Positive Gastric Cancer with Sequential Genomic Profiling
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Targeting Synthetic Lethality Between the SRC Kinase and the EPHB6 Receptor May Benefit Cancer Treatment
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 7, No. 31 Research Paper Targeting synthetic lethality between the SRC kinase and the EPHB6 receptor may benefit cancer treatment James M. Paul1,*, Behzad Toosi2,*, Frederick S. Vizeacoumar2,*, Kalpana Kalyanasundaram Bhanumathy2, Yue Li3,4,5, Courtney Gerger2, Amr El Zawily2,6, Tanya Freywald7, Deborah H. Anderson7, Darrell Mousseau8, Rani Kanthan2, Zhaolei Zhang3,4, Franco J. Vizeacoumar2,7, Andrew Freywald2 1Department of Biochemistry, University of Saskatchewan, Saskatoon, SK, S7N 5E5, Canada 2Department of Pathology and Laboratory Medicine, College of Medicine, University of Saskatchewan, Royal University Hospital, Saskatoon, SK, S7N 0W8, Canada 3Department of Computer Science, University of Toronto, Toronto, ON, M5S 3G4, Canada 4The Donnelly Centre, University of Toronto, Toronto, ON, M5S 3E1, Canada 5Present address: Computer Science and Artificial Intelligence Laboratory, Massachusetts Institute of Technology, Cambridge, MA, 02139, USA 6Faculty of Science, Damanhour University, Damanhour, 22516, Egypt 7Cancer Research, Saskatchewan Cancer Agency, Saskatoon, SK, S7N 5E5, Canada 8Cell Signaling Laboratory, Neuroscience Cluster, University of Saskatchewan, Saskatoon, SK, S7N 5E5, Canada *These authors contributed equally to this work Correspondence to: Franco J. Vizeacoumar, email: [email protected] Andrew Freywald, email: [email protected] Keywords: breast cancer, genetic interaction, synthetic lethality, EPHB6, SRC kinase Received: April 22, 2016 Accepted: June 17, 2016 Published: July 13, 2016 ABSTRACT Application of tumor genome sequencing has identified numerous loss-of-function alterations in cancer cells. While these alterations are difficult to target using direct interventions, they may be attacked with the help of the synthetic lethality (SL) approach. In this approach, inhibition of one gene causes lethality only when another gene is also completely or partially inactivated. -

Human RET Kinase Protein (His Tag)
Human RET Kinase Protein (His Tag) Catalog Number: 11997-H08H1 General Information SDS-PAGE: Gene Name Synonym: CDHF12; CDHR16; HSCR1; MEN2A; MEN2B; MTC1; PTC; RET-ELE1; RET51 Protein Construction: A DNA sequence encoding the extracellular domain of human RET (P07949-1) (Met 1-Arg 635) was fused with a polyhistidine tag at the N- terminus. Source: Human Expression Host: HEK293 Cells QC Testing Purity: > 92 % as determined by SDS-PAGE Protein Description Endotoxin: RET proto-oncogene, also known as RET, is a cell-surface molecule that < 1.0 EU per μg of the protein as determined by the LAL method transduce signals for cell growth and differentiation. It contains 1 cadherin domain and 1 protein kinase domain. RET proto-oncogene belongs to the Stability: protein kinase superfamily, tyr protein kinase family. RET proto-oncogene is involved in numerous cellular mechanisms including cell proliferation, ℃ Samples are stable for up to twelve months from date of receipt at -70 neuronal navigation, cell migration, and cell differentiation upon binding with glial cell derived neurotrophic factor family ligands. It phosphorylates Leu 29 Predicted N terminal: PTK2/FAK1 and regulates both cell death/survival balance and positional Molecular Mass: information. RET is required for the molecular mechanisms orchestration during intestine organogenesis; involved in the development of enteric The recombinant human RET consists of 618 amino acids and has a nervous system and renal organogenesis during embryonic life; promotes calculated molecular mass of 69.1 kDa. The apparent molecular mass of the formation of Peyer's patch-like structures; modulates cell adhesion via the protein is approximately 110-120 kDa in SDS-PAGE under reducing its cleavage; involved in the development of the neural crest. -

Constitutive Scaffolding of Multiple Wnt Enhanceosome Components By
RESEARCH ARTICLE Constitutive scaffolding of multiple Wnt enhanceosome components by Legless/ BCL9 Laurens M van Tienen, Juliusz Mieszczanek, Marc Fiedler, Trevor J Rutherford, Mariann Bienz* MRC Laboratory of Molecular Biology, Cambridge, United Kingdom Abstract Wnt/b-catenin signaling elicits context-dependent transcription switches that determine normal development and oncogenesis. These are mediated by the Wnt enhanceosome, a multiprotein complex binding to the Pygo chromatin reader and acting through TCF/LEF- responsive enhancers. Pygo renders this complex Wnt-responsive, by capturing b-catenin via the Legless/BCL9 adaptor. We used CRISPR/Cas9 genome engineering of Drosophila legless (lgs) and human BCL9 and B9L to show that the C-terminus downstream of their adaptor elements is crucial for Wnt responses. BioID proximity labeling revealed that BCL9 and B9L, like PYGO2, are constitutive components of the Wnt enhanceosome. Wnt-dependent docking of b-catenin to the enhanceosome apparently causes a rearrangement that apposes the BCL9/B9L C-terminus to TCF. This C-terminus binds to the Groucho/TLE co-repressor, and also to the Chip/LDB1-SSDP enhanceosome core complex via an evolutionary conserved element. An unexpected link between BCL9/B9L, PYGO2 and nuclear co-receptor complexes suggests that these b-catenin co-factors may coordinate Wnt and nuclear hormone responses. DOI: 10.7554/eLife.20882.001 *For correspondence: mb2@mrc- Introduction lmb.cam.ac.uk The Wnt/b-catenin signaling cascade is an ancient cell communication pathway that operates con- Competing interests: The text-dependent transcriptional switches to control animal development and tissue homeostasis authors declare that no (Cadigan and Nusse, 1997). -

SUPPLEMENTARY FIGURES and TABLES Genetic Hallmarks of Recurrent/Metastatic Adenoid Cystic Carcinoma
SUPPLEMENTARY FIGURES AND TABLES Genetic Hallmarks Of Recurrent/Metastatic Adenoid Cystic Carcinoma SUPPLEMENTARY FIGURES Figure S1. Flow diagram of study design. Figure S2. Primary and recurrent/metastatic adenoid cystic carcinoma distribution by anatomic site. Figure S3. Variant allelic frequency (VAF) density histogram for NOTCH1 mutations observed in recurrent/metastatic adenoid cystic carcinoma (R/M ACC). Figure S4. Downsampling analysis of R/M MSK-IMPACT cohort (n=94) to simulate mutation detection at 100x coverage. Figure S5. Representative PyClone plots demonstrating intratumoral heterogeneity quantified as number of genomically distinct subclonal populations in adenoid cystic carcinoma. Figure S6. MRI of the neck with contrast of adenoid cystic carcinoma of right parotid gland involving masseter muscle and ascending ramus. Figure S7. Histologic confirmation of 6 representative distant metastatic sites of a single case with parotid adenoid cystic carcinoma. Figure S8. Fluorescence in situ hybridization (FISH) of distant lung metastatic lesions in a single case of parotid adenoid cystic carcinoma. Figure S9. Two-way plots of cancer cell fraction in a single case of parotid adenoid cystic carcinoma, comparing primary tumor with eight metastatic lesions. Figure S10. Multiregion clonal evolution heatmap analysis of two breast adenoid cystic carcinoma cases with transformation to high grade triple-negative breast cancer (TNBC) histology. SUPPLEMENTARY TABLES Table S1. Study distribution of primary and recurrent/metastatic (R/M) adenoid cystic carcinoma (ACC) cases. Mixed entails head and neck, lung, and breast disease sites. Table S2. Top gene alteration incidence by tumor site (includes primary and recurrent/metastatic cases). Table S3. Top gene alteration incidence of recurrent/metastatic adenoid cystic carcinoma (R/M ACC) cases comparing primary site with distant metastatic site. -

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -
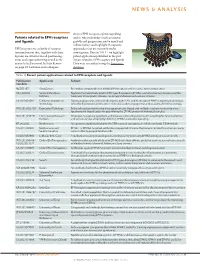
Patents Related to EPH Receptors and Ligands
NEWS & ANALYSIS discuss EPH receptor–ephrin signalling Patents related to EPH receptors and its role in disorders such as tumour and ligands growth and progression, nerve injury and inflammation, and highlight therapeutic EPH receptors are a family of receptor approaches that are currently under tyrosine kinases that, together with their investigation. Here in TABLE 1 we highlight ligands, are involved in cell positioning, patent applications published in the past tissue and organ patterning as well as the 3 years related to EPH receptors and ligands. control of cell survival. In their Review Data were researched using the Espacenet on page 39, Lackman and colleagues database. Table 1 | Recent patent applications related to EPH receptors and ligands Nature Reviews | Drug Discovery Publication Applicants Subject numbers NZ 581397 AstraZeneca Pyrimidine compounds that inhibit EPH receptors and are useful for treating cancer HK 1108702 Sanford-Burnham Peptides that selectively bind to EPH type-B receptors (EPHBs); useful for tumour imaging and the Institute treatment of neoplastic disease, neurological disease and vascular disease US 2013091591 California Institute of During angiogenesis, arterial cells express ephrin B2, and its receptor EPHB4 is expressed on venous Technology cells; this distinction can be used in methods to alter angiogenesis and to assess the effect of drugs WO 2013052710 Expression Pathology Selected reaction monitoring mass spectrometry-based and multiple reaction monitoring mass spectrometry-based assays for quantifying -

Human FLT4 / VEGFR3 ELISA Kit (ARG82047)
Product datasheet [email protected] ARG82047 Package: 96 wells Human FLT4 / VEGFR3 ELISA Kit Store at: 4°C Summary Product Description Human FLT4 / VEGFR3 ELISA Kit is an Enzyme Immunoassay kit for the quantification of Human FLT4 / VEGFR3 in serum, plasma and cell culture supernatants. Tested Reactivity Hu Tested Application ELISA Target Name FLT4 / VEGFR3 Conjugation HRP Conjugation Note Substrate: TMB and read at 450 nm. Sensitivity 78 pg/ml Sample Type Serum, plasma and cell culture supernatants. Standard Range 156 - 10000 pg/ml Sample Volume 100 µl Alternate Names FLT-4; FLT41; Vascular endothelial growth factor receptor 3; VEGFR3; VEGFR-3; PCL; Tyrosine-protein kinase receptor FLT4; LMPH1A; EC 2.7.10.1; Fms-like tyrosine kinase 4 Application Instructions Assay Time 4.5 hours Properties Form 96 well Storage instruction Store the kit at 2-8°C. Keep microplate wells sealed in a dry bag with desiccants. Do not expose test reagents to heat, sun or strong light during storage and usage. Please refer to the product user manual for detail temperatures of the components. Note For laboratory research only, not for drug, diagnostic or other use. Bioinformation Gene Symbol FLT4 Gene Full Name fms-related tyrosine kinase 4 Background This gene encodes a tyrosine kinase receptor for vascular endothelial growth factors C and D. The protein is thought to be involved in lymphangiogenesis and maintenance of the lymphatic endothelium. Mutations in this gene cause hereditary lymphedema type IA. [provided by RefSeq, Jul 2008] Function Tyrosine-protein kinase that acts as a cell-surface receptor for VEGFC and VEGFD, and plays an essential role in adult lymphangiogenesis and in the development of the vascular network and the cardiovascular system during embryonic development. -

The Kinase Defective EPHB6 Receptor Tyrosine Kinase Activates MAP Kinase Signaling in Lung Adenocarcinoma
175-179.qxd 29/5/2009 01:21 ÌÌ ™ÂÏ›‰·175 INTERNATIONAL JOURNAL OF ONCOLOGY 35: 175-179, 2009 175 The kinase defective EPHB6 receptor tyrosine kinase activates MAP kinase signaling in lung adenocarcinoma JUN YU1,2, ETMAR BULK1, PING JI1, ANTJE HASCHER1, STEFFEN KOSCHMIEDER1, WOLFGANG E. BERDEL1 and CARSTEN MÜLLER-TIDOW1 1Department of Medicine, Hematology and Oncology, University of Münster, Münster, Germany; 2Department of Preclinical Experiment Center, Fourth Military Medical University, Xi'an, P.R. China Received January 28, 2009; Accepted March 13, 2009 DOI: 10.3892/ijo_00000326 Abstract. Decreased expression levels of EPHB6, a member Elk-1 (8,9). ERK1/2 are negatively regulated by a family of of the receptor tyrosine kinases (RTKs), are associated with dual-specificity (Thr/Tyr) MAPK phosphatases, known as an increased risk of metastasis development in early stage DUSPs or MKPs, and pharmacologically by MEK inhibitors non-small cell lung cancer (NSCLC). However, the signaling such as U0126 and PD98059 (10). properties of the kinase-defective EPHB6 receptor are not EPH receptors form the largest known subfamily of well-understood. Here, we show that expression of EPHB6 receptor tyrosine kinases, and to date, the EPH subfamily in A549 lung adenocarinoma cells led to phosphorylation of contains 16 members in vertebrates (11,12). The EPH receptors the MAP kinase ERK. Conversely, siRNA based knockdown interact with a family of ligands located on the surfaces of of EPHB6 reversed ERK phosphorylation. Intriguingly, adjacent cells, named Ephrins including Ephrin-As and EPHB6-induced phosphorylation of ERK was uncoupled Ephrin-Bs subgroups. The EPH receptors are also grouped by activation of the Elk-1 transcriptional factor. -

A Review of VEGF/VEGFR-Targeted Therapeutics for Recurrent Glioblastoma
414 Original Article A Review of VEGF/VEGFR-Targeted Therapeutics for Recurrent Glioblastoma David A. Reardon, MDa,b; Scott Turner, MDc; Katherine B. Peters, MD, PhDc; Annick Desjardins, MDc; Sridharan Gururangan, MDa,b; John H. Sampson, MD, PhDa; Roger E. McLendon, MDd; James E. Herndon II, PhDe; Lee W. Jones, PhDf; John P. Kirkpatrick, MD, PhDf; Allan H. Friedman, MDa; James J. Vredenburgh, MDc; Darell D. Bigner, MD, PhDd; and Henry S. Friedman, MDa,b; Durham, North Carolina Key Words ability effect of VEGF inhibitors, the Radiologic Assessment in Neu- Glioblastoma, angiogenesis, vascular endothelial growth factor, ro-Oncology (RANO) criteria were recently implemented to bet- malignant glioma ter assess response among patients with glioblastoma. Although bevacizumab improves survival and quality of life, eventual tumor progression is the norm. Better understanding of resistance mech- Abstract anisms to VEGF inhibitors and identification of effective therapy Glioblastoma, the most common primary malignant brain tumor after bevacizumab progression are currently a critical need for pa- among adults, is a highly angiogenic and deadly tumor. Angiogen- tients with glioblastoma. (JNCCN 2011;9:414–427) esis in glioblastoma, driven by hypoxia-dependent and indepen- dent mechanisms, is primarily mediated by vascular endothelial growth factor (VEGF), and generates blood vessels with distinctive features. The outcome for patients with recurrent glioblastoma is poor because of ineffective therapies. However, recent encourag- ing rates of radiographic response and progression-free survival, Malignant gliomas, including the most common sub- and adequate safety, led the FDA to grant accelerated approval of type of glioblastoma, are rapidly growing destructive tu- bevacizumab, a humanized monoclonal antibody against VEGF, for mors that extensively invade locally but rarely metasta- the treatment of recurrent glioblastoma in May 2009. -

Nucleo-Cytoplasmic Distribution of ß-Catenin Is Regulated by Retention
Research Article 1453 Nucleo-cytoplasmic distribution of -catenin is regulated by retention Eva Krieghoff, Jürgen Behrens* and Bernhard Mayr Nikolaus-Fiebiger-Center for Molecular Medicine, University of Erlangen-Nürnberg, Glückstr. 6, 91054 Erlangen, Germany *Author for correspondence (e-mail: [email protected]) Accepted 19 December 2005 Journal of Cell Science 119, 1453-1463 Published by The Company of Biologists 2006 doi:10.1242/jcs.02864 Summary -catenin is the central signalling molecule of the canonical of -catenin, i.e. increases the rate of -catenin nuclear Wnt pathway, where it activates target genes in a complex import or export. Moreover, the cytoplasmic enrichment of with LEF/TCF transcription factors in the nucleus. The -catenin by APC and axin is not abolished by inhibition regulation of -catenin activity is thought to occur mainly of CRM-1-dependent nuclear export. TCF4, APC, axin and on the level of protein degradation, but it has been axin2 move more slowly than -catenin in their respective suggested that -catenin nuclear localization and hence its compartment, and concomitantly decrease -catenin transcriptional activity may additionally be regulated via mobility. Together, these data indicate that -catenin nuclear import by TCF4 and BCL9 and via nuclear export interaction partners mainly regulate -catenin subcellular by APC and axin. Using live-cell microscopy and localization by retaining it in the compartment in which fluorescence recovery after photobleaching (FRAP), we they are localized, rather than by active transport into or have directly analysed the impact of these factors on the out of the nucleus. subcellular localization of -catenin, its nucleo-cytoplasmic shuttling and its mobility within the nucleus and the Supplementary material available online at cytoplasm. -

Single-Cell RNA Sequencing Demonstrates the Molecular and Cellular Reprogramming of Metastatic Lung Adenocarcinoma
ARTICLE https://doi.org/10.1038/s41467-020-16164-1 OPEN Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma Nayoung Kim 1,2,3,13, Hong Kwan Kim4,13, Kyungjong Lee 5,13, Yourae Hong 1,6, Jong Ho Cho4, Jung Won Choi7, Jung-Il Lee7, Yeon-Lim Suh8,BoMiKu9, Hye Hyeon Eum 1,2,3, Soyean Choi 1, Yoon-La Choi6,10,11, Je-Gun Joung1, Woong-Yang Park 1,2,6, Hyun Ae Jung12, Jong-Mu Sun12, Se-Hoon Lee12, ✉ ✉ Jin Seok Ahn12, Keunchil Park12, Myung-Ju Ahn 12 & Hae-Ock Lee 1,2,3,6 1234567890():,; Advanced metastatic cancer poses utmost clinical challenges and may present molecular and cellular features distinct from an early-stage cancer. Herein, we present single-cell tran- scriptome profiling of metastatic lung adenocarcinoma, the most prevalent histological lung cancer type diagnosed at stage IV in over 40% of all cases. From 208,506 cells populating the normal tissues or early to metastatic stage cancer in 44 patients, we identify a cancer cell subtype deviating from the normal differentiation trajectory and dominating the metastatic stage. In all stages, the stromal and immune cell dynamics reveal ontological and functional changes that create a pro-tumoral and immunosuppressive microenvironment. Normal resident myeloid cell populations are gradually replaced with monocyte-derived macrophages and dendritic cells, along with T-cell exhaustion. This extensive single-cell analysis enhances our understanding of molecular and cellular dynamics in metastatic lung cancer and reveals potential diagnostic and therapeutic targets in cancer-microenvironment interactions. 1 Samsung Genome Institute, Samsung Medical Center, Seoul 06351, Korea. -

Profiling Data
Compound Name DiscoveRx Gene Symbol Entrez Gene Percent Compound Symbol Control Concentration (nM) JNK-IN-8 AAK1 AAK1 69 1000 JNK-IN-8 ABL1(E255K)-phosphorylated ABL1 100 1000 JNK-IN-8 ABL1(F317I)-nonphosphorylated ABL1 87 1000 JNK-IN-8 ABL1(F317I)-phosphorylated ABL1 100 1000 JNK-IN-8 ABL1(F317L)-nonphosphorylated ABL1 65 1000 JNK-IN-8 ABL1(F317L)-phosphorylated ABL1 61 1000 JNK-IN-8 ABL1(H396P)-nonphosphorylated ABL1 42 1000 JNK-IN-8 ABL1(H396P)-phosphorylated ABL1 60 1000 JNK-IN-8 ABL1(M351T)-phosphorylated ABL1 81 1000 JNK-IN-8 ABL1(Q252H)-nonphosphorylated ABL1 100 1000 JNK-IN-8 ABL1(Q252H)-phosphorylated ABL1 56 1000 JNK-IN-8 ABL1(T315I)-nonphosphorylated ABL1 100 1000 JNK-IN-8 ABL1(T315I)-phosphorylated ABL1 92 1000 JNK-IN-8 ABL1(Y253F)-phosphorylated ABL1 71 1000 JNK-IN-8 ABL1-nonphosphorylated ABL1 97 1000 JNK-IN-8 ABL1-phosphorylated ABL1 100 1000 JNK-IN-8 ABL2 ABL2 97 1000 JNK-IN-8 ACVR1 ACVR1 100 1000 JNK-IN-8 ACVR1B ACVR1B 88 1000 JNK-IN-8 ACVR2A ACVR2A 100 1000 JNK-IN-8 ACVR2B ACVR2B 100 1000 JNK-IN-8 ACVRL1 ACVRL1 96 1000 JNK-IN-8 ADCK3 CABC1 100 1000 JNK-IN-8 ADCK4 ADCK4 93 1000 JNK-IN-8 AKT1 AKT1 100 1000 JNK-IN-8 AKT2 AKT2 100 1000 JNK-IN-8 AKT3 AKT3 100 1000 JNK-IN-8 ALK ALK 85 1000 JNK-IN-8 AMPK-alpha1 PRKAA1 100 1000 JNK-IN-8 AMPK-alpha2 PRKAA2 84 1000 JNK-IN-8 ANKK1 ANKK1 75 1000 JNK-IN-8 ARK5 NUAK1 100 1000 JNK-IN-8 ASK1 MAP3K5 100 1000 JNK-IN-8 ASK2 MAP3K6 93 1000 JNK-IN-8 AURKA AURKA 100 1000 JNK-IN-8 AURKA AURKA 84 1000 JNK-IN-8 AURKB AURKB 83 1000 JNK-IN-8 AURKB AURKB 96 1000 JNK-IN-8 AURKC AURKC 95 1000 JNK-IN-8