstein DOBRENIS EMMONS ESKANDER COEN-CAGLI College of Medicine FISHMAN CHUA FRANCESCONI CASTILLO 2019–2020 FRENETTE BUSCHKE FRICKER GALANOPOULOU BUELOW GONÇALVES BERGMAN HALL BENNETT HÉBERT BATISTA-BRITO JORDAN BALLABH KHODAKHAH AUTRY ASCHNER AREZZO KURSHAN ALPERT AKABAS LACHMAN LAROCCA LEGATT LIPTON MEHLER MOLHOLM MOSHÉ NICOLA PEÑA PEREDA ROSS RUDOLPH SCHWARTZ SECOMBE SHARP SINGER SJULSON SOLDNER SPRAY STEINSCHNEIDER SUADICANI SUSSMAN VERSELIS WALKLEY WILSON YANG ZHENG ZUKIN Dominick P. Purpura Department of Neuroscience Faculty Research Interests at the Albert Einstein College of Medicine 2019–2020 Myles Akabas, M.D., Ph.D. 1 Peri Kurshan, Ph.D. 68 Joseph C. Arezzo, Ph.D. 6 Herbert M. Lachman, M.D. 70 Michael Aschner, Ph.D. 8 Jorge N. Larocca, Ph.D. 73 Anita E. Autry, Ph.D. 10 Alan D. Legatt, M.D., Ph.D. 75 Praveen Ballabh, M.D. 12 Michael L. Lipton, M.D., Ph.D. 77 Renata Batista-Brito, Ph.D. 14 Mark F. Mehler, M.D. 81 Michael V. L. Bennett, D.Phil. 16 Sophie Molholm, Ph.D. 83 Aviv Bergman, Ph.D. 18 Solomon L. Moshé, M.D. 89 Herman Buschke, M.D. 24 Saleem M. Nicola, Ph.D. 93 Pablo E. Castillo, M.D., Ph.D. 25 José L. Peña, M.D., Ph.D. 96 Streamson C. Chua, Jr., M.D., Ph.D. 27 Alberto E. Pereda, M.D., Ph.D. 98 Ruben Coen-Cagli, Ph.D. 29 Rachel A. Ross, M.D., Ph.D. 100 Kostantin Dobrenis, Ph.D. 31 Stephanie Rudolph, Ph.D. 102 Scott W. -

Analysis of the Indacaterol-Regulated Transcriptome in Human Airway
Supplemental material to this article can be found at: http://jpet.aspetjournals.org/content/suppl/2018/04/13/jpet.118.249292.DC1 1521-0103/366/1/220–236$35.00 https://doi.org/10.1124/jpet.118.249292 THE JOURNAL OF PHARMACOLOGY AND EXPERIMENTAL THERAPEUTICS J Pharmacol Exp Ther 366:220–236, July 2018 Copyright ª 2018 by The American Society for Pharmacology and Experimental Therapeutics Analysis of the Indacaterol-Regulated Transcriptome in Human Airway Epithelial Cells Implicates Gene Expression Changes in the s Adverse and Therapeutic Effects of b2-Adrenoceptor Agonists Dong Yan, Omar Hamed, Taruna Joshi,1 Mahmoud M. Mostafa, Kyla C. Jamieson, Radhika Joshi, Robert Newton, and Mark A. Giembycz Departments of Physiology and Pharmacology (D.Y., O.H., T.J., K.C.J., R.J., M.A.G.) and Cell Biology and Anatomy (M.M.M., R.N.), Snyder Institute for Chronic Diseases, Cumming School of Medicine, University of Calgary, Calgary, Alberta, Canada Received March 22, 2018; accepted April 11, 2018 Downloaded from ABSTRACT The contribution of gene expression changes to the adverse and activity, and positive regulation of neutrophil chemotaxis. The therapeutic effects of b2-adrenoceptor agonists in asthma was general enriched GO term extracellular space was also associ- investigated using human airway epithelial cells as a therapeu- ated with indacaterol-induced genes, and many of those, in- tically relevant target. Operational model-fitting established that cluding CRISPLD2, DMBT1, GAS1, and SOCS3, have putative jpet.aspetjournals.org the long-acting b2-adrenoceptor agonists (LABA) indacaterol, anti-inflammatory, antibacterial, and/or antiviral activity. Numer- salmeterol, formoterol, and picumeterol were full agonists on ous indacaterol-regulated genes were also induced or repressed BEAS-2B cells transfected with a cAMP-response element in BEAS-2B cells and human primary bronchial epithelial cells by reporter but differed in efficacy (indacaterol $ formoterol . -

Csmd2 Is a Synaptic Transmembrane Protein That Interacts with PSD-95 and Is Required for Neuronal Maturation
New Research Development Csmd2 Is a Synaptic Transmembrane Protein that Interacts with PSD-95 and Is Required for Neuronal Maturation Mark A. Gutierrez,1,2 Brett E. Dwyer,1 and Santos J. Franco1,2,3 https://doi.org/10.1523/ENEURO.0434-18.2019 1Department of Pediatrics, University of Colorado School of Medicine, Aurora, CO 80045, 2Cell Biology, Stem Cells and Development Graduate Program, University of Colorado School of Medicine, Aurora, CO 80045, and 3Program of Pediatric Stem Cell Biology, Children’s Hospital Colorado, Aurora, CO 80045 Abstract Mutations and copy number variants of the CUB and Sushi multiple domains 2 (CSMD2) gene are associated with neuropsychiatric disease. CSMD2 encodes a single-pass transmembrane protein with a large extracel- lular domain comprising repeats of CUB and Sushi domains. High expression of CSMD2 in the developing and mature brain suggests possible roles in neuron development or function, but the cellular functions of CSMD2 are not known. In this study, we show that mouse Csmd2 is expressed in excitatory and inhibitory neurons in the forebrain. Csmd2 protein exhibits a somatodendritic localization in the neocortex and hippocampus, with smaller puncta localizing to the neuropil. Using immunohistochemical and biochemical methods, we demonstrate that Csmd2 localizes to dendritic spines and is enriched in the postsynaptic density (PSD). Accordingly, we show that the cytoplasmic tail domain of Csmd2 interacts with synaptic scaffolding proteins of the membrane-associated guanylate kinase (MAGUK) family. The association be- tween Csmd2 and MAGUK member PSD-95 is dependent on a PDZ-binding domain on the Csmd2 tail, which is also required for synaptic targeting of Csmd2. -

Interval of Mouse Chromosome 18
Genetics and Molecular Biology, 28, 2, 201-204 (2005) Copyright by the Brazilian Society of Genetics. Printed in Brazil www.sbg.org.br Research Article The paralysé (par) mouse neurological mutation maps to a 9 Mbp (4 cM) interval of mouse chromosome 18 Lino Silva Neto1, Asadollah Aghaie2, Jean-Louis Guénet2 and Ana Lúcia Brunialti Godard1 1Universidade Federal de Minas Gerais, ICB, Laboratório de Genética Animal e Humana, Belo Horizonte, MG, Brazil. 2Institut Pasteur, Unité de Génétique des Mammifères, Paris, France. Abstract The Paralysé mutation is a spontaneous neuromuscular mutation, first observed in 1980 at the Pasteur Institute, which is transmitted by the autosomal recessive par allele. Affected homozygote par/par mice rarely survive beyond 16 days of age and at the end of their life they are emaciated and completely paralyzed. Several concordant histological and physiological observations indicate that mutant mice might be good models for studying early-onset human motor neuron diseases such as spinal muscular atrophy. Linkage analysis using a set of molecular markers and two F2 crosses indicate that the mutation maps to mouse chromosome 18 in a region spanning 4 cM (or 9 megabase pairs, Mbp) between the microsatellites D18Mit140 and D18Mit33. These results positioned the par locus in a region homologous to human chromosome 18p11.22 to 18q21.32. Key words: mouse model, neuromuscular disease, mouse genetic map, Paralysé mutation. Received: April 26, 2004; Accepted: October 14, 2004. Introduction mutant mice, indicating that the normal process of regres- The spontaneous neuromuscular mutation Paralysé sion of redundant innervation did not occur. Intramuscular was discovered in mice at the Pasteur Institute in the early axons failed to become myelinated in mutant animals but 1980’s. -

18Q22.1-Qter Deletion and 4P16.3 Microduplication in a Boy With
Wang et al. Molecular Cytogenetics (2018) 11:55 https://doi.org/10.1186/s13039-018-0404-2 CASE REPORT Open Access 18q22.1-qter deletion and 4p16.3 microduplication in a boy with speech delay and mental retardation: case report and review of the literature Chunjing Wang1†, Huanhuan Ren1†, Huaifu Dong2, Meng Liang1,QiWu1 and Yaping Liao1* Abstract Background: Deletions involving the long arm of chromosome 18 have been associated with a highly variable phenotypic spectrum that is related to the extent of the deleted region. Duplications in chromosomal region 4p16.3 have also been shown to cause 4p16.3 microduplication syndrome. Most reported patients of trisomy 4p16.3 have more duplications, including the Wolf-Hirschhorn critical region (WHSCR). Here, we present a patient with speech delay and mental retardation caused by a deletion of 18q (18q22.1-qter) and terminal microduplication of 4p (4p16.3-pter) distal to WHSCR. Case presentation: The patient was a 23-month-old boy with moderate growth retardation, severe speech delay, mental retardation, and dysmorphic features. Single nucleotide polymorphism (SNP) array analysis confirmed an 11.2-Mb terminal deletion at 18q22.1 and revealed a 1.8-Mb terminal duplication of 4p16.3. Our patient showed clinical overlap with these two syndromes, although his overall features were milder than what had been previously described. Some dosage-sensitive genes on the 18q terminal deleted region and 4p16.3 duplicated region of the present case may have contributed to his phenotype. Conclusions: This is the first report of a patient with combined terminal deletion of 18q22.1 and duplication of 4p16.3. -

Damaging Coding Variants Within Kainate Receptor Channel Genes Are Enriched in Individuals with Schizophrenia, Autism and Intell
www.nature.com/scientificreports OPEN Damaging coding variants within kainate receptor channel genes are enriched in individuals with schizophrenia, autism and intellectual disabilities Maria Koromina, Miles Flitton, Alix Blockley, Ian R. Mellor & Helen M. Knight* Schizophrenia (Scz), autism spectrum disorder (ASD) and intellectual disability are common complex neurodevelopmental disorders. Kainate receptors (KARs) are ionotropic glutamate ion channels involved in synaptic plasticity which are modulated by auxiliary NETO proteins. Using UK10K exome sequencing data, we interrogated the coding regions of KAR and NETO genes in individuals with Scz, ASD or intellectual disability and population controls; performed follow-up genetic replication studies; and, conducted in silico and in vitro functional studies. We found an excess of Loss-of-Function and missense variants in individuals with Scz compared with control individuals (p = 1.8 × 10−10), and identifed a signifcant burden of functional variants for Scz (p < 1.6 × 10−11) and ASD (p = 6.9 × 10−18). Single allele associations for 6 damaging missense variants were signifcantly replicated (p < 5.0 × 10−15) and confrmed GRIK3 S310A as a protective genetic factor. Functional studies demonstrated that three missense variants located within GluK2 and GluK4, GluK2 (K525E) and GluK4 (Y555N, L825W), afect agonist sensitivity and current decay rates. These fndings establish that genetic variation in KAR receptor ion channels confers risk for schizophrenia, autism and intellectual disability and provide new genetic and pharmacogenetic biomarkers for neurodevelopmental disease. Schizophrenia and autism are common, highly heritable debilitating neurodevelopmental disorders which are ofen comorbid with intellectual disability. Advances in genomic technology suggest a role for both common and rare variants contributing to genetic risk1–4. -

Heterogeneity Between Primary Colon Carcinoma and Paired Lymphatic and Hepatic Metastases
MOLECULAR MEDICINE REPORTS 6: 1057-1068, 2012 Heterogeneity between primary colon carcinoma and paired lymphatic and hepatic metastases HUANRONG LAN1, KETAO JIN2,3, BOJIAN XIE4, NA HAN5, BINBIN CUI2, FEILIN CAO2 and LISONG TENG3 Departments of 1Gynecology and Obstetrics, and 2Surgical Oncology, Taizhou Hospital, Wenzhou Medical College, Linhai, Zhejiang; 3Department of Surgical Oncology, First Affiliated Hospital, College of Medicine, Zhejiang University, Hangzhou, Zhejiang; 4Department of Surgical Oncology, Sir Run Run Shaw Hospital, College of Medicine, Zhejiang University, Hangzhou, Zhejiang; 5Cancer Chemotherapy Center, Zhejiang Cancer Hospital, Zhejiang University of Chinese Medicine, Hangzhou, Zhejiang, P.R. China Received January 26, 2012; Accepted May 8, 2012 DOI: 10.3892/mmr.2012.1051 Abstract. Heterogeneity is one of the recognized characteris- Introduction tics of human tumors, and occurs on multiple levels in a wide range of tumors. A number of studies have focused on the Intratumor heterogeneity is one of the recognized charac- heterogeneity found in primary tumors and related metastases teristics of human tumors, which occurs on multiple levels, with the consideration that the evaluation of metastatic rather including genetic, protein and macroscopic, in a wide range than primary sites could be of clinical relevance. Numerous of tumors, including breast, colorectal cancer (CRC), non- studies have demonstrated particularly high rates of hetero- small cell lung cancer (NSCLC), prostate, ovarian, pancreatic, geneity between primary colorectal tumors and their paired gastric, brain and renal clear cell carcinoma (1). Over the past lymphatic and hepatic metastases. It has also been proposed decade, a number of studies have focused on the heterogeneity that the heterogeneity between primary colon carcinomas and found in primary tumors and related metastases with the their paired lymphatic and hepatic metastases may result in consideration that the evaluation of metastatic rather than different responses to anticancer therapies. -

Robles JTO Supplemental Digital Content 1
Supplementary Materials An Integrated Prognostic Classifier for Stage I Lung Adenocarcinoma based on mRNA, microRNA and DNA Methylation Biomarkers Ana I. Robles1, Eri Arai2, Ewy A. Mathé1, Hirokazu Okayama1, Aaron Schetter1, Derek Brown1, David Petersen3, Elise D. Bowman1, Rintaro Noro1, Judith A. Welsh1, Daniel C. Edelman3, Holly S. Stevenson3, Yonghong Wang3, Naoto Tsuchiya4, Takashi Kohno4, Vidar Skaug5, Steen Mollerup5, Aage Haugen5, Paul S. Meltzer3, Jun Yokota6, Yae Kanai2 and Curtis C. Harris1 Affiliations: 1Laboratory of Human Carcinogenesis, NCI-CCR, National Institutes of Health, Bethesda, MD 20892, USA. 2Division of Molecular Pathology, National Cancer Center Research Institute, Tokyo 104-0045, Japan. 3Genetics Branch, NCI-CCR, National Institutes of Health, Bethesda, MD 20892, USA. 4Division of Genome Biology, National Cancer Center Research Institute, Tokyo 104-0045, Japan. 5Department of Chemical and Biological Working Environment, National Institute of Occupational Health, NO-0033 Oslo, Norway. 6Genomics and Epigenomics of Cancer Prediction Program, Institute of Predictive and Personalized Medicine of Cancer (IMPPC), 08916 Badalona (Barcelona), Spain. List of Supplementary Materials Supplementary Materials and Methods Fig. S1. Hierarchical clustering of based on CpG sites differentially-methylated in Stage I ADC compared to non-tumor adjacent tissues. Fig. S2. Confirmatory pyrosequencing analysis of DNA methylation at the HOXA9 locus in Stage I ADC from a subset of the NCI microarray cohort. 1 Fig. S3. Methylation Beta-values for HOXA9 probe cg26521404 in Stage I ADC samples from Japan. Fig. S4. Kaplan-Meier analysis of HOXA9 promoter methylation in a published cohort of Stage I lung ADC (J Clin Oncol 2013;31(32):4140-7). Fig. S5. Kaplan-Meier analysis of a combined prognostic biomarker in Stage I lung ADC. -

Sheep Genome Functional Annotation Reveals Proximal Regulatory Elements Contributed to the Evolution of Modern Breeds
ARTICLE DOI: 10.1038/s41467-017-02809-1 OPEN Sheep genome functional annotation reveals proximal regulatory elements contributed to the evolution of modern breeds Marina Naval-Sanchez1, Quan Nguyen1, Sean McWilliam1, Laercio R. Porto-Neto1, Ross Tellam1, Tony Vuocolo1, Antonio Reverter1, Miguel Perez-Enciso 2,3, Rudiger Brauning4, Shannon Clarke4, Alan McCulloch4, Wahid Zamani5, Saeid Naderi 6, Hamid Reza Rezaei7, Francois Pompanon 8, Pierre Taberlet8, Kim C. Worley9, Richard A. Gibbs9, Donna M. Muzny9, Shalini N. Jhangiani9, Noelle Cockett10, Hans Daetwyler11,12 & James Kijas1 1234567890():,; Domestication fundamentally reshaped animal morphology, physiology and behaviour, offering the opportunity to investigate the molecular processes driving evolutionary change. Here we assess sheep domestication and artificial selection by comparing genome sequence from 43 modern breeds (Ovis aries) and their Asian mouflon ancestor (O. orientalis)to identify selection sweeps. Next, we provide a comparative functional annotation of the sheep genome, validated using experimental ChIP-Seq of sheep tissue. Using these annotations, we evaluate the impact of selection and domestication on regulatory sequences and find that sweeps are significantly enriched for protein coding genes, proximal regulatory elements of genes and genome features associated with active transcription. Finally, we find individual sites displaying strong allele frequency divergence are enriched for the same regulatory features. Our data demonstrate that remodelling of gene expression is likely to have been one of the evolutionary forces that drove phenotypic diversification of this common livestock species. 1 CSIRO Agriculture and Food, 306 Carmody Road, St. Lucia 4067 QLD, Australia. 2 Centre for Research in Agricultural Genomics (CRAG), Bellaterra 08193, Spain. 3 ICREA, Carrer de Lluís Companys 23, Barcelona 08010, Spain. -
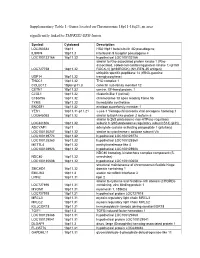
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

Investigating Contributions of Trisomy 21 in Down Syndrome to Alzheimer Disease Phenotypes in a Novel Mouse Cross
Investigating contributions of trisomy 21 in Down syndrome to Alzheimer disease phenotypes in a novel mouse cross A thesis presented in partial fulfilment of the requirements for the degree of Doctor of Philosophy from University College London Xun Yu Choong Department of Neurodegenerative Disease Institute of Neurology University College London 2016 1 Declaration I, Xun Yu Choong, confirm that the work presented in this thesis is my own. Where information has been derived from other sources, I confirm that this has been indicated in the thesis. 2 Acknowledgements The work done in this project was only possible with the support of numerous friends and colleagues, with whom I have grown an incredible amount. Foremost thanks go to Prof. Elizabeth Fisher and Dr. Frances Wiseman, who have been inexhaustibly dedicated supervisors and inspirational figures. Special mention also goes to two unofficial mentors who have looked out for me throughout the PhD, Dr. Karen Cleverley and Dr. Sarah Mizielinska. The Fisher and Isaacs groups have been a joy to work with and have always unhesitatingly offered time and help. Thank you to the Down syndrome group: Olivia Sheppard, Dr. Toby Collins, Dr. Suzanna Noy, Amy Nick, Laura Pulford, Justin Tosh, Matthew Rickman; other members of Lizzy’s group: Dr. Anny Devoy, Dr. Rosie Bunton-Stasyshyn, Dr. Rachele Saccon, Julian Pietrzyk, Dr. Beverley Burke, Heesoon Park, Julian Jaeger; the Isaacs group: Dr. Adrian Isaacs, Dr. Emma Clayton, Dr. Roberto Simone, Charlotte Ridler, Frances Norona. The PhD office has also been a second home in more ways than one, thanks to: Angelos Armen, Dr. -

Cysteine Proteases in Protozoan Parasites
REVIEW Cysteine proteases in protozoan parasites Jair L. Siqueira-Neto1*, Anjan Debnath1, Laura-Isobel McCall1¤, Jean A. Bernatchez1, Momar Ndao2,3, Sharon L. Reed4, Philip J. Rosenthal5 1 Center for Discovery and Innovation in Parasitic Diseases, Skaggs School of Pharmacy and Pharmaceutical Sciences, University of California San Diego, La Jolla, California, United States of America, 2 National Reference Centre for Parasitology, The Research Institute of the McGill University Health Center, Montreal, Canada, 3 Program in Infectious Diseases and Immunity in Global Health, The Research Institute of the McGill University Health Centre, Montreal, Quebec, Canada, 4 Departments of Pathology and Medicine, University of California San Diego School of Medicine, La Jolla, California, United States of America, 5 Department of Medicine, University of California, San Francisco, San Francisco, California, a1111111111 United States of America a1111111111 a1111111111 ¤ Current address: Department of Chemistry and Biochemistry, University of Oklahoma, Norman, Oklahoma, a1111111111 United States of America a1111111111 * [email protected] Abstract OPEN ACCESS Cysteine proteases (CPs) play key roles in the pathogenesis of protozoan parasites, includ- Citation: Siqueira-Neto JL, Debnath A, McCall L-I, ing cell/tissue penetration, hydrolysis of host or parasite proteins, autophagy, and evasion Bernatchez JA, Ndao M, Reed SL, et al. (2018) or modulation of the host immune response, making them attractive chemotherapeutic and Cysteine proteases in protozoan parasites. PLoS vaccine targets. This review highlights current knowledge on clan CA cysteine proteases, Negl Trop Dis 12(8): e0006512. https://doi.org/ 10.1371/journal.pntd.0006512 the best-characterized group of cysteine proteases, from 7 protozoan organisms causing human diseases with significant impact: Entamoeba histolytica, Leishmania species (sp.), Editor: Photini Sinnis, Johns Hopkins Bloomberg School of Public Health, UNITED STATES Trypanosoma brucei, T.