Single-Cell Transcriptomics Identifies an Effectorness Gradient
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

HCST (NM 001007469) Human Recombinant Protein Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TP319253 HCST (NM_001007469) Human Recombinant Protein Product data: Product Type: Recombinant Proteins Description: Recombinant protein of human hematopoietic cell signal transducer (HCST), transcript variant 2 Species: Human Expression Host: HEK293T Tag: C-Myc/DDK Predicted MW: 7.3 kDa Concentration: >50 ug/mL as determined by microplate BCA method Purity: > 80% as determined by SDS-PAGE and Coomassie blue staining Buffer: 25 mM Tris.HCl, pH 7.3, 100 mM glycine, 10% glycerol Preparation: Recombinant protein was captured through anti-DDK affinity column followed by conventional chromatography steps. Storage: Store at -80°C. Stability: Stable for 12 months from the date of receipt of the product under proper storage and handling conditions. Avoid repeated freeze-thaw cycles. RefSeq: NP_001007470 Locus ID: 10870 UniProt ID: Q9UBK5 RefSeq Size: 521 Cytogenetics: 19q13.12 RefSeq ORF: 279 Synonyms: DAP10; KAP10; PIK3AP This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 HCST (NM_001007469) Human Recombinant Protein – TP319253 Summary: This gene encodes a transmembrane signaling adaptor that contains a YxxM motif in its cytoplasmic domain. The encoded protein may form part of the immune recognition receptor complex with the C-type lectin-like receptor NKG2D. As part of this receptor complex, this protein may activate phosphatidylinositol 3-kinase dependent signaling pathways through its intracytoplasmic YxxM motif. -
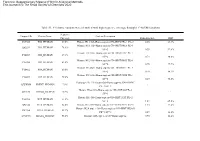
Table S1. 49 Histone Variants Were Identified with High Sequence Coverage Through LC-MS/MS Analysis Electronic Supplementary
Electronic Supplementary Material (ESI) for Analytical Methods. This journal is © The Royal Society of Chemistry 2020 Table S1. 49 histone variants were identified with high sequence coverage through LC-MS/MS analysis Sequence Uniprot IDs Protein Name Protein Description Coverage Ratio E2+/E2- RSD P07305 H10_HUMAN 67.5% Histone H1.0 OS=Homo sapiens GN=H1F0 PE=1 SV=3 4.85 23.3% Histone H1.1 OS=Homo sapiens GN=HIST1H1A PE=1 Q02539 H11_HUMAN 74.4% SV=3 0.35 92.6% Histone H1.2 OS=Homo sapiens GN=HIST1H1C PE=1 P16403 H12_HUMAN 67.1% SV=2 0.73 80.6% Histone H1.3 OS=Homo sapiens GN=HIST1H1D PE=1 P16402 H13_HUMAN 63.8% SV=2 0.75 77.7% Histone H1.4 OS=Homo sapiens GN=HIST1H1E PE=1 P10412 H14_HUMAN 69.0% SV=2 0.70 80.3% Histone H1.5 OS=Homo sapiens GN=HIST1H1B PE=1 P16401 H15_HUMAN 79.6% SV=3 0.29 98.3% Testis-specific H1 histone OS=Homo sapiens GN=H1FNT Q75WM6 H1FNT_HUMAN 7.8% \ \ PE=2 SV=3 Histone H1oo OS=Homo sapiens GN=H1FOO PE=2 Q8IZA3 H1FOO_HUMAN 5.2% \ \ SV=1 Histone H1t OS=Homo sapiens GN=HIST1H1T PE=2 P22492 H1T_HUMAN 31.4% SV=4 1.42 65.0% Q92522 H1X_HUMAN 82.6% Histone H1x OS=Homo sapiens GN=H1FX PE=1 SV=1 1.15 33.2% Histone H2A type 1 OS=Homo sapiens GN=HIST1H2AG P0C0S8 H2A1_HUMAN 99.2% PE=1 SV=2 0.57 26.8% Q96QV6 H2A1A_HUMAN 58.0% Histone H2A type 1-A OS=Homo sapiens 0.90 11.2% GN=HIST1H2AA PE=1 SV=3 Histone H2A type 1-B/E OS=Homo sapiens P04908 H2A1B_HUMAN 99.2% GN=HIST1H2AB PE=1 SV=2 0.92 30.2% Histone H2A type 1-C OS=Homo sapiens Q93077 H2A1C_HUMAN 100.0% GN=HIST1H2AC PE=1 SV=3 0.76 27.6% Histone H2A type 1-D OS=Homo sapiens P20671 -

UNIVERSITY of CALIFORNIA, SAN DIEGO Functional Analysis of Sall4
UNIVERSITY OF CALIFORNIA, SAN DIEGO Functional analysis of Sall4 in modulating embryonic stem cell fate A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Molecular Pathology by Pei Jen A. Lee Committee in charge: Professor Steven Briggs, Chair Professor Geoff Rosenfeld, Co-Chair Professor Alexander Hoffmann Professor Randall Johnson Professor Mark Mercola 2009 Copyright Pei Jen A. Lee, 2009 All rights reserved. The dissertation of Pei Jen A. Lee is approved, and it is acceptable in quality and form for publication on microfilm and electronically: ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ Co-Chair ______________________________________________________________ Chair University of California, San Diego 2009 iii Dedicated to my parents, my brother ,and my husband for their love and support iv Table of Contents Signature Page……………………………………………………………………….…iii Dedication…...…………………………………………………………………………..iv Table of Contents……………………………………………………………………….v List of Figures…………………………………………………………………………...vi List of Tables………………………………………………….………………………...ix Curriculum vitae…………………………………………………………………………x Acknowledgement………………………………………………….……….……..…...xi Abstract………………………………………………………………..…………….....xiii Chapter 1 Introduction ..…………………………………………………………………………….1 Chapter 2 Materials and Methods……………………………………………………………..…12 -

Genome-Wide Approach to Identify Risk Factors for Therapy-Related Myeloid Leukemia
Leukemia (2006) 20, 239–246 & 2006 Nature Publishing Group All rights reserved 0887-6924/06 $30.00 www.nature.com/leu ORIGINAL ARTICLE Genome-wide approach to identify risk factors for therapy-related myeloid leukemia A Bogni1, C Cheng2, W Liu2, W Yang1, J Pfeffer1, S Mukatira3, D French1, JR Downing4, C-H Pui4,5,6 and MV Relling1,6 1Department of Pharmaceutical Sciences, The University of Tennessee, Memphis, TN, USA; 2Department of Biostatistics, The University of Tennessee, Memphis, TN, USA; 3Hartwell Center, The University of Tennessee, Memphis, TN, USA; 4Department of Pathology, The University of Tennessee, Memphis, TN, USA; 5Department of Hematology/Oncology St Jude Children’s Research Hospital, The University of Tennessee, Memphis, TN, USA; and 6Colleges of Medicine and Pharmacy, The University of Tennessee, Memphis, TN, USA Using a target gene approach, only a few host genetic risk therapy increases, the importance of identifying host factors for factors for treatment-related myeloid leukemia (t-ML) have been secondary neoplasms increases. defined. Gene expression microarrays allow for a more 4 genome-wide approach to assess possible genetic risk factors Because DNA microarrays interrogate multiple ( 10 000) for t-ML. We assessed gene expression profiles (n ¼ 12 625 genes in one experiment, they allow for a ‘genome-wide’ probe sets) in diagnostic acute lymphoblastic leukemic cells assessment of genes that may predispose to leukemogenesis. from 228 children treated on protocols that included leukemo- DNA microarray analysis of gene expression has been used to genic agents such as etoposide, 13 of whom developed t-ML. identify distinct expression profiles that are characteristic of Expression of 68 probes, corresponding to 63 genes, was different leukemia subtypes.13,14 Studies using this method have significantly related to risk of t-ML. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Integrating Single-Step GWAS and Bipartite Networks Reconstruction Provides Novel Insights Into Yearling Weight and Carcass Traits in Hanwoo Beef Cattle
animals Article Integrating Single-Step GWAS and Bipartite Networks Reconstruction Provides Novel Insights into Yearling Weight and Carcass Traits in Hanwoo Beef Cattle Masoumeh Naserkheil 1 , Abolfazl Bahrami 1 , Deukhwan Lee 2,* and Hossein Mehrban 3 1 Department of Animal Science, University College of Agriculture and Natural Resources, University of Tehran, Karaj 77871-31587, Iran; [email protected] (M.N.); [email protected] (A.B.) 2 Department of Animal Life and Environment Sciences, Hankyong National University, Jungang-ro 327, Anseong-si, Gyeonggi-do 17579, Korea 3 Department of Animal Science, Shahrekord University, Shahrekord 88186-34141, Iran; [email protected] * Correspondence: [email protected]; Tel.: +82-31-670-5091 Received: 25 August 2020; Accepted: 6 October 2020; Published: 9 October 2020 Simple Summary: Hanwoo is an indigenous cattle breed in Korea and popular for meat production owing to its rapid growth and high-quality meat. Its yearling weight and carcass traits (backfat thickness, carcass weight, eye muscle area, and marbling score) are economically important for the selection of young and proven bulls. In recent decades, the advent of high throughput genotyping technologies has made it possible to perform genome-wide association studies (GWAS) for the detection of genomic regions associated with traits of economic interest in different species. In this study, we conducted a weighted single-step genome-wide association study which combines all genotypes, phenotypes and pedigree data in one step (ssGBLUP). It allows for the use of all SNPs simultaneously along with all phenotypes from genotyped and ungenotyped animals. Our results revealed 33 relevant genomic regions related to the traits of interest. -

SARS-Cov-2 Entry Protein TMPRSS2 and Its Homologue, TMPRSS4
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.26.441280; this version posted April 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 SARS-CoV-2 Entry Protein TMPRSS2 and Its 2 Homologue, TMPRSS4 Adopts Structural Fold Similar 3 to Blood Coagulation and Complement Pathway 4 Related Proteins ∗,a ∗∗,b b 5 Vijaykumar Yogesh Muley , Amit Singh , Karl Gruber , Alfredo ∗,a 6 Varela-Echavarría a 7 Instituto de Neurobiología, Universidad Nacional Autónoma de México, Querétaro, México b 8 Institute of Molecular Biosciences, University of Graz, Graz, Austria 9 Abstract The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) utilizes TMPRSS2 receptor to enter target human cells and subsequently causes coron- avirus disease 19 (COVID-19). TMPRSS2 belongs to the type II serine proteases of subfamily TMPRSS, which is characterized by the presence of the serine- protease domain. TMPRSS4 is another TMPRSS member, which has a domain architecture similar to TMPRSS2. TMPRSS2 and TMPRSS4 have been shown to be involved in SARS-CoV-2 infection. However, their normal physiological roles have not been explored in detail. In this study, we analyzed the amino acid sequences and predicted 3D structures of TMPRSS2 and TMPRSS4 to under- stand their functional aspects at the protein domain level. Our results suggest that these proteins are likely to have common functions based on their conserved domain organization. -

CD29 Identifies IFN-Γ–Producing Human CD8+ T Cells With
+ CD29 identifies IFN-γ–producing human CD8 T cells with an increased cytotoxic potential Benoît P. Nicoleta,b, Aurélie Guislaina,b, Floris P. J. van Alphenc, Raquel Gomez-Eerlandd, Ton N. M. Schumacherd, Maartje van den Biggelaarc,e, and Monika C. Wolkersa,b,1 aDepartment of Hematopoiesis, Sanquin Research, 1066 CX Amsterdam, The Netherlands; bLandsteiner Laboratory, Oncode Institute, Amsterdam University Medical Center, University of Amsterdam, 1105 AZ Amsterdam, The Netherlands; cDepartment of Research Facilities, Sanquin Research, 1066 CX Amsterdam, The Netherlands; dDivision of Molecular Oncology and Immunology, Oncode Institute, The Netherlands Cancer Institute, 1066 CX Amsterdam, The Netherlands; and eDepartment of Molecular and Cellular Haemostasis, Sanquin Research, 1066 CX Amsterdam, The Netherlands Edited by Anjana Rao, La Jolla Institute for Allergy and Immunology, La Jolla, CA, and approved February 12, 2020 (received for review August 12, 2019) Cytotoxic CD8+ T cells can effectively kill target cells by producing therefore developed a protocol that allowed for efficient iso- cytokines, chemokines, and granzymes. Expression of these effector lation of RNA and protein from fluorescence-activated cell molecules is however highly divergent, and tools that identify and sorting (FACS)-sorted fixed T cells after intracellular cytokine + preselect CD8 T cells with a cytotoxic expression profile are lacking. staining. With this top-down approach, we performed an un- + Human CD8 T cells can be divided into IFN-γ– and IL-2–producing biased RNA-sequencing (RNA-seq) and mass spectrometry cells. Unbiased transcriptomics and proteomics analysis on cytokine- γ– – + + (MS) analyses on IFN- and IL-2 producing primary human producing fixed CD8 T cells revealed that IL-2 cells produce helper + + + CD8 Tcells. -

Análise Integrativa De Perfis Transcricionais De Pacientes Com
UNIVERSIDADE DE SÃO PAULO FACULDADE DE MEDICINA DE RIBEIRÃO PRETO PROGRAMA DE PÓS-GRADUAÇÃO EM GENÉTICA ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Ribeirão Preto – 2012 ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Tese apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo para obtenção do título de Doutor em Ciências. Área de Concentração: Genética Orientador: Prof. Dr. Eduardo Antonio Donadi Co-orientador: Prof. Dr. Geraldo A. S. Passos Ribeirão Preto – 2012 AUTORIZO A REPRODUÇÃO E DIVULGAÇÃO TOTAL OU PARCIAL DESTE TRABALHO, POR QUALQUER MEIO CONVENCIONAL OU ELETRÔNICO, PARA FINS DE ESTUDO E PESQUISA, DESDE QUE CITADA A FONTE. FICHA CATALOGRÁFICA Evangelista, Adriane Feijó Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. Ribeirão Preto, 2012 192p. Tese de Doutorado apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo. Área de Concentração: Genética. Orientador: Donadi, Eduardo Antonio Co-orientador: Passos, Geraldo A. 1. Expressão gênica – microarrays 2. Análise bioinformática por module maps 3. Diabetes mellitus tipo 1 4. Diabetes mellitus tipo 2 5. Diabetes mellitus gestacional FOLHA DE APROVAÇÃO ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. -

Genetic Correlations Reveal the Shared Genetic Architecture of Transcription in Human Peripheral Blood
Hemani, G. (2017). Genetic correlations reveal the shared genetic architecture of transcription in human peripheral blood. Nature Communications, 8, [483]. https://doi.org/10.1038/s41467-017-00473- z Publisher's PDF, also known as Version of record License (if available): CC BY Link to published version (if available): 10.1038/s41467-017-00473-z Link to publication record in Explore Bristol Research PDF-document This is the final published version of the article (version of record). It first appeared online via Nature Publishing Group at https://www.nature.com/articles/s41467-017-00473-z . Please refer to any applicable terms of use of the publisher. University of Bristol - Explore Bristol Research General rights This document is made available in accordance with publisher policies. Please cite only the published version using the reference above. Full terms of use are available: http://www.bristol.ac.uk/red/research-policy/pure/user-guides/ebr-terms/ ARTICLE DOI: 10.1038/s41467-017-00473-z OPEN Genetic correlations reveal the shared genetic architecture of transcription in human peripheral blood Samuel W. Lukowski 1, Luke R. Lloyd-Jones1,2, Alexander Holloway2, Holger Kirsten3,4, Gibran Hemani 5,6, Jian Yang 1,2, Kerrin Small 7, Jing Zhao8, Andres Metspalu9, Emmanouil T. Dermitzakis10, Greg Gibson 8, Timothy D. Spector7, Joachim Thiery4,11, Markus Scholz 3,4, Grant W. Montgomery1,12, Tonu Esko 9, Peter M. Visscher 1,2 & Joseph E. Powell1,2 Transcript co-expression is regulated by a combination of shared genetic and environmental factors. Here, we estimate the proportion of co-expression that is due to shared genetic variance. -

1A Multiple Sclerosis Treatment
The Pharmacogenomics Journal (2012) 12, 134–146 & 2012 Macmillan Publishers Limited. All rights reserved 1470-269X/12 www.nature.com/tpj ORIGINAL ARTICLE Network analysis of transcriptional regulation in response to intramuscular interferon-b-1a multiple sclerosis treatment M Hecker1,2, RH Goertsches2,3, Interferon-b (IFN-b) is one of the major drugs for multiple sclerosis (MS) 3 2 treatment. The purpose of this study was to characterize the transcriptional C Fatum , D Koczan , effects induced by intramuscular IFN-b-1a therapy in patients with relapsing– 2 1 H-J Thiesen , R Guthke remitting form of MS. By using Affymetrix DNA microarrays, we obtained and UK Zettl3 genome-wide expression profiles of peripheral blood mononuclear cells of 24 MS patients within the first 4 weeks of IFN-b administration. We identified 1Leibniz Institute for Natural Product Research 121 genes that were significantly up- or downregulated compared with and Infection Biology—Hans-Knoell-Institute, baseline, with stronger changed expression at 1 week after start of therapy. Jena, Germany; 2University of Rostock, Institute of Immunology, Rostock, Germany and Eleven transcription factor-binding sites (TFBS) are overrepresented in the 3University of Rostock, Department of Neurology, regulatory regions of these genes, including those of IFN regulatory factors Rostock, Germany and NF-kB. We then applied TFBS-integrating least angle regression, a novel integrative algorithm for deriving gene regulatory networks from gene Correspondence: M Hecker, Leibniz Institute for Natural Product expression data and TFBS information, to reconstruct the underlying network Research and Infection Biology—Hans-Knoell- of molecular interactions. An NF-kB-centered sub-network of genes was Institute, Beutenbergstr. -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine