Investigation of the Active Ingredients and Mechanism of Polygonum Cuspidatum in Asthma Based on Network Pharmacology and Experimental Verification
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Essential Genes and Their Role in Autism Spectrum Disorder
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2017 Essential Genes And Their Role In Autism Spectrum Disorder Xiao Ji University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Bioinformatics Commons, and the Genetics Commons Recommended Citation Ji, Xiao, "Essential Genes And Their Role In Autism Spectrum Disorder" (2017). Publicly Accessible Penn Dissertations. 2369. https://repository.upenn.edu/edissertations/2369 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/2369 For more information, please contact [email protected]. Essential Genes And Their Role In Autism Spectrum Disorder Abstract Essential genes (EGs) play central roles in fundamental cellular processes and are required for the survival of an organism. EGs are enriched for human disease genes and are under strong purifying selection. This intolerance to deleterious mutations, commonly observed haploinsufficiency and the importance of EGs in pre- and postnatal development suggests a possible cumulative effect of deleterious variants in EGs on complex neurodevelopmental disorders. Autism spectrum disorder (ASD) is a heterogeneous, highly heritable neurodevelopmental syndrome characterized by impaired social interaction, communication and repetitive behavior. More and more genetic evidence points to a polygenic model of ASD and it is estimated that hundreds of genes contribute to ASD. The central question addressed in this dissertation is whether genes with a strong effect on survival and fitness (i.e. EGs) play a specific oler in ASD risk. I compiled a comprehensive catalog of 3,915 mammalian EGs by combining human orthologs of lethal genes in knockout mice and genes responsible for cell-based essentiality. -

A Tool for Clinical Management of Genetic Variants
Wang et al. Genome Medicine (2015) 7:77 DOI 10.1186/s13073-015-0207-6 SOFTWARE Open Access ClinLabGeneticist: a tool for clinical management of genetic variants from whole exome sequencing in clinical genetic laboratories Jinlian Wang, Jun Liao, Jinglan Zhang, Wei-Yi Cheng, Jörg Hakenberg, Meng Ma, Bryn D. Webb, Rajasekar Ramasamudram-chakravarthi, Lisa Karger, Lakshmi Mehta, Ruth Kornreich, George A. Diaz, Shuyu Li, Lisa Edelmann* and Rong Chen* Abstract Routine clinical application of whole exome sequencing remains challenging due to difficulties in variant interpretation, large dataset management, and workflow integration. We describe a tool named ClinLabGeneticist to implement a workflow in clinical laboratories for management of variant assessment in genetic testing and disease diagnosis. We established an extensive variant annotation data source for the identification of pathogenic variants. A dashboard was deployed to aid a multi-step, hierarchical review process leading to final clinical decisions on genetic variant assessment. In addition, a central database was built to archive all of the genetic testing data, notes, and comments throughout the review process, variant validation data by Sanger sequencing as well as the final clinical reports for future reference. The entire workflow including data entry, distribution of work assignments, variant evaluation and review, selection of variants for validation, report generation, and communications between various personnel is integrated into a single data management platform. Three case studies are presented to illustrate the utility of ClinLabGeneticist. ClinLabGeneticist is freely available to academia at http://rongchenlab.org/software/clinlabgeneticist. Background effects, and testing of tumor biopsies to determine somatic Molecular genetic testing is playing an increasingly im- alterations for cancer classification, prognosis, and devel- portant role in medicine. -

Nº Ref Uniprot Proteína Péptidos Identificados Por MS/MS 1 P01024
Document downloaded from http://www.elsevier.es, day 26/09/2021. This copy is for personal use. Any transmission of this document by any media or format is strictly prohibited. Nº Ref Uniprot Proteína Péptidos identificados 1 P01024 CO3_HUMAN Complement C3 OS=Homo sapiens GN=C3 PE=1 SV=2 por 162MS/MS 2 P02751 FINC_HUMAN Fibronectin OS=Homo sapiens GN=FN1 PE=1 SV=4 131 3 P01023 A2MG_HUMAN Alpha-2-macroglobulin OS=Homo sapiens GN=A2M PE=1 SV=3 128 4 P0C0L4 CO4A_HUMAN Complement C4-A OS=Homo sapiens GN=C4A PE=1 SV=1 95 5 P04275 VWF_HUMAN von Willebrand factor OS=Homo sapiens GN=VWF PE=1 SV=4 81 6 P02675 FIBB_HUMAN Fibrinogen beta chain OS=Homo sapiens GN=FGB PE=1 SV=2 78 7 P01031 CO5_HUMAN Complement C5 OS=Homo sapiens GN=C5 PE=1 SV=4 66 8 P02768 ALBU_HUMAN Serum albumin OS=Homo sapiens GN=ALB PE=1 SV=2 66 9 P00450 CERU_HUMAN Ceruloplasmin OS=Homo sapiens GN=CP PE=1 SV=1 64 10 P02671 FIBA_HUMAN Fibrinogen alpha chain OS=Homo sapiens GN=FGA PE=1 SV=2 58 11 P08603 CFAH_HUMAN Complement factor H OS=Homo sapiens GN=CFH PE=1 SV=4 56 12 P02787 TRFE_HUMAN Serotransferrin OS=Homo sapiens GN=TF PE=1 SV=3 54 13 P00747 PLMN_HUMAN Plasminogen OS=Homo sapiens GN=PLG PE=1 SV=2 48 14 P02679 FIBG_HUMAN Fibrinogen gamma chain OS=Homo sapiens GN=FGG PE=1 SV=3 47 15 P01871 IGHM_HUMAN Ig mu chain C region OS=Homo sapiens GN=IGHM PE=1 SV=3 41 16 P04003 C4BPA_HUMAN C4b-binding protein alpha chain OS=Homo sapiens GN=C4BPA PE=1 SV=2 37 17 Q9Y6R7 FCGBP_HUMAN IgGFc-binding protein OS=Homo sapiens GN=FCGBP PE=1 SV=3 30 18 O43866 CD5L_HUMAN CD5 antigen-like OS=Homo -

Epigenome-450K-Wide Methylation Signatures of Active Cigarette Smoking: the Young Finns Study
Bioscience Reports (2020) 40 BSR20200596 https://doi.org/10.1042/BSR20200596 Research Article Epigenome-450K-wide methylation signatures of active cigarette smoking: The Young Finns Study Pashupati P. Mishra1,2,3,*, Ismo Hanninen¨ 1,2,3,*, Emma Raitoharju1,2,3, Saara Marttila1,2,3,4, Binisha H. Mishra1,2,3, Nina Mononen1,2,3,MikaKah¨ onen¨ 2,5, Mikko Hurme4,6, Olli Raitakari7,8,9, Petri Tor¨ onen¨ 10, Liisa Holm10,11 and Terho Lehtimaki¨ 1,2,3 Downloaded from http://portlandpress.com/bioscirep/article-pdf/40/7/BSR20200596/887717/bsr-2020-0596.pdf by guest on 27 September 2021 1Department of Clinical Chemistry, Faculty of Medicine and Health Technology, Tampere University, Tampere, Finland; 2Finnish Cardiovascular Research Center-Tampere, Faculty of Medicine and Health Technology, Tampere University, Tampere, Finland; 3Department of Clinical Chemistry, Fimlab Laboratories, Tampere, Finland; 4Gerontology Research Center (GEREC), Tampere University, Tampere, Finland; 5Department of Clinical Physiology, Tampere University Hospital, Tampere, Finland; 6Department of Microbiology and Immunology, Faculty of Medicine and Health Technology, Tampere University, Tampere, Finland; 7Centre for Population Health Research, University of Turku and Turku University Hospital, Turku, Finland; 8Research Centre of Applied and Preventive Cardiovascular Medicine, University of Turku, Turku, Finland; 9Department of Clinical Physiology and Nuclear Medicine, Turku University Hospital, Turku, Finland; 10Institute of Biotechnology, Helsinki Institute of Life Sciences (HiLife), University of Helsinki, Helsinki, Finland; 11Organismal and Evolutionary Biology Research Program, Faculty of Biological and Environmental Sciences, University of Helsinki, Helsinki, Finland Correspondence: Pashupati P. Mishra (pashupati.mishra@tuni.fi) Smoking as a major risk factor for morbidity affects numerous regulatory systems of the human body including DNA methylation. -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

Comprehensive Gene Expression Analysis of Prostate Cancer Reveals Distinct Transcriptional Programs Associated with Metastatic Disease1
[CANCER RESEARCH 62, 4499–4506, August 1, 2002] Comprehensive Gene Expression Analysis of Prostate Cancer Reveals Distinct Transcriptional Programs Associated with Metastatic Disease1 Eva LaTulippe, Jaya Satagopan, Alex Smith, Howard Scher, Peter Scardino, Victor Reuter, and William L. Gerald2 Departments of Pathology [E. L., V. R., W. L. G.], Epidemiology and Biostatistics [J. S., A. S.], Urology [P. S.], and Genitourinary Oncology Service, Department of Medicine [H. S.], Memorial Sloan-Kettering Cancer Center, New York, New York 10021, and Department of Medicine, Joan and Sanford I. Weill College of Medicine of Cornell University, New York, New York 10021 [H. S.] ABSTRACT tissues. To form clinically significant tumors, metastatic cells must proliferate in the new microenvironment and recruit a blood supply. The identification of genes that contribute to the biological basis for Those tumor cells growing at metastatic sites are then continually clinical heterogeneity and progression of prostate cancer is critical to accurate classification and appropriate therapy. We performed a compre- selected for growth advantage. This is a complex and dynamic process hensive gene expression analysis of prostate cancer using oligonucleotide that is expected to involve alterations in many genes and transcrip- arrays with 63,175 probe sets to identify genes and expressed sequences tional programs. Identification of genes, gene expression profiles, and with strong and uniform differential expression between nonrecurrent biological pathways that contribute to metastasis will be of significant primary prostate cancers and metastatic prostate cancers. The mean benefit to improved tumor classification and therapy. To address this expression value for >3,000 tumor-intrinsic genes differed by at least question, we performed a genome-wide expression analysis and iden- 3-fold between the two groups. -

Genome-Wide Association Study Identifies Candidate Genes
animals Article Genome-Wide Association Study Identifies Candidate Genes Associated with Feet and Leg Conformation Traits in Chinese Holstein Cattle Ismail Mohamed Abdalla 1, Xubin Lu 1 , Mudasir Nazar 1, Abdelaziz Adam Idriss Arbab 1,2, Tianle Xu 3 , Mohammed Husien Yousif 4, Yongjiang Mao 1 and Zhangping Yang 1,* 1 College of Animal Science and Technology, Yangzhou University, Yangzhou 225009, China; [email protected] (I.M.A.); [email protected] (X.L.); [email protected] (M.N.); [email protected] (A.A.I.A.); [email protected] (Y.M.) 2 Biomedical Research Institute, Darfur College, Nyala 63313, Sudan 3 Joint International Research Laboratory of Agriculture and Agri-Product Safety, Yangzhou University, Yangzhou 225009, China; [email protected] 4 Faculty of Animal Production, West Kordufan University, Alnuhud City 12942, Sudan; [email protected] * Correspondence: [email protected]; Tel.: +86-0514-87979269 Simple Summary: Feet and leg problems are among the major reasons for dairy cows leaving the herd, as well as having direct association with production and reproduction efficiency, health (e.g., claw disorders and lameness) and welfare. Hence, understanding the genetic architecture underlying feet and conformation traits in dairy cattle offers new opportunities toward the genetic improvement and long-term selection. Through a genome-wide association study on Chinese Holstein cattle, we identified several candidate genes associated with feet and leg conformation traits. These results could provide useful information about the molecular breeding basis of feet and leg traits, thus Citation: Abdalla, I.M.; Lu, X.; Nazar, improving the longevity and productivity of dairy cattle. -
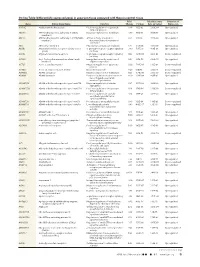
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

Number 11 November 2020 Atlas of Genetics and Cytogenetics in Oncology and Haematology
Volume 1 - Number 1 May - September 1997 Volume 24 - Number 11 November 2020 Atlas of Genetics and Cytogenetics in Oncology and Haematology OPEN ACCESS JOURNAL INIST-CNRS Scope The Atlas of Genetics and Cytogenetics in Oncology and Haematology is a peer reviewed on-line journal in open access, devoted to genes, cytogenetics, and clinical entities in cancer, and cancer-prone diseases. It is made for and by: clinicians and researchers in cytogenetics, molecular biology, oncology, haematology, and pathology. One main scope of the Atlas is to conjugate the scientific information provided by cytogenetics/molecular genetics to the clinical setting (diagnostics, prognostics and therapeutic design), another is to provide an encyclopedic knowledge in cancer genetics. The Atlas deals with cancer research and genomics. It is at the crossroads of research, virtual medical university (university and post-university e-learning), and telemedicine. It contributes to "meta-medicine", this mediation, using information technology, between the increasing amount of knowledge and the individual, having to use the information. Towards a personalized medicine of cancer. It presents structured review articles ("cards") on: 1- Genes, 2- Leukemias, 3- Solid tumors, 4- Cancer-prone diseases, and also 5- "Deep insights": more traditional review articles on the above subjects and on surrounding topics. It also present 6- Case reports in hematology and 7- Educational items in the various related topics for students in Medicine and in Sciences. The Atlas of Genetics -

Genetic Variations in Olfactory Receptor Gene OR2AG2 in a Large
www.nature.com/scientificreports OPEN Genetic variations in olfactory receptor gene OR2AG2 in a large multigenerational family with asthma Samarpana Chakraborty1,2, Pushkar Dakle1, Anirban Sinha 1, Sangeetha Vishweswaraiah3, Aditya Nagori1,2, Shivalingaswamy Salimath4, Y. S. Prakash5, R. Lodha6, S. K. Kabra6, Balaram Ghosh1,2, Mohammed Faruq1,2, P. A. Mahesh4 & Anurag Agrawal1,2* It is estimated from twin studies that heritable factors account for at-least half of asthma-risk, of which genetic variants identifed through population studies explain only a small fraction. Multi-generation large families with high asthma prevalence can serve as a model to identify highly penetrant genetic variants in closely related individuals that are missed by population studies. To achieve this, a four- generation Indian family with asthma was identifed and recruited for examination and genetic testing. Twenty subjects representing all generations were selected for whole genome genotyping, of which eight were subjected to exome sequencing. Non-synonymous and deleterious variants, segregating with the afected individuals, were identifed by exome sequencing. A prioritized deleterious missense common variant in the olfactory receptor gene OR2AG2 that segregated with a risk haplotype in asthma, was validated in an asthma cohort of diferent ethnicity. Phenotypic tests were conducted to verify expected defcits in terms of reduced ability to sense odors. Pathway-level relevance to asthma biology was tested in model systems and unrelated human lung samples. Our study suggests that OR2AG2 and other olfactory receptors may contribute to asthma pathophysiology. Genetic studies on large families of interest can lead to efcient discovery. Asthma is a chronic disorder of the airways caused by a strong genetic predisposition, in addition to environmen- tal insults1. -

Downloaded from the Gene Expression Omnibus to Uncover Diferentially Expressed Long Non-Coding Rnas (Lncrnas), Mrnas, and Micrornas (Mirnas)
Shi and Yao BMC Med Genomics (2021) 14:133 https://doi.org/10.1186/s12920-021-00931-0 RESEARCH ARTICLE Open Access Signature RNAS and related regulatory roles in type 1 diabetes mellitus based on competing endogenous RNA regulatory network analysis Qinghong Shi1 and Hanxin Yao2* Abstract Background: Our study aimed to investigate signature RNAs and their potential roles in type 1 diabetes mellitus (T1DM) using a competing endogenous RNA regulatory network analysis. Methods: Expression profles of GSE55100, deposited from peripheral blood mononuclear cells of 12 T1DM patients and 10 normal controls, were downloaded from the Gene Expression Omnibus to uncover diferentially expressed long non-coding RNAs (lncRNAs), mRNAs, and microRNAs (miRNAs). The ceRNA regulatory network was constructed, then functional and pathway enrichment analysis was conducted. AT1DM-related ceRNA regulatory network was established based on the Human microRNA Disease Database to carry out pathway enrichment analysis. Meanwhile, the T1DM-related pathways were retrieved from the Comparative Toxicogenomics Database (CTD). Results: In total, 847 mRNAs, 41 lncRNAs, and 38 miRNAs were signifcantly diferentially expressed. The ceRNA regu- latory network consisted of 12 lncRNAs, 10 miRNAs, and 24 mRNAs. Two miRNAs (hsa-miR-181a and hsa-miR-1275) were screened as T1DM-related miRNAs to build the T1DM-related ceRNA regulatory network, in which genes were considerably enriched in seven pathways. Moreover, three overlapping pathways, including the phosphatidylinosi- tol signaling system (involving PIP4K2A, INPP4A, PIP4K2C, and CALM1); dopaminergic synapse (involving CALM1 and PPP2R5C); and the insulin signaling pathway (involving CBLB and CALM1) were revealed by comparing with T1DM- related pathways in the CTD, which involved four lncRNAs (LINC01278, TRG-AS1, MIAT, and GAS5-AS1). -

Chimeric Transcript Discovery by Paired-End Transcriptome Sequencing
Chimeric transcript discovery by paired-end transcriptome sequencing Christopher A. Mahera,b, Nallasivam Palanisamya,b, John C. Brennera,b, Xuhong Caoa,c, Shanker Kalyana-Sundarama,b, Shujun Luod, Irina Khrebtukovad, Terrence R. Barrettea,b, Catherine Grassoa,b, Jindan Yua,b, Robert J. Lonigroa,b, Gary Schrothd, Chandan Kumar-Sinhaa,b, and Arul M. Chinnaiyana,b,c,e,f,1 aMichigan Center for Translational Pathology, Ann Arbor, MI 48109; Departments of bPathology and eUrology, University of Michigan, Ann Arbor, MI 48109; cHoward Hughes Medical Institute and fComprehensive Cancer Center, University of Michigan Medical School, Ann Arbor, MI 48109; and dIllumina Inc., 25861 Industrial Boulevard, Hayward, CA 94545 Communicated by David Ginsburg, University of Michigan Medical School, Ann Arbor, MI, May 4, 2009 (received for review March 16, 2009) Recurrent gene fusions are a prevalent class of mutations arising from (SAGE)-like sequencing (13), and next-generation DNA sequenc- the juxtaposition of 2 distinct regions, which can generate novel ing (14). Despite unveiling many novel genomic rearrangements, functional transcripts that could serve as valuable therapeutic targets solid tumors accumulate multiple nonspecific aberrations through- in cancer. Therefore, we aim to establish a sensitive, high-throughput out tumor progression; thus, making causal and driver aberrations methodology to comprehensively catalog functional gene fusions in indistinguishable from secondary and insignificant mutations, cancer by evaluating a paired-end transcriptome sequencing strategy. respectively. Not only did a paired-end approach provide a greater dynamic range The deep unbiased view of a cancer cell enabled by massively in comparison with single read based approaches, but it clearly parallel transcriptome sequencing has greatly facilitated gene fu- distinguished the high-level ‘‘driving’’ gene fusions, such as BCR-ABL1 sion discovery.