Title Systems Level Analysis of Transgenerational Spermatogenic
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Gene Expression Resource Generated by Genome-Wide Lacz
© 2015. Published by The Company of Biologists Ltd | Disease Models & Mechanisms (2015) 8, 1467-1478 doi:10.1242/dmm.021238 RESOURCE ARTICLE A gene expression resource generated by genome-wide lacZ profiling in the mouse Elizabeth Tuck1,**, Jeanne Estabel1,*,**, Anika Oellrich1, Anna Karin Maguire1, Hibret A. Adissu2, Luke Souter1, Emma Siragher1, Charlotte Lillistone1, Angela L. Green1, Hannah Wardle-Jones1, Damian M. Carragher1,‡, Natasha A. Karp1, Damian Smedley1, Niels C. Adams1,§, Sanger Institute Mouse Genetics Project1,‡‡, James N. Bussell1, David J. Adams1, Ramiro Ramırez-Soliś 1, Karen P. Steel1,¶, Antonella Galli1 and Jacqueline K. White1,§§ ABSTRACT composite of RNA-based expression data sets. Strong agreement was observed, indicating a high degree of specificity in our data. Knowledge of the expression profile of a gene is a critical piece of Furthermore, there were 1207 observations of expression of a information required to build an understanding of the normal and particular gene in an anatomical structure where Bgee had no essential functions of that gene and any role it may play in the data, indicating a large amount of novelty in our data set. development or progression of disease. High-throughput, large- Examples of expression data corroborating and extending scale efforts are on-going internationally to characterise reporter- genotype-phenotype associations and supporting disease gene tagged knockout mouse lines. As part of that effort, we report an candidacy are presented to demonstrate the potential of this open access adult mouse expression resource, in which the powerful resource. expression profile of 424 genes has been assessed in up to 47 different organs, tissues and sub-structures using a lacZ reporter KEY WORDS: Gene expression, lacZ reporter, Mouse, Resource gene. -

Mouse Dym Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Dym Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Dym conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Dym gene (NCBI Reference Sequence: NM_027727 ; Ensembl: ENSMUSG00000035765 ) is located on Mouse chromosome 18. 17 exons are identified, with the ATG start codon in exon 2 and the TGA stop codon in exon 17 (Transcript: ENSMUST00000039608). Exon 3 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Dym gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP23-46L22 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for a gene trapped allele display decreased body size with short tubular bones, chondrodysplasia, partial penetrance of obstructive hydronephrosis and impaired vesicular transport. Exon 3 starts from about 7.03% of the coding region. The knockout of Exon 3 will result in frameshift of the gene. The size of intron 2 for 5'-loxP site insertion: 9711 bp, and the size of intron 3 for 3'-loxP site insertion: 2247 bp. The size of effective cKO region: ~553 bp. The cKO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele gRNA region 5' gRNA region 3' 1 3 4 17 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Dym Homology arm cKO region loxP site Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

An Association Study Between the Dymeclin Gene and Schizophrenia in the Japanese Population
Journal of Human Genetics (2010) 55, 631–634 & 2010 The Japan Society of Human Genetics All rights reserved 1434-5161/10 $32.00 www.nature.com/jhg SHORT COMMUNICATION An association study between the dymeclin gene and schizophrenia in the Japanese population Saori Yazaki1, Minori Koga1,2, Hiroki Ishiguro1,2, Toshiya Inada3, Hiroshi Ujike4, Masanari Itokawa5, Takeshi Otowa6, Yuichiro Watanabe7, Toshiyuki Someya7, Nakao Iwata8, Hiroshi Kunugi9, Norio Ozaki10 and Tadao Arinami1,2 Many gene variants are involved in the susceptibility to schizophrenia and some of them are expected to be associated with other human characters. Recently reported meta-analysis of genetic associations revealed nucleotide variants in synaptic vesicular transport/Golgi apparatus genes with schizophrenia. In this study, we selected the dymeclin gene (DYM) as a candidate gene for schizophrenia. The DYM gene encodes dymeclin that has been identified to be associated with the Golgi apparatus and with transitional vesicles of the reticulum–Golgi interface. A three-step case–control study of total of 2105 Japanese cases of schizophrenia and 2087 Japanese control subjects was carried out for tag single-nucleotide polymorphisms (SNPs) in the DYM gene and an association between an SNP, rs833497, and schizophrenia was identified (allelic P¼2Â10À5, in the total sample). DYM is the causal gene for Dyggve–Melchior–Clausen syndrome and this study shows the second neuropsychiatric disorder in which the DYM gene is involved. The present data support the involvement of Golgi function and vesicular transport in the presynapse in schizophrenia. Journal of Human Genetics (2010) 55, 631–634; doi:10.1038/jhg.2010.72; published online 17 June 2010 Keywords: association; Dyggve–Melchior–Clausen syndrome; Golgi; postmortem study; SNP INTRODUCTION endoplasmic reticulum in some cell bodies of GABAergic neurons Schizophrenia is a chronic, severe and disabling brain disorder that in the cortex and hippocampus,5and genetic associations of the reelin affects approximately 1% of the world’s population. -

The Active Compounds and Therapeutic Mechanisms of Pentaherbs Formula for Oral and Topical Treatment of Atopic Dermatitis Based on Network Pharmacology
plants Article The Active Compounds and Therapeutic Mechanisms of Pentaherbs Formula for Oral and Topical Treatment of Atopic Dermatitis Based on Network Pharmacology Man Chu 1, Miranda Sin-Man Tsang 2,3, Ru He 1, Christopher Wai-Kei Lam 4, Zhi Bo Quan 1,* and Chun Kwok Wong 2,3,5,* 1 Faulty of Medical Technology, Shaanxi University of Chinese Medicine, Xianyang 712046, China; [email protected] (M.C.); [email protected] (R.H.) 2 Department of Chemical Pathology, The Chinese University of Hong Kong, Prince of Wales Hospital, Shatin, NT, Hong Kong 999077, China; [email protected] 3 State Key Laboratory of Research on Bioactivities and Clinical Applications of Medicinal Plants and Institute of Chinese Medicine, The Chinese University of Hong Kong, Shatin, NT, Hong Kong 999077, China 4 State Key Laboratory of Quality Research in Chinese Medicines and Faculty of Medicine, Macau University of Science and Technology, Macau 999078, China; [email protected] 5 Li Dak Sum Yip Yio Chin R & D Centre for Chinese Medicine, The Chinese University of Hong Kong, Shatin, NT, Hong Kong 999077, China * Correspondence: [email protected] (Z.B.Q.); [email protected] (C.K.W.); Tel.: +86-29-3818-5196 (Z.B.Q.); +852-3505-2964 (C.K.W.); Fax: +86-29-3818-5196 (Z.B.Q.); +852-2636-5090 (C.K.W.) Received: 23 July 2020; Accepted: 7 September 2020; Published: 9 September 2020 Abstract: To examine the molecular targets and therapeutic mechanism of a clinically proven Chinese medicinal pentaherbs formula (PHF) in atopic dermatitis (AD), we analyzed the active compounds and core targets, performed network and molecular docking analysis, and investigated interacting pathways. -

Dual Specificity Phosphatases from Molecular Mechanisms to Biological Function
International Journal of Molecular Sciences Dual Specificity Phosphatases From Molecular Mechanisms to Biological Function Edited by Rafael Pulido and Roland Lang Printed Edition of the Special Issue Published in International Journal of Molecular Sciences www.mdpi.com/journal/ijms Dual Specificity Phosphatases Dual Specificity Phosphatases From Molecular Mechanisms to Biological Function Special Issue Editors Rafael Pulido Roland Lang MDPI • Basel • Beijing • Wuhan • Barcelona • Belgrade Special Issue Editors Rafael Pulido Roland Lang Biocruces Health Research Institute University Hospital Erlangen Spain Germany Editorial Office MDPI St. Alban-Anlage 66 4052 Basel, Switzerland This is a reprint of articles from the Special Issue published online in the open access journal International Journal of Molecular Sciences (ISSN 1422-0067) from 2018 to 2019 (available at: https: //www.mdpi.com/journal/ijms/special issues/DUSPs). For citation purposes, cite each article independently as indicated on the article page online and as indicated below: LastName, A.A.; LastName, B.B.; LastName, C.C. Article Title. Journal Name Year, Article Number, Page Range. ISBN 978-3-03921-688-8 (Pbk) ISBN 978-3-03921-689-5 (PDF) c 2019 by the authors. Articles in this book are Open Access and distributed under the Creative Commons Attribution (CC BY) license, which allows users to download, copy and build upon published articles, as long as the author and publisher are properly credited, which ensures maximum dissemination and a wider impact of our publications. The book as a whole is distributed by MDPI under the terms and conditions of the Creative Commons license CC BY-NC-ND. Contents About the Special Issue Editors .................................... -
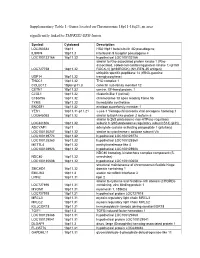
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

393LN V 393P 344SQ V 393P Probe Set Entrez Gene
393LN v 393P 344SQ v 393P Entrez fold fold probe set Gene Gene Symbol Gene cluster Gene Title p-value change p-value change chemokine (C-C motif) ligand 21b /// chemokine (C-C motif) ligand 21a /// chemokine (C-C motif) ligand 21c 1419426_s_at 18829 /// Ccl21b /// Ccl2 1 - up 393 LN only (leucine) 0.0047 9.199837 0.45212 6.847887 nuclear factor of activated T-cells, cytoplasmic, calcineurin- 1447085_s_at 18018 Nfatc1 1 - up 393 LN only dependent 1 0.009048 12.065 0.13718 4.81 RIKEN cDNA 1453647_at 78668 9530059J11Rik1 - up 393 LN only 9530059J11 gene 0.002208 5.482897 0.27642 3.45171 transient receptor potential cation channel, subfamily 1457164_at 277328 Trpa1 1 - up 393 LN only A, member 1 0.000111 9.180344 0.01771 3.048114 regulating synaptic membrane 1422809_at 116838 Rims2 1 - up 393 LN only exocytosis 2 0.001891 8.560424 0.13159 2.980501 glial cell line derived neurotrophic factor family receptor alpha 1433716_x_at 14586 Gfra2 1 - up 393 LN only 2 0.006868 30.88736 0.01066 2.811211 1446936_at --- --- 1 - up 393 LN only --- 0.007695 6.373955 0.11733 2.480287 zinc finger protein 1438742_at 320683 Zfp629 1 - up 393 LN only 629 0.002644 5.231855 0.38124 2.377016 phospholipase A2, 1426019_at 18786 Plaa 1 - up 393 LN only activating protein 0.008657 6.2364 0.12336 2.262117 1445314_at 14009 Etv1 1 - up 393 LN only ets variant gene 1 0.007224 3.643646 0.36434 2.01989 ciliary rootlet coiled- 1427338_at 230872 Crocc 1 - up 393 LN only coil, rootletin 0.002482 7.783242 0.49977 1.794171 expressed sequence 1436585_at 99463 BB182297 1 - up 393 -

Signatures of Adaptive Evolution in Platyrrhine Primate Genomes 5 6 Hazel Byrne*, Timothy H
1 2 Supplementary Materials for 3 4 Signatures of adaptive evolution in platyrrhine primate genomes 5 6 Hazel Byrne*, Timothy H. Webster, Sarah F. Brosnan, Patrícia Izar, Jessica W. Lynch 7 *Corresponding author. Email [email protected] 8 9 10 This PDF file includes: 11 Section 1: Extended methods & results: Robust capuchin reference genome 12 Section 2: Extended methods & results: Signatures of selection in platyrrhine genomes 13 Section 3: Extended results: Robust capuchins (Sapajus; H1) positive selection results 14 Section 4: Extended results: Gracile capuchins (Cebus; H2) positive selection results 15 Section 5: Extended results: Ancestral Cebinae (H3) positive selection results 16 Section 6: Extended results: Across-capuchins (H3a) positive selection results 17 Section 7: Extended results: Ancestral Cebidae (H4) positive selection results 18 Section 8: Extended results: Squirrel monkeys (Saimiri; H5) positive selection results 19 Figs. S1 to S3 20 Tables S1–S3, S5–S7, S10, and S23 21 References (94 to 172) 22 23 Other Supplementary Materials for this manuscript include the following: 24 Tables S4, S8, S9, S11–S22, and S24–S44 1 25 1) Extended methods & results: Robust capuchin reference genome 26 1.1 Genome assembly: versions and accessions 27 The version of the genome assembly used in this study, Sape_Mango_1.0, was uploaded to a 28 Zenodo repository (see data availability). An assembly (Sape_Mango_1.1) with minor 29 modifications including the removal of two short scaffolds and the addition of the mitochondrial 30 genome assembly was uploaded to NCBI under the accession JAGHVQ. The BioProject and 31 BioSample NCBI accessions for this project and sample (Mango) are PRJNA717806 and 32 SAMN18511585. -

Sheet1 Page 1 Gene Symbol Gene Description Entrez Gene ID
Sheet1 RefSeq ID ProbeSets Gene Symbol Gene Description Entrez Gene ID Sequence annotation Seed matches location(s) Ago-2 binding specific enrichment (replicate 1) Ago-2 binding specific enrichment (replicate 2) OE lysate log2 fold change (replicate 1) OE lysate log2 fold change (replicate 2) Probability NM_022823 218843_at FNDC4 Homo sapiens fibronectin type III domain containing 4 (FNDC4), mRNA. 64838 TR(1..1649)CDS(367..1071) 1523..1530 3.73 1.77 -1.91 -0.39 1 NM_003919 204688_at SGCE Homo sapiens sarcoglycan, epsilon (SGCE), transcript variant 2, mRNA. 8910 TR(1..1709)CDS(112..1425) 1495..1501 3.09 1.56 -1.02 -0.27 1 NM_006982 206837_at ALX1 Homo sapiens ALX homeobox 1 (ALX1), mRNA. 8092 TR(1..1320)CDS(5..985) 916..923 2.99 1.93 -0.19 -0.33 1 NM_019024 233642_s_at HEATR5B Homo sapiens HEAT repeat containing 5B (HEATR5B), mRNA. 54497 TR(1..6792)CDS(97..6312) 5827..5834,4309..4315 3.28 1.51 -0.92 -0.23 1 NM_018366 223431_at CNO Homo sapiens cappuccino homolog (mouse) (CNO), mRNA. 55330 TR(1..1546)CDS(96..749) 1062..1069,925..932 2.89 1.51 -1.2 -0.41 1 NM_032436 226194_at C13orf8 Homo sapiens chromosome 13 open reading frame 8 (C13orf8), mRNA. 283489 TR(1..3782)CDS(283..2721) 1756..1762,3587..3594,1725..1731,3395..3402 2.75 1.72 -1.38 -0.34 1 NM_031450 221534_at C11orf68 Homo sapiens chromosome 11 open reading frame 68 (C11orf68), mRNA. 83638 TR(1..1568)CDS(153..908) 967..973 3.07 1.35 -0.72 -0.06 1 NM_033318 225795_at,225794_s_at C22orf32 Homo sapiens chromosome 22 open reading frame 32 (C22orf32), mRNA. -

ALKBH5 Is a Mammalian RNA Demethylase That Impacts RNA Metabolism and Mouse Fertility
Molecular Cell Article ALKBH5 Is a Mammalian RNA Demethylase that Impacts RNA Metabolism and Mouse Fertility Guanqun Zheng,1,11 John Arne Dahl,3,11 Yamei Niu,2,11 Peter Fedorcsak,4 Chun-Min Huang,2 Charles J. Li,1 Cathrine B. Va˚ gbø,6 Yue Shi,2,7 Wen-Ling Wang,2,7 Shu-Hui Song,5 Zhike Lu,1 Ralph P.G. Bosmans,1 Qing Dai,1 Ya-Juan Hao,2,7 Xin Yang,2,7 Wen-Ming Zhao,5 Wei-Min Tong,8 Xiu-Jie Wang,9 Florian Bogdan,3 Kari Furu,3 Ye Fu,1 Guifang Jia,1 Xu Zhao,2,7 Jun Liu,10 Hans E. Krokan,6 Arne Klungland,3,* Yun-Gui Yang,2,7,* and Chuan He1,* 1Department of Chemistry, Institute for Biophysical Dynamics, The University of Chicago, 929 East 57th Street, Chicago, IL 60637, USA 2Genome Structure & Stability Group, BIG CAS-OSLO Genome Research Cooperation, Disease Genomics and Individualized Medicine Laboratory, Beijing Institute of Genomics, Chinese Academy of Sciences, No.7 Beitucheng West Road, Chaoyang District, Beijing 100029, P.R. China 3Centre for Molecular Biology and Neuroscience, Institute of Medical Microbiology, BIG CAS-OSLO Genome Research Cooperation, Oslo University Hospital, Oslo 0027, Norway 4Department of Gynecology, Oslo University Hospital, Rikshospitalet, Oslo 0027, Norway 5Core Genomic Facility, Beijing Institute of Genomics, Chinese Academy of Sciences, No.7 Beitucheng West Road, Chaoyang District, Beijing 100029, P.R. China 6Department of Cancer Research and Molecular Medicine, Norwegian University of Science and Technology, 7489 Trondheim, Norway 7University of Chinese Academy of Sciences, 19A Yuquan Road, Beijing 100049, P.R. -
Expression Profiling of WD40 Family Genes Including DDB1- and CUL4
Mistry et al. BMC Genomics (2020) 21:602 https://doi.org/10.1186/s12864-020-07016-9 RESEARCH ARTICLE Open Access Expression profiling of WD40 family genes including DDB1- and CUL4- associated factor (DCAF) genes in mice and human suggests important regulatory roles in testicular development and spermatogenesis Bhavesh V. Mistry1, Maha Alanazi1, Hanae Fitwi1,2, Olfat Al-Harazi3, Mohamed Rajab1, Abdullah Altorbag1, Falah Almohanna1, Dilek Colak2 and Abdullah M. Assiri1,3,4* Abstract Background: The WD40-repeat containing proteins, including DDB1–CUL4-associated factors (DCAFs), are abundant and conserved proteins that play important roles in different cellular processes including spermatogenesis. DCAFs are subset of WD40 family proteins that contain WDxR motif and have been proposed to function as substrate receptor for Cullin4-RING-based E3 ubiquitin ligase complexes to recruit diverse proteins for ubiquitination, a vital process in spermatogenesis. Large number of WD40 genes has been identified in different species including mouse and human. However, a systematic expression profiling of WD40 genes in different tissues of mouse and human has not been investigated. We hypothesize that large number of WD40 genes may express highly or specifically in the testis, where their expression is uniquely regulated during testis development and spermatogenesis. Therefore, the objective of this study is to mine and characterize expression patterns of WD40 genes in different tissues of mouse and human with particular emphasis on DCAF genes expressions during mouse testicular development. (Continued on next page) * Correspondence: [email protected] 1Department of Comparative Medicine, King Faisal Specialist Hospital & Research Centre, Riyadh, Saudi Arabia 3Biostatistics, Epidemiology and Scientific Computing Department, King Faisal Specialist Hospital & Research Centre, Riyadh, Saudi Arabia Full list of author information is available at the end of the article © The Author(s). -

Mouse Endogenous Retroviruses Can Trigger Premature Transcriptional Termination at a Distance
Downloaded from genome.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Research Mouse endogenous retroviruses can trigger premature transcriptional termination at a distance Jingfeng Li,1 Keiko Akagi,1 Yongjun Hu,2 Anna L. Trivett,3 Christopher J.W. Hlynialuk,1 Deborah A. Swing,4 Natalia Volfovsky,5 Tamara C. Morgan,6 Yelena Golubeva,6 Robert M. Stephens,5 David E. Smith,2 and David E. Symer1,7,8 1Human Cancer Genetics Program and Department of Molecular Virology, Immunology and Medical Genetics, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio 43210, USA; 2Department of Pharmaceutical Sciences, University of Michigan, Ann Arbor, Michigan 48109, USA; 3Laboratory of Molecular Immunoregulation and 4Mouse Cancer Genetics Program, National Cancer Institute, Frederick, Maryland 21702, USA; 5Advanced Biomedical Computing Center, Information Systems Program and 6Histotechnology Laboratory, SAIC-Frederick, Inc., National Cancer Institute, Frederick, Maryland 21702, USA; 7Department of Internal Medicine and Department of Biomedical Informatics, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio 43210, USA Endogenous retrotransposons have caused extensive genomic variation within mammalian species, but the functional implications of such mobilization are mostly unknown. We mapped thousands of endogenous retrovirus (ERV) germline integrants in highly divergent, previously unsequenced mouse lineages, facilitating a comparison of gene expression in the presence or absence of