Downloaded Its Corresponding Raw and Summarized Ephys Data (Summary Measurements Included Input Resistance and Resting Potential)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Exploring the Relationship Between Genetic Variation in Taste Receptor Genes and Salt Taste Perception Among People with Hypertension
Mississippi State University Scholars Junction Theses and Dissertations Theses and Dissertations 11-25-2020 Exploring the relationship between genetic variation in taste receptor genes and salt taste perception among people with hypertension Pradtana Tapanee Follow this and additional works at: https://scholarsjunction.msstate.edu/td Recommended Citation Tapanee, Pradtana, "Exploring the relationship between genetic variation in taste receptor genes and salt taste perception among people with hypertension" (2020). Theses and Dissertations. 2176. https://scholarsjunction.msstate.edu/td/2176 This Dissertation - Open Access is brought to you for free and open access by the Theses and Dissertations at Scholars Junction. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of Scholars Junction. For more information, please contact [email protected]. Template B v4.1 (beta): Created by L. Threet 11/15/19 Exploring the relationship between genetic variation in taste receptor genes and salt taste perception among people with hypertension By TITLE PAGE Pradtana Tapanee Approved by: Terezie Tolar-Peterson (Major Professor) Daniel G. Peterson Diane K. Tidwell M. Wes Schilling Wen-Hsing Cheng (Committee Member/Graduate Coordinator) Scott T. Willard (Dean, College of Agriculture and Life Sciences) A Dissertation Submitted to the Faculty of Mississippi State University in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy in Nutrition in the Department of Food Science, -

Update on the Implication of Potassium Channels in Autism: K+ Channelautism Spectrum Disorder
REVIEW ARTICLE published: 02 March 2015 doi: 10.3389/fncel.2015.00034 Update on the implication of potassium channels in autism: K+ channelautism spectrum disorder Luca Guglielmi 1*, Ilenio Servettini 1, Martino Caramia 2 , Luigi Catacuzzeno 2 , Fabio Franciolini 2 , Maria Cristina D’Adamo1 and Mauro Pessia1* 1 Section of Physiology and Biochemistry, Department of Experimental Medicine, University of Perugia School of Medicine, Perugia, Italy 2 Department of Chemistry, Biology and Biotechnology, University of Perugia, Perugia, Italy Edited by: Autism spectrum disorders (ASDs) are characterized by impaired ability to properly Tycho M. Hoogland, Netherlands implement environmental stimuli that are essential to achieve a state of social and Institute for Neuroscience, Netherlands cultural exchange. Indeed, the main features of ASD are impairments of interpersonal relationships, verbal and non-verbal communication and restricted and repetitive behaviors. Reviewed by: Noritaka Ichinohe, National Institute of These aspects are often accompanied by several comorbidities such as motor delay, Neuroscience – National Center of praxis impairment, gait abnormalities, insomnia, and above all epilepsy. Genetic analyses Neurology and Psychiatry, Japan of autistic individuals uncovered deleterious mutations in several K+ channel types Mohamed Jaber, University of Poitiers, France strengthening the notion that their intrinsic dysfunction may play a central etiologic role in ASD. However, indirect implication of K+ channels in ASD has been also reported. *Correspondence: + Luca Guglielmi and Mauro Pessia, For instance, loss of fragile X mental retardation protein (FMRP) results in K channels Section of Physiology and deregulation, network dysfunction and ASD-like cognitive and behavioral symptoms. This Biochemistry, Department of review provides an update on direct and indirect implications of K+ channels in ASDs. -

The Mineralocorticoid Receptor Leads to Increased Expression of EGFR
www.nature.com/scientificreports OPEN The mineralocorticoid receptor leads to increased expression of EGFR and T‑type calcium channels that support HL‑1 cell hypertrophy Katharina Stroedecke1,2, Sandra Meinel1,2, Fritz Markwardt1, Udo Kloeckner1, Nicole Straetz1, Katja Quarch1, Barbara Schreier1, Michael Kopf1, Michael Gekle1 & Claudia Grossmann1* The EGF receptor (EGFR) has been extensively studied in tumor biology and recently a role in cardiovascular pathophysiology was suggested. The mineralocorticoid receptor (MR) is an important efector of the renin–angiotensin–aldosterone‑system and elicits pathophysiological efects in the cardiovascular system; however, the underlying molecular mechanisms are unclear. Our aim was to investigate the importance of EGFR for MR‑mediated cardiovascular pathophysiology because MR is known to induce EGFR expression. We identifed a SNP within the EGFR promoter that modulates MR‑induced EGFR expression. In RNA‑sequencing and qPCR experiments in heart tissue of EGFR KO and WT mice, changes in EGFR abundance led to diferential expression of cardiac ion channels, especially of the T‑type calcium channel CACNA1H. Accordingly, CACNA1H expression was increased in WT mice after in vivo MR activation by aldosterone but not in respective EGFR KO mice. Aldosterone‑ and EGF‑responsiveness of CACNA1H expression was confrmed in HL‑1 cells by Western blot and by measuring peak current density of T‑type calcium channels. Aldosterone‑induced CACNA1H protein expression could be abrogated by the EGFR inhibitor AG1478. Furthermore, inhibition of T‑type calcium channels with mibefradil or ML218 reduced diameter, volume and BNP levels in HL‑1 cells. In conclusion the MR regulates EGFR and CACNA1H expression, which has an efect on HL‑1 cell diameter, and the extent of this regulation seems to depend on the SNP‑216 (G/T) genotype. -

Pharmacogenomics of Drug-Induced Conditions
Pharmacogenomics of Drug-Induced Conditions Dan M. Roden, M.D. Professor of Medicine and Pharmacology Director, Oates Institute for Experimental Therapeutics Assistant Vice-Chancellor for Personalized Medicine Vanderbilt University School of Medicine !1 An “idiosyncratic” drug response AR, 78 year old male • Coronary artery disease; 13 years s/p bypass surgery • 2 days after starting dofetilide (potent IKr blocker)… No personal or family history of syncope, sudden death • KCNQ1 variant leading to R583C identified • In vitro: ↓IKs • Not found in >400 ethnically-matched controls • ∴this is likely subclinical congenital Long QT Syndrome !2 Another face of the congenital long QT syndrome !3 Father’s ECG !4 Mother’s ECG !5 9-year-old sister’s ECG !6 Family tree Aunt, cousin, niece with 402 426 syncope 462 405 439 428 (RBBB) 436 428 492 411 419 404 • One episode of syncope, age 20 • Multiple ER visits for acute allergic reactions à epinephrine without incident !7 Mother’s ECG post-exercise !8 Identifying a mutation in KCNQ1 mothersister …YI G FLGLI… …YI D FLGLI… G269D !9 Family tree Aunt, cousin, niece with 402 426 syncope 462 405 439 428 (RBBB) 436 428 492 411 419 404 Incomplete penetrance !10 Grandfather’s ECG !11 Defining the “drug-induced long QT syndrome” • Marked QT prolongation and typical pause-dependent torsades de pointes with drug challenge, both resolving with drug withdrawal • Variants: • Marked QT prolongation (e.g. >520 or 550 or 600 msec), recognized and drug withdrawn (no torsades) • pause-dependent polymorphic VT after administration -

Towards Mutation-Specific Precision Medicine in Atypical Clinical
International Journal of Molecular Sciences Review Towards Mutation-Specific Precision Medicine in Atypical Clinical Phenotypes of Inherited Arrhythmia Syndromes Tadashi Nakajima * , Shuntaro Tamura, Masahiko Kurabayashi and Yoshiaki Kaneko Department of Cardiovascular Medicine, Gunma University Graduate School of Medicine, Maebashi 371-8511, Gunma, Japan; [email protected] (S.T.); [email protected] (M.K.); [email protected] (Y.K.) * Correspondence: [email protected]; Tel.: +81-27-220-8145; Fax: +81-27-220-8158 Abstract: Most causal genes for inherited arrhythmia syndromes (IASs) encode cardiac ion channel- related proteins. Genotype-phenotype studies and functional analyses of mutant genes, using heterol- ogous expression systems and animal models, have revealed the pathophysiology of IASs and enabled, in part, the establishment of causal gene-specific precision medicine. Additionally, the utilization of induced pluripotent stem cell (iPSC) technology have provided further insights into the patho- physiology of IASs and novel promising therapeutic strategies, especially in long QT syndrome. It is now known that there are atypical clinical phenotypes of IASs associated with specific mutations that have unique electrophysiological properties, which raises a possibility of mutation-specific precision medicine. In particular, patients with Brugada syndrome harboring an SCN5A R1632C mutation exhibit exercise-induced cardiac events, which may be caused by a marked activity-dependent loss of R1632C-Nav1.5 availability due to a marked delay of recovery from inactivation. This suggests that the use of isoproterenol should be avoided. Conversely, the efficacy of β-blocker needs to be examined. Patients harboring a KCND3 V392I mutation exhibit both cardiac (early repolarization syndrome and Citation: Nakajima, T.; Tamura, S.; paroxysmal atrial fibrillation) and cerebral (epilepsy) phenotypes, which may be associated with a Kurabayashi, M.; Kaneko, Y. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
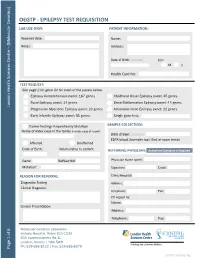
Oegtp - Epilepsy Test Requisition Lab Use Only: Patient Information
OEGTP - EPILEPSY TEST REQUISITION LAB USE ONLY: PATIENT INFORMATION: Received date: Name: Notes: Address: Date of Birth: YY/MM/DD Sex: M F Health Card No: TEST REQUEST: See page 2 for gene list for each of the panels below Epilepsy Comprehensive panel: 167 genes Childhood Onset Epilepsy panel: 45 genes Focal Epilepsy panel: 14 genes Brain Malformation Epilepsy panel: 44 genes London Health Sciences Centre – (Molecular Genetics) London Health Sciences Centre Progressive Myoclonic Epilepsy panel: 20 genes Actionable Gene Epilepsy panel: 22 genes Early Infantile Epilepsy panel: 51 genes Single gene test: Carrier Testing/ KnownFamily Mutation SAMPLE COLLECTION: Name of index case in the family (include copy of report) Date drawn: YY/MM/DD EDTA blood (lavender top) (5ml at room temp) Affected Unaffected Date of Birth: Relationship to patient: REFERRING PHYSICIAN: Authorized Signature is Required Gene: RefSeq:NM Physician Name (print): Mutation: Signature: Email: REASON FOR REFERRAL: Clinic/Hospital: Diagnostic Testing Address: Clinical Diagnosis: Telephone: Fax: CC report to: Name: Clinical Presentation: Address: Telephone: Fax: Molecular Genetics Laboratory Victoria Hospital, Room B10-123A 800 Commissioners Rd. E. London, Ontario | N6A 5W9 Pathology and Laboratory Medicine Ph: 519-685-8122 | Fax: 519-685-8279 Page 1 of 6 Page OEGTP (2021/05/28) OEGTP - EPILEPSY TEST PANELS Patient Identifier: COMPREHENSIVE EPILEPSY PANEL: 167 Genes ACTB, ACTG1, ADSL, AKT3, ALDH7A1, AMT, AP3B2, ARFGEF2, ARHGEF9, ARV1, ARX, ASAH1, ASNS, ATP1A3, ATP6V0A2, ATP7A, -

Transcriptomic Analysis of Native Versus Cultured Human and Mouse Dorsal Root Ganglia Focused on Pharmacological Targets Short
bioRxiv preprint doi: https://doi.org/10.1101/766865; this version posted September 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Transcriptomic analysis of native versus cultured human and mouse dorsal root ganglia focused on pharmacological targets Short title: Comparative transcriptomics of acutely dissected versus cultured DRGs Andi Wangzhou1, Lisa A. McIlvried2, Candler Paige1, Paulino Barragan-Iglesias1, Carolyn A. Guzman1, Gregory Dussor1, Pradipta R. Ray1,#, Robert W. Gereau IV2, # and Theodore J. Price1, # 1The University of Texas at Dallas, School of Behavioral and Brain Sciences and Center for Advanced Pain Studies, 800 W Campbell Rd. Richardson, TX, 75080, USA 2Washington University Pain Center and Department of Anesthesiology, Washington University School of Medicine # corresponding authors [email protected], [email protected] and [email protected] Funding: NIH grants T32DA007261 (LM); NS065926 and NS102161 (TJP); NS106953 and NS042595 (RWG). The authors declare no conflicts of interest Author Contributions Conceived of the Project: PRR, RWG IV and TJP Performed Experiments: AW, LAM, CP, PB-I Supervised Experiments: GD, RWG IV, TJP Analyzed Data: AW, LAM, CP, CAG, PRR Supervised Bioinformatics Analysis: PRR Drew Figures: AW, PRR Wrote and Edited Manuscript: AW, LAM, CP, GD, PRR, RWG IV, TJP All authors approved the final version of the manuscript. 1 bioRxiv preprint doi: https://doi.org/10.1101/766865; this version posted September 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Rescue of Motor Coordination by Purkinje Cell-Targeted Restoration of Kv3.3 Channels in Kcnc3-Null Mice Requires Kcnc1
The Journal of Neuroscience, December 16, 2009 • 29(50):15735–15744 • 15735 Cellular/Molecular Rescue of Motor Coordination by Purkinje Cell-Targeted Restoration of Kv3.3 Channels in Kcnc3-Null Mice Requires Kcnc1 Edward C. Hurlock, Mitali Bose, Ganon Pierce, and Rolf H. Joho Department of Neuroscience, The University of Texas Southwestern Medical Center, Dallas, Texas 75390-9111 The role of cerebellar Kv3.1 and Kv3.3 channels in motor coordination was examined with an emphasis on the deep cerebellar nuclei (DCN). Kv3 channel subunits encoded by Kcnc genes are distinguished by rapid activation and deactivation kinetics that support high-frequency, narrow action potential firing. Previously we reported that increased lateral deviation while ambulating and slips while traversing a narrow beam of ataxic Kcnc3-null mice were corrected by restoration of Kv3.3 channels specifically to Purkinje cells, whereas Kcnc3-mutant mice additionally lacking one Kcnc1 allele were partially rescued. Here, we report mice lacking all Kcnc1 and Kcnc3 alleles exhibit no such rescue. For Purkinje cell output to reach the rest of the brain it must be conveyed by neurons of the DCN or vestibular nuclei. As Kcnc1, but not Kcnc3, alleles are lost, mutant mice exhibit increasing gait ataxia accompanied by spike broadening and deceleration in DCN neurons, suggesting the facet of coordination rescued by Purkinje-cell-restricted Kv3.3 restoration in mice lacking just Kcnc3 is hypermetria, while gait ataxia emerges when additionally Kcnc1 alleles are lost. Thus, fast repolarization in Purkinje cells appears important for normal movement velocity, whereas DCN neurons are a prime candidate locus where fast repolarization is necessary for normal gait patterning. -

Supplemental Table 1. Complete Gene Lists and GO Terms from Figure 3C
Supplemental Table 1. Complete gene lists and GO terms from Figure 3C. Path 1 Genes: RP11-34P13.15, RP4-758J18.10, VWA1, CHD5, AZIN2, FOXO6, RP11-403I13.8, ARHGAP30, RGS4, LRRN2, RASSF5, SERTAD4, GJC2, RHOU, REEP1, FOXI3, SH3RF3, COL4A4, ZDHHC23, FGFR3, PPP2R2C, CTD-2031P19.4, RNF182, GRM4, PRR15, DGKI, CHMP4C, CALB1, SPAG1, KLF4, ENG, RET, GDF10, ADAMTS14, SPOCK2, MBL1P, ADAM8, LRP4-AS1, CARNS1, DGAT2, CRYAB, AP000783.1, OPCML, PLEKHG6, GDF3, EMP1, RASSF9, FAM101A, STON2, GREM1, ACTC1, CORO2B, FURIN, WFIKKN1, BAIAP3, TMC5, HS3ST4, ZFHX3, NLRP1, RASD1, CACNG4, EMILIN2, L3MBTL4, KLHL14, HMSD, RP11-849I19.1, SALL3, GADD45B, KANK3, CTC- 526N19.1, ZNF888, MMP9, BMP7, PIK3IP1, MCHR1, SYTL5, CAMK2N1, PINK1, ID3, PTPRU, MANEAL, MCOLN3, LRRC8C, NTNG1, KCNC4, RP11, 430C7.5, C1orf95, ID2-AS1, ID2, GDF7, KCNG3, RGPD8, PSD4, CCDC74B, BMPR2, KAT2B, LINC00693, ZNF654, FILIP1L, SH3TC1, CPEB2, NPFFR2, TRPC3, RP11-752L20.3, FAM198B, TLL1, CDH9, PDZD2, CHSY3, GALNT10, FOXQ1, ATXN1, ID4, COL11A2, CNR1, GTF2IP4, FZD1, PAX5, RP11-35N6.1, UNC5B, NKX1-2, FAM196A, EBF3, PRRG4, LRP4, SYT7, PLBD1, GRASP, ALX1, HIP1R, LPAR6, SLITRK6, C16orf89, RP11-491F9.1, MMP2, B3GNT9, NXPH3, TNRC6C-AS1, LDLRAD4, NOL4, SMAD7, HCN2, PDE4A, KANK2, SAMD1, EXOC3L2, IL11, EMILIN3, KCNB1, DOK5, EEF1A2, A4GALT, ADGRG2, ELF4, ABCD1 Term Count % PValue Genes regulation of pathway-restricted GDF3, SMAD7, GDF7, BMPR2, GDF10, GREM1, BMP7, LDLRAD4, SMAD protein phosphorylation 9 6.34 1.31E-08 ENG pathway-restricted SMAD protein GDF3, SMAD7, GDF7, BMPR2, GDF10, GREM1, BMP7, LDLRAD4, phosphorylation -

Expression Profiling of Ion Channel Genes Predicts Clinical Outcome in Breast Cancer
UCSF UC San Francisco Previously Published Works Title Expression profiling of ion channel genes predicts clinical outcome in breast cancer Permalink https://escholarship.org/uc/item/1zq9j4nw Journal Molecular Cancer, 12(1) ISSN 1476-4598 Authors Ko, Jae-Hong Ko, Eun A Gu, Wanjun et al. Publication Date 2013-09-22 DOI http://dx.doi.org/10.1186/1476-4598-12-106 Peer reviewed eScholarship.org Powered by the California Digital Library University of California Ko et al. Molecular Cancer 2013, 12:106 http://www.molecular-cancer.com/content/12/1/106 RESEARCH Open Access Expression profiling of ion channel genes predicts clinical outcome in breast cancer Jae-Hong Ko1, Eun A Ko2, Wanjun Gu3, Inja Lim1, Hyoweon Bang1* and Tong Zhou4,5* Abstract Background: Ion channels play a critical role in a wide variety of biological processes, including the development of human cancer. However, the overall impact of ion channels on tumorigenicity in breast cancer remains controversial. Methods: We conduct microarray meta-analysis on 280 ion channel genes. We identify candidate ion channels that are implicated in breast cancer based on gene expression profiling. We test the relationship between the expression of ion channel genes and p53 mutation status, ER status, and histological tumor grade in the discovery cohort. A molecular signature consisting of ion channel genes (IC30) is identified by Spearman’s rank correlation test conducted between tumor grade and gene expression. A risk scoring system is developed based on IC30. We test the prognostic power of IC30 in the discovery and seven validation cohorts by both Cox proportional hazard regression and log-rank test. -

Non-Coding Rnas in the Cardiac Action Potential and Their Impact on Arrhythmogenic Cardiac Diseases
Review Non-Coding RNAs in the Cardiac Action Potential and Their Impact on Arrhythmogenic Cardiac Diseases Estefania Lozano-Velasco 1,2 , Amelia Aranega 1,2 and Diego Franco 1,2,* 1 Cardiovascular Development Group, Department of Experimental Biology, University of Jaén, 23071 Jaén, Spain; [email protected] (E.L.-V.); [email protected] (A.A.) 2 Fundación Medina, 18016 Granada, Spain * Correspondence: [email protected] Abstract: Cardiac arrhythmias are prevalent among humans across all age ranges, affecting millions of people worldwide. While cardiac arrhythmias vary widely in their clinical presentation, they possess shared complex electrophysiologic properties at cellular level that have not been fully studied. Over the last decade, our current understanding of the functional roles of non-coding RNAs have progressively increased. microRNAs represent the most studied type of small ncRNAs and it has been demonstrated that miRNAs play essential roles in multiple biological contexts, including normal development and diseases. In this review, we provide a comprehensive analysis of the functional contribution of non-coding RNAs, primarily microRNAs, to the normal configuration of the cardiac action potential, as well as their association to distinct types of arrhythmogenic cardiac diseases. Keywords: cardiac arrhythmia; microRNAs; lncRNAs; cardiac action potential Citation: Lozano-Velasco, E.; Aranega, A.; Franco, D. Non-Coding RNAs in the Cardiac Action Potential 1. The Electrical Components of the Adult Heart and Their Impact on Arrhythmogenic The adult heart is a four-chambered organ that propels oxygenated blood to the entire Cardiac Diseases. Hearts 2021, 2, body. It is composed of atrial and ventricular chambers, each of them with distinct left and 307–330.