EMBL-EBI Annual Scientific Report 2013
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Crystallography News British Crystallographic Association
Crystallography News British Crystallographic Association Issue No. 100 March 2007 ISSN 1467-2790 BCA Spring Meeting 2007 - Canterbury p8-17 Patrick Tollin (1938 - 2006) p7 The Z’ > 1 Phenomenon p18-19 History p21-23 Meetings of Interest p32 March 2007 Crystallography News Contents 2 . From the President 3 . Council Members 4 . BCA Letters to the Editor 5 Administrative Office, . Elaine Fulton, From the Editor 6 Northern Networking Events Ltd. 7 1 Tennant Avenue, Puzzle Corner College Milton South, . East Kilbride, Glasgow G74 5NA Scotland, UK Patrick Tollin (1938 - 2006) 8-17 Tel: + 44 1355 244966 Fax: + 44 1355 249959 . e-mail: [email protected] BCA 2007 Spring Meeting 16-17 . CRYSTALLOGRAPHY NEWS is published quarterly (March, June, BCA 2007 Meeting Timetable 18-19 September and December) by the British Crystallographic Association, . and printed by William Anderson and Sons Ltd, Glasgow. Text should The Z’ > 1 Phenomenon 20 preferably be sent electronically as MSword documents (any version - . .doc, .rtf or .txt files) or else on a PC disk. Diagrams and figures are most IUCr Computing Commission 21-23 welcome, but please send them separately from text as .jpg, .gif, .tif, or .bmp files. Items may include technical articles, news about people (e.g. History . 24-27 awards, honours, retirements etc.), reports on past meetings of interest to crystallographers, notices of future meetings, historical reminiscences, Groups .......................................................... 28-31 letters to the editor, book, hardware or software reviews. Please ensure that items for inclusion in the June 2007 issue are sent to the Editor to arrive Meetings . 32 before 25th April 2007. -

Ccp4 Newsletter on Protein Crystallography
CCP4 NEWSLETTER ON PROTEIN CRYSTALLOGRAPHY An informal Newsletter associated with the BBSRC Collaborative Computational Project No. 4 on Protein Crystallography. Number 41 Fall 2002 Contents CCP4 News 1. News from CCP4. Alun Ashton, Charles Ballard, Peter Briggs, Maeri Howard-Eales, Pryank Patel and Martyn Winn. 2. Developments with CCP4i: October 2002. Peter Briggs. 3. Developments with the CCP4 library II. Martyn Winn, Charles Ballard and Eugene Krissinel. General News 4. BioMed College at Daresbury. Gareth Jones. Software 5. Handling reflection data using the Clipper libraries. Kevin Cowtan, Department of Chemistry, University of York, UK. Theory and Techniques 6. Atomic displacement in incomplete models caused by optimisation of crystallographic criteria. P.V Afonine, Université Henri Poincaré, Nancy, France, Centre Charles Hermite, LORIA, Villers-lès-Nancy, France. 7. Modelling of bond electron density by Gaussian scatters at subatomic resolution. P. Afonine(1,2), V. Pichon-Pesme(1), N. Muzet(1), C. Jelsh(1), C Lecomte(1) and A. Urzhumtsev(1), (1) Université Henri Poincaré, Nancy, France, (2) Centre Charles Hermite, LORIA, Villers-lès-Nancy, France. 8. Bulk-solvent correction for use with the CCP4 version of AMoRe. Guido Capitani(1) and Andrei Fokine(2), (1) University of Zürich, Switzerland, (2) Université Henry Poincaré, Nancy, France. 9. Variation of solvent density and low-resolution ab initio phasing. Andrei Fokine, Université Henry Poincaré, Nancy, France. 10. Retrieval of lost reflections in high resolution Fourier syntheses by 'soft' solvent flattening. Natalia L. Lunina(1), Vladimir Y. Lunin(1) and Alberto D. Podjarny(2), (1) Russian Academy of Sciences, Russia, (2) CU Strasbourg, France. Bulletin Board 11. -

The EMBL-European Bioinformatics Institute the Hub for Bioinformatics in Europe
The EMBL-European Bioinformatics Institute The hub for bioinformatics in Europe Blaise T.F. Alako, PhD [email protected] www.ebi.ac.uk What is EMBL-EBI? • Part of the European Molecular Biology Laboratory • International, non-profit research institute • Europe’s hub for biological data, services and research The European Molecular Biology Laboratory Heidelberg Hamburg Hinxton, Cambridge Basic research Structural biology Bioinformatics Administration Grenoble Monterotondo, Rome EMBO EMBL staff: 1500 people Structural biology Mouse biology >60 nationalities EMBL member states Austria, Belgium, Croatia, Denmark, Finland, France, Germany, Greece, Iceland, Ireland, Israel, Italy, Luxembourg, the Netherlands, Norway, Portugal, Spain, Sweden, Switzerland and the United Kingdom Associate member state: Australia Who we are ~500 members of staff ~400 work in services & support >53 nationalities ~120 focus on basic research EMBL-EBI’s mission • Provide freely available data and bioinformatics services to all facets of the scientific community in ways that promote scientific progress • Contribute to the advancement of biology through basic investigator-driven research in bioinformatics • Provide advanced bioinformatics training to scientists at all levels, from PhD students to independent investigators • Help disseminate cutting-edge technologies to industry • Coordinate biological data provision throughout Europe Services Data and tools for molecular life science www.ebi.ac.uk/services Browse our services 9 What services do we provide? Labs around the -

Functional Effects Detailed Research Plan
GeCIP Detailed Research Plan Form Background The Genomics England Clinical Interpretation Partnership (GeCIP) brings together researchers, clinicians and trainees from both academia and the NHS to analyse, refine and make new discoveries from the data from the 100,000 Genomes Project. The aims of the partnerships are: 1. To optimise: • clinical data and sample collection • clinical reporting • data validation and interpretation. 2. To improve understanding of the implications of genomic findings and improve the accuracy and reliability of information fed back to patients. To add to knowledge of the genetic basis of disease. 3. To provide a sustainable thriving training environment. The initial wave of GeCIP domains was announced in June 2015 following a first round of applications in January 2015. On the 18th June 2015 we invited the inaugurated GeCIP domains to develop more detailed research plans working closely with Genomics England. These will be used to ensure that the plans are complimentary and add real value across the GeCIP portfolio and address the aims and objectives of the 100,000 Genomes Project. They will be shared with the MRC, Wellcome Trust, NIHR and Cancer Research UK as existing members of the GeCIP Board to give advance warning and manage funding requests to maximise the funds available to each domain. However, formal applications will then be required to be submitted to individual funders. They will allow Genomics England to plan shared core analyses and the required research and computing infrastructure to support the proposed research. They will also form the basis of assessment by the Project’s Access Review Committee, to permit access to data. -
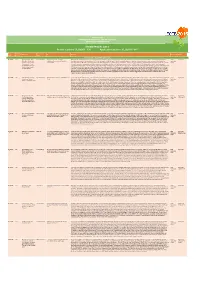
DATA Poster Numbers: P Da001 - 130 Application Posters: P Da001 - 041
POSTER LIST ORDERED ALPHABETICALLY BY POSTER TITLE GROUPED BY THEME/TRACK THEME/TRACK: DATA Poster numbers: P_Da001 - 130 Application posters: P_Da001 - 041 Poster EasyChair Presenting Author list Title Abstract Theme/track Topics number number author APPLICATION POSTERS WITHIN DATA THEME P_Da001 773 Benoît Carrères, Anne Benoît Carrères A systems approach to explore Microalgae are promising platforms for sustainable biofuel production. They produce triacyl-glycerides (TAG) which are easily converted into biofuel. When exposed to nitrogen limitation, Data/ Application Klok, Maria Suarez Diez, triacylglycerol production in Neochloris Neochloris oleoabundans accumulates up to 40% of its dry weight in TAG. However, a feasible production requires a decrease of production costs, which can be partially reached by Application Lenny de Jaeger, Mark oleoabundans increasing TAG yield.We built a constraint-based model describing primary metabolism of N. oleoabundans. It was grown in combinations of light absorption and nitrate supply rates and the poster Sturme, Packo Lamers, parameters needed for modeling of metabolism were measured. Fluxes were then calculated by flux balance analysis. cDNA samples of 16 experimental conditions were sequenced, Rene' Wijffels, Vitor Dos assembled and functionally annotated. Relative expression changes and relative flux changes for all reactions in the model were compared.The model predicts a maximum TAG yield on light Santos, Peter Schaap of 1.07g (mol photons)-1, more than 3 times current yield under optimal conditions. Furthermore, from optimization scenarios we concluded that increasing light efficiency has much higher and Dirk Martens potential to increase TAG yield than blocking entire pathways.Certain reaction expression patterns suggested an interdependence of the response to nitrogen and light supply. -

Exploring the Structure and Function Paradigm Oliver C Redfern, Benoit Dessailly and Christine a Orengo
Available online at www.sciencedirect.com Exploring the structure and function paradigm Oliver C Redfern, Benoit Dessailly and Christine A Orengo Advances in protein structure determination, led by the Figure 1 shows that as the international genomics initiat- structural genomics initiatives have increased the proportion of ives gather pace, both the number of sequences and novel folds deposited in the Protein Data Bank. However, these protein families are still growing at an exponential rate, structures are often not accompanied by functional annotations although the rate of expansion of protein families is with experimental confirmation. In this review, we reassess the substantially less. This trend is also observed among meaning of structural novelty and examine its relevance domain families, which are 10-fold fewer (<10 000) than to the complexity of the structure-function paradigm. the number of protein families. By targeting these, the Recent advances in the prediction of protein function from structural genomics initiatives can aim to characterise the structure are discussed, as well as new sequence-based major building blocks of whole proteins and since these methods for partitioning large, diverse superfamilies into domains recur in different combination in the genomes, it biologically meaningful clusters. Obtaining structural data for will be an important step towards understanding the these functionally coherent groups of proteins will allow us to complete structural repertoire in nature. Furthermore, better understand the relationship between structure and the complement of molecular functions found within function. an organism is likely to be even fewer. For example, 97% of proteins in yeast can be assigned one or more of Address 4000 unique GO terms. -

A Molecular Phylogenetic Toolkit Using Node.Js 1 2 3 Damien M
bioRxiv preprint doi: https://doi.org/10.1101/075101; this version posted October 9, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Phylo-Node: a molecular phylogenetic toolkit using Node.js 2 3 4 Damien M. O’Halloran1,2* 5 6 1. Department of Biological Sciences, The George Washington University, Science 7 and Engineering Hall, Rm 6000, 800 22nd Street N.W., Washington DC 20052, USA. 8 2. Institute for Neuroscience, The George Washington University, 636 Ross Hall, 9 2300 I St. N.W. Washington DC 20052, USA. 10 11 12 13 *Corresponding author address: 14 Damien O’Halloran, The George Washington University, 636 Ross Hall, 2300 I St. 15 N.W. Washington DC 20052, USA. 16 Tel: 202-994-8955 17 Fax: 202-994-6100 18 EMAIL: [email protected] 19 20 21 22 23 24 25 1 bioRxiv preprint doi: https://doi.org/10.1101/075101; this version posted October 9, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 26 ABSTRACT 27 Background: Node.js is an open-source and cross-platform environment that 28 provides a JavaScript codebase for back-end server-side applications. JavaScript 29 has been used to develop very fast, and user-friendly front-end tools for bioinformatic 30 and phylogenetic analyses. However, no such toolkits are available using Node.js to 31 conduct comprehensive molecular phylogenetic analysis. -

TRINITY COLLEGE Cambridge Trinity College Cambridge College Trinity Annual Record Annual
2016 TRINITY COLLEGE cambridge trinity college cambridge annual record annual record 2016 Trinity College Cambridge Annual Record 2015–2016 Trinity College Cambridge CB2 1TQ Telephone: 01223 338400 e-mail: [email protected] website: www.trin.cam.ac.uk Contents 5 Editorial 11 Commemoration 12 Chapel Address 15 The Health of the College 18 The Master’s Response on Behalf of the College 25 Alumni Relations & Development 26 Alumni Relations and Associations 37 Dining Privileges 38 Annual Gatherings 39 Alumni Achievements CONTENTS 44 Donations to the College Library 47 College Activities 48 First & Third Trinity Boat Club 53 Field Clubs 71 Students’ Union and Societies 80 College Choir 83 Features 84 Hermes 86 Inside a Pirate’s Cookbook 93 “… Through a Glass Darkly…” 102 Robert Smith, John Harrison, and a College Clock 109 ‘We need to talk about Erskine’ 117 My time as advisor to the BBC’s War and Peace TRINITY ANNUAL RECORD 2016 | 3 123 Fellows, Staff, and Students 124 The Master and Fellows 139 Appointments and Distinctions 141 In Memoriam 155 A Ninetieth Birthday Speech 158 An Eightieth Birthday Speech 167 College Notes 181 The Register 182 In Memoriam 186 Addresses wanted CONTENTS TRINITY ANNUAL RECORD 2016 | 4 Editorial It is with some trepidation that I step into Boyd Hilton’s shoes and take on the editorship of this journal. He managed the transition to ‘glossy’ with flair and panache. As historian of the College and sometime holder of many of its working offices, he also brought a knowledge of its past and an understanding of its mysteries that I am unable to match. -

Multi-Class Protein Classification Using Adaptive Codes
Journal of Machine Learning Research 8 (2007) 1557-1581 Submitted 8/06; Revised 4/07; Published 7/07 Multi-class Protein Classification Using Adaptive Codes Iain Melvin∗ [email protected] NEC Laboratories of America Princeton, NJ 08540, USA Eugene Ie∗ [email protected] Department of Computer Science and Engineering University of California San Diego, CA 92093-0404, USA Jason Weston [email protected] NEC Laboratories of America Princeton, NJ 08540, USA William Stafford Noble [email protected] Department of Genome Sciences Department of Computer Science and Engineering University of Washington Seattle, WA 98195, USA Christina Leslie [email protected] Center for Computational Learning Systems Columbia University New York, NY 10115, USA Editor: Nello Cristianini Abstract Predicting a protein’s structural class from its amino acid sequence is a fundamental problem in computational biology. Recent machine learning work in this domain has focused on develop- ing new input space representations for protein sequences, that is, string kernels, some of which give state-of-the-art performance for the binary prediction task of discriminating between one class and all the others. However, the underlying protein classification problem is in fact a huge multi- class problem, with over 1000 protein folds and even more structural subcategories organized into a hierarchy. To handle this challenging many-class problem while taking advantage of progress on the binary problem, we introduce an adaptive code approach in the output space of one-vs- the-rest prediction scores. Specifically, we use a ranking perceptron algorithm to learn a weight- ing of binary classifiers that improves multi-class prediction with respect to a fixed set of out- put codes. -

Annual Scientific Report 2013 on the Cover Structure 3Fof in the Protein Data Bank, Determined by Laponogov, I
EMBL-European Bioinformatics Institute Annual Scientific Report 2013 On the cover Structure 3fof in the Protein Data Bank, determined by Laponogov, I. et al. (2009) Structural insight into the quinolone-DNA cleavage complex of type IIA topoisomerases. Nature Structural & Molecular Biology 16, 667-669. © 2014 European Molecular Biology Laboratory This publication was produced by the External Relations team at the European Bioinformatics Institute (EMBL-EBI) A digital version of the brochure can be found at www.ebi.ac.uk/about/brochures For more information about EMBL-EBI please contact: [email protected] Contents Introduction & overview 3 Services 8 Genes, genomes and variation 8 Molecular atlas 12 Proteins and protein families 14 Molecular and cellular structures 18 Chemical biology 20 Molecular systems 22 Cross-domain tools and resources 24 Research 26 Support 32 ELIXIR 36 Facts and figures 38 Funding & resource allocation 38 Growth of core resources 40 Collaborations 42 Our staff in 2013 44 Scientific advisory committees 46 Major database collaborations 50 Publications 52 Organisation of EMBL-EBI leadership 61 2013 EMBL-EBI Annual Scientific Report 1 Foreword Welcome to EMBL-EBI’s 2013 Annual Scientific Report. Here we look back on our major achievements during the year, reflecting on the delivery of our world-class services, research, training, industry collaboration and European coordination of life-science data. The past year has been one full of exciting changes, both scientifically and organisationally. We unveiled a new website that helps users explore our resources more seamlessly, saw the publication of ground-breaking work in data storage and synthetic biology, joined the global alliance for global health, built important new relationships with our partners in industry and celebrated the launch of ELIXIR. -

Establishing Incentives and Changing Cultures to Support Data Access
EXPERT ADVISORY GROUP ON DATA ACCESS ESTABLISHING INCENTIVES AND CHANGING CULTURES TO SUPPORT DATA ACCESS May 2014 ACKNOWLEDGEMENT This is a report of the Expert Advisory Group on Data Access (EAGDA). EAGDA was established by the MRC, ESRC, Cancer Research UK and the Wellcome Trust in 2012 to provide strategic advice on emerging scientific, ethical and legal issues in relation to data access for cohort and longitudinal studies. The report is based on work undertaken by the EAGDA secretariat at the Group’s request. The contributions of David Carr, Natalie Banner, Grace Gottlieb, Joanna Scott and Katherine Littler at the Wellcome Trust are gratefully acknowledged. EAGDA would also like to thank the representatives of the MRC, ESRC and Cancer Research UK for their support and input throughout the project. Most importantly, EAGDA owes a considerable debt of gratitude to the many individuals from the research community who contributed to this study through feeding in their expert views via surveys, interviews and focus groups. The Expert Advisory Group on Data Access Martin Bobrow (Chair) Bartha Maria Knoppers James Banks Mark McCarthy Paul Burton Andrew Morris George Davey Smith Onora O'Neill Rosalind Eeles Nigel Shadbolt Paul Flicek Chris Skinner Mark Guyer Melanie Wright Tim Hubbard 1 EXECUTIVE SUMMARY This project was developed as a key component of the workplan of the Expert Advisory Group on Data Access (EAGDA). EAGDA wished to understand the factors that help and hinder individual researchers in making their data (both published and unpublished) available to other researchers, and to examine the potential need for new types of incentives to enable data access and sharing. -

EYAL AKIVA, Phd
Eyal Akiva CV, Nov. 2017 EYAL AKIVA, PhD Department of Bioengineering and Therapeutic Sciences Phone +1-650-504-9008 University of California at San Francisco Email [email protected] 1700 4th street, San Francisco, Web www.babbittlab.ucsf.edu/eakiva CA, USA EDUCATION 2012-2017 Post-doctoral fellowship at UCSF, Dept. Of Bioengineering and Therapeutic Sciences. Host: Prof. Patricia Babbitt. 2010-2012 Post-doctoral fellowship at UCSF, Dept. Of Bioengineering and Therapeutic Sciences. Host: Prof. Tanja Kortemme. 2004-2010 PhD at The Hebrew University of Jerusalem (Israel), bioinformatics. Host: Prof. Hanah Margalit. “Various Aspects of Modularity in Protein-Protein Interaction". 2001-2004 MSc at The Hebrew University of Jerusalem (Israel), bioinformatics and human genetics. Host: Prof. Muli Ben-Sasson. “Exploiting the Exploiters: Identification of Virus-Host Pep- tide Mimicry as a Source for Modules of Functional Significance”. MAGNA CUM LAUDE. 1997-2000 BSc at Bar-Ilan University (Israel), biology (major) and computer science (minor). Final project advisor: Prof. Ramit Mehr “Modeling the Evolution of the Immune System: a Sim- ulation of the Evolution of Genes that Encode the Variable Regions of Immunoglobulins”. MAGNA CUM LAUDE. 1996-1997 First year of "Industrial Engineering and Management" studies, Tel-Aviv University, Israel. OTHER WORK EXPERIENCE 2000-01 ‘Do-coop technologies’: Team leader and chemistry/microbiology researcher; development of biological applications and manufacture of proprietary nanoparticles (Or Yehuda, Israel and Tel-Aviv University (Prof. Eshel Ben-Jacob’s lab at the school of physics)). FUNDING, HONORS AND AWARDS 2017 Grant: Co-PI, “Utilizing metagenomic sequences for enzyme function prediction”, Joint Genome Institute (US Department of Energy) (http://jgi.doe.gov/doe-user-facilities-ficus- join-forces-to-tackle-biology-big-data/).