Research Design and Methods
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Pancreatic Beta Cells Express a Diverse Set Ofhomeobox Genes
Proc. Nati. Acad. Sci. USA Vol. 91, pp. 12203-12207, December 1994 Biochemistry Pancreatic beta cells express a diverse set of homeobox genes (Lim motif/Lmx gene/Nkx gene/Alx gene/Vdx homeobox) ABRAHAM RUDNICK*t, THAI YEN LING*, HIROKI ODAGIRI*, WILLIAM J. RUTTER*t, AND MICHAEL S. GERMAN*t§ *Hormone Research Institute and Departments of tMedicine and tBiochemistry and Biophysics, University of California, San Francisco, CA 94143-0534 Contributed by William J. Rutter, August 22, 1994 ABSTRACT Homeobox genes, which are found in all RIPE3B element (16) and the P1 element (8) [also called CT1 eukaryotic organisms, encode transcriptional regulators in- (9)] lie on either side of the IEB1 element. The A/T elements volved in cell-type differentiation and development. Several and the E boxes function synergistically: none of the ele- homeobox genes encoding homeodomain proteins that bind and ments can function in isolation, but combination of an E box activate the insulin gene promoter have been described. In an and an A/T element results in dramatic activation of tran- attempt to identify additional beta-cell homeodomain proteins, scription (11, 16, 19). A number of complexes from beta-cell we designed primers based on the sequences of beta-cell nuclei bind to the A/T elements (6, 8-11, 16, 19). Some homeobox genes cdx3 and lmxl and the Drosophia homeodo- proteins in these complexes have been cloned, and they all main protein Antennapedia and used these primers to amplffy contain homeodomains. The A/T-binding proteins that have inserts by PCR from an insulinoma cDNA library. -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine -

Mouse Alx3: an Aristaless-Like Homeobox Gene Expressed During Embryogenesis in Ectomesenchyme and Lateral Plate Mesoderm
DEVELOPMENTAL BIOLOGY 199, 11–25 (1998) ARTICLE NO. DB988921 View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Mouse Alx3: An aristaless-like Homeobox Gene Expressed during Embryogenesis in Ectomesenchyme and Lateral Plate Mesoderm Derk ten Berge, Antje Brouwer, Sophia El Bahi,* Jean-Louis Gue´net,† Benoıˆt Robert,* and Frits Meijlink Hubrecht Laboratory, Netherlands Institute for Developmental Biology, Uppsalalaan 8, 3584CT Utrecht, The Netherlands; *Institut Pasteur, De´partement de Biologie Mole´culaireGe´ne´tique Mole´culaire du De´veloppement, 28, rue du Dr. Roux, 75724 Paris Cedex 15, France; and †Institut Pasteur, Ge´ne´tique des Mammife`res, De´partement d’ Immunologie, 28, rue du Dr. Roux, 75724 Paris Cedex 15, France Mouse Alx3 is a homeobox gene that is related to the Drosophila aristaless gene and to a group of vertebrate genes including Prx1, Prx2, Cart1, and Alx4. The protein encoded contains a diverged variant of a conserved peptide sequence present near the carboxyl terminus of at least 15 different paired-class-homeodomain proteins. Alx3 is expressed in mouse embryos from 8 days of gestation onward in a characteristic pattern, predominantly in neural crest-derived mesenchyme and in lateral plate mesoderm. We detected prominent expression in frontonasal head mesenchyme and in the first and second pharyngeal arches and some of their derivatives. High expression was also seen in the tail and in many derivatives of the lateral plate mesoderm including the limbs, the body wall, and the genital tubercle. aristaless-related genes like Alx3, Cart1, and Prx2 are expressed in overlapping proximodistal patterns in the pharyngeal arches. -

Dynamic Regulatory Module Networks for Inference of Cell Type
bioRxiv preprint doi: https://doi.org/10.1101/2020.07.18.210328; this version posted July 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Dynamic regulatory module networks for inference of cell type specific transcriptional networks Alireza Fotuhi Siahpirani1,2,+, Deborah Chasman1,8,+, Morten Seirup3,4, Sara Knaack1, Rupa Sridharan1,5, Ron Stewart3, James Thomson3,5,6, and Sushmita Roy1,2,7* 1Wisconsin Institute for Discovery, University of Wisconsin-Madison 2Department of Computer Sciences, University of Wisconsin-Madison 3Morgridge Institute for Research 4Molecular and Environmental Toxicology Program, University of Wisconsin-Madison 5Department of Cell and Regenerative Biology, University of Wisconsin-Madison 6Department of Molecular, Cellular, & Developmental Biology, University of California Santa Barbara 7Department of Biostatistics and Medical Informatics, University of Wisconsin-Madison 8Present address: Division of Reproductive Sciences, Department of Obstetrics and Gynecology, University of Wisconsin-Madison +These authors contributed equally. *To whom correspondence should be addressed. 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.07.18.210328; this version posted July 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Abstract Changes in transcriptional regulatory networks can significantly alter cell fate. To gain insight into transcriptional dynamics, several studies have profiled transcriptomes and epigenomes at different stages of a developmental process. -

Transcriptome Profiling of Cleft Palate Intgf-Beta3 Knockout Mice Alleles: RNA-SEQ Analysis of TGF-Beta3 Mice" (2017)
University of Nebraska Medical Center DigitalCommons@UNMC Theses & Dissertations Graduate Studies Fall 12-15-2017 Transcriptome Profiling of Cleft alateP inTGF-beta3 Knockout Mice Alleles: RNA-SEQ Analysis of TGF-beta3 Mice Kelsey White University of Nebraska Medical Center Follow this and additional works at: https://digitalcommons.unmc.edu/etd Part of the Bioinformatics Commons, and the Developmental Biology Commons Recommended Citation White, Kelsey, "Transcriptome Profiling of Cleft Palate inTGF-beta3 Knockout Mice Alleles: RNA-SEQ Analysis of TGF-beta3 Mice" (2017). Theses & Dissertations. 239. https://digitalcommons.unmc.edu/etd/239 This Thesis is brought to you for free and open access by the Graduate Studies at DigitalCommons@UNMC. It has been accepted for inclusion in Theses & Dissertations by an authorized administrator of DigitalCommons@UNMC. For more information, please contact [email protected]. TRANSCRIPTOME PROFILING OF CLEFT PALATE IN TGF-3 KNOCKOUT MICE ALLELES: RNA-SEQ ANALYSIS OF TGF-3 MICE By Kelsey Marie White, D.D.S. A THESIS Presented to the Faculty of the University of Nebraska Graduate College in Partial Fulfillment of Requirements for the Degree of Master of Science Medical Sciences Interdepartmental Area Graduate Program (Oral Biology) Under the Supervision of Ali Nawshad, Ph.D. University of Nebraska Medical Center Omaha, Nebraska December, 2017 Advisory Committee: Peter Giannini, D.D.S., M.S Ali Nawshad, Ph.D. S. Prem Premaraj, B.D.S., M.S., Ph. D. FRCD(C) Hasan Otu, Ph.D. i ACKNOWLEDGEMENTS First, I would like to sincerely thank my leader and mentor, Dr. Ali Nawshad, for the opportunity to work with and learn from him. -
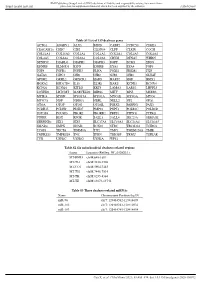
Table S1 List of 159 Deafness Genes ACTG1 ADGRV1 ALX3 BSND
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Med Genet Table S1 List of 159 deafness genes ACTG1 ADGRV1 ALX3 BSND CABP2 CCDC50 CDH23 CEACAM16 CHD7 CIB2 CLDN14 CLPP CLRN1 COCH COL11A1 COL11A2 COL1A1 COL1A2 COL2A1 COL4A3 COL4A4 COL4A5 COL4A6 COL9A1 COL9A2 CRYM DFNA5 WHRN DFNB59 DIABLO DIAPH1 DIAPH3 DSPP ECM1 EDN3 EDNRB ELMOD3 ESPN ESRRB EYA1 EYA4 FGF3 FGF8 FGFR1 FGFR3 FLNA FOXI1 FREM1 FXN GATA3 GIPC3 GJB1 GJB2 GJB3 GJB6 GLYAT GPSM2 GRHL2 GRXCR1 HARS HARS2 HGF HMX1 HOXA2 HSD17B4 IL13 ILDR1 KARS KCNE1 KCNJ10 KCNQ1 KCNQ4 KITLG KRT9 LAMA3 LARS2 LHFPL5 LOXHD1 LRTOMT MARVELD2 MIR96 MITF MPZ MSRB3 MYH14 MYH9 MYO15A MYO1A MYO1E MYO3A MYO6 MYO7A NDP NDRG1 NEFL NELL2 NF2 OPA1 OTOA OTOF OTOG OTOGL P2RX2 PABPN1 PAX3 PCDH15 PCDH9 PDZD7 PMP22 PNPT1 POLR1C POLR1D POU3F4 POU4F3 PROK2 PROKR2 PRPS1 PTPN11 PTPRQ PTPRR RDX RPGR SALL1 SALL4 SEC23A SEMA3E SERPINB6 SIX1 SIX5 SLC17A8 SLC19A2 SLC26A4 SLC26A5 SMAD4 SMPX SNAI2 SOX10 STRC TBC1D24 TCIRG1 TCOF1 TECTA TIMM8A TJP2 TMC1 TMEM126A TMIE TMPRSS3 TMPRSS4 TNC TPRN TRIOBP TRMU TSPEAR TYR USH1C USH1G USH2A WFS1 Table S2 Six mitochondrial deafness-related regions Locus Location (RefSeq: NC_012920.1) MT-RNR1 chrM:648-1601 MT-TL1 chrM:3230-3304 MT-CO1 chrM:5904-7445 MT-TS1 chrM:7446-7514 MT-TK chrM:8295-8364 MT-TE chrM:14674-14742 Table S3 Three deafness-related miRNAs Name Chromosome Position (hg19) miR-96 chr7: 129414532-129414609 miR-182 chr7: 129410223-129410332 miR-183 chr7: 129414745-129414854 Gao X, et al. -

Supporting Information
Supporting Information Li et al. 10.1073/pnas.1617802113 SI Materials and Methods transfection. Cells were transfected with reporter constructs TOP- Plasmids. Mutant Tet3 (H950D and Y952A) was generated by or FOP-Flash using Lipofectamine 2000 (Invitrogen). Cell extracts PCR using Pfu polymerase, Dpn I treatment, and transformation were prepared 48 h after transfection. The luciferase activity was (Stratagene). The ORF of mouse Tet3, mutant Tet3 (H950D and evaluated by using the Dual-Luciferase Reporter Assay System ’ Y952A), Sfpr4, Pcdha4, and Pcdha7 were cloned into pPyCAGIP (Promega) according to the manufacturer s recommendations. vector (60). qRT-PCR. qRT-PCR was performed by using Universal SYBR Green Generation of Tet3 KO and Tet1/2/3 TKO mESCs. Tet3 KO and Tet1/2/3 Master Mix (Roche) and analyzed by a StepOne Plus real-time PCR ’ TKO mouse ESCs were generated from mice (C57BL/6 back- system (Applied Biosystems), according to the manufacturer sin- ground) bearing the individual floxed alleles (Fig. S1C) (14, 18), structions, and the data were normalized for Gapdh expression. followed by excision of the floxed exon by transient expression of The primers used for qRT-PCR are listed in Dataset S4. Cre recombinase. Immunohistochemistry. Immunohistochemistry was performed as mESC Culture and Differentiation. mESCs were maintained on mi- described (61) with the primary antibodies described below. For ∼ tomycin C-treated mouse embryonic fibroblasts (MEFs; feeders) in statistical analysis, 300 cells were examined for each experiment, standard medium (61). Neural differentiation in SFEB culture was which was repeated four times. Mouse embryos were fixed over- night in 4% (wt/vol) paraformaldehyde, saturated with 20% (wt/vol) performed as described (31) with a minor modification. -
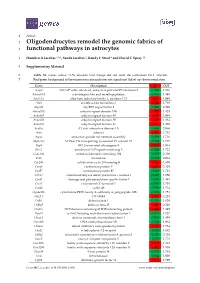
Oligodendrocytes Remodel the Genomic Fabrics of Functional Pathways in Astrocytes
1 Article 2 Oligodendrocytes remodel the genomic fabrics of 3 functional pathways in astrocytes 4 Dumitru A Iacobas 1,2,*, Sanda Iacobas 3, Randy F Stout 4 and David C Spray 2,5 5 Supplementary Material 6 Table S1. Genes whose >1.5x absolute fold-change did not meet the individual CUT criterion. 7 Red/green background of the expression ratio indicates not significant (false) up-/down-regulation. Gene Description X CUT Acap2 ArfGAP with coiled-coil, ankyrin repeat and PH domains 2 -1.540 1.816 Adamts18 a disintegrin-like and metallopeptidase -1.514 1.594 Akr1c12 aldo-keto reductase family 1, member C12 1.866 1.994 Alx3 aristaless-like homeobox 3 1.536 1.769 Alyref2 Aly/REF export factor 2 -1.880 2.208 Ankrd33b ankyrin repeat domain 33B 1.593 1.829 Ankrd45 ankyrin repeat domain 45 1.514 1.984 Ankrd50 ankyrin repeat domain 50 1.628 1.832 Ankrd61 ankyrin repeat domain 61 1.645 1.802 Arid1a AT rich interactive domain 1A -1.668 2.066 Artn artemin 1.524 1.732 Aspm abnormal spindle microtubule assembly -1.693 1.716 Atp6v1e1 ATPase, H+ transporting, lysosomal V1 subunit E1 -1.679 1.777 Bag4 BCL2-associated athanogene 4 1.723 1.914 Birc3 baculoviral IAP repeat-containing 3 -1.588 1.722 Ccdc104 coiled-coil domain containing 104 -1.819 2.130 Ccl2 chemokine -1.699 2.034 Cdc20b cell division cycle 20 homolog B 1.512 1.605 Cenpf centromere protein F 2.041 2.128 Cep97 centrosomal protein 97 -1.641 1.723 COX1 mitochondrially encoded cytochrome c oxidase I -1.607 1.650 Cpsf7 cleavage and polyadenylation specific factor 7 -1.635 1.891 Crct1 cysteine-rich -

Qt7s99h7j8 Nosplash 33D25b6
! ii! Acknowledgements Lots of people have helped me get to where I am today and I apologize if I have left anyone out. First, I would like to thank my mother and father for always pushing me to try harder and encouraging me to persist in my scientific career. Both of my parents encouraged me to think like a scientist from a young age. I don’t think I would be here today without that early upbringing. Joining the Pollard lab was the best decision I made in graduate school. I can’t emphasize the importance of finding a good mentor in school. My advisor, Katherine Pollard, provided me with a model of leadership that I will carry with me for the rest of my career. Additionally, I would like to emphasize that the lab is full of driven individuals that have supported me in all my scientific endeavors. Fellow graduate students, Aram Avila-Herrera and Genevieve Erwin Haliburton, guided much of my direction during the early years. In addition, postdoctoral scholars Nandita Garud, Hassan Samee, Patrick Bradley, and Geoffrey Fudenberg were key in helping me prepare for my future steps in my career. I want to thank my committee members for sitting through long meetings and giving me the feedback I needed. Nadav Ahituv, Benoit Bruneau, and Jeff Wall have given me valuable advice on whether to pursue certain directions in my research. I also want to acknowledge all the individuals that supported me personally in school. In particular, I have always valued Sara Calhoun’s well-thought-out advice and Rose ! iii! Citron’s insight when making important decisions. -

Human Transcription Factors Responsive to Initial Reprogramming Predominantly Undergo Legitimate Reprogramming During Fbroblast Conversion to Ipscs Ricardo R
www.nature.com/scientificreports OPEN Human transcription factors responsive to initial reprogramming predominantly undergo legitimate reprogramming during fbroblast conversion to iPSCs Ricardo R. Cevallos1, Yvonne J. K. Edwards1,2, John M. Parant3, Bradley K. Yoder2 & Kejin Hu1* The four transcription factors OCT4, SOX2, KLF4, and MYC (OSKM) together can convert human fbroblasts to induced pluripotent stem cells (iPSCs). It is, however, perplexing that they can do so only for a rare population of the starting cells with a long latency. Transcription factors (TFs) defne identities of both the starting fbroblasts and the end product, iPSCs, and are also of paramount importance for the reprogramming process. It is critical to upregulate or activate the iPSC-enriched TFs while downregulate or silence the fbroblast-enriched TFs. This report explores the initial TF responses to OSKM as the molecular underpinnings for both the potency aspects and the limitation sides of the OSKM reprogramming. The authors frst defned the TF reprogramome, i.e., the full complement of TFs to be reprogrammed. Most TFs were resistant to OSKM reprogramming at the initial stages, an observation consistent with the inefciency and long latency of iPSC reprogramming. Surprisingly, the current analyses also revealed that most of the TFs (at least 83 genes) that did respond to OSKM induction underwent legitimate reprogramming. The initial legitimate transcriptional responses of TFs to OSKM reprogramming were also observed in the reprogramming fbroblasts from a diferent individual. Such early biased legitimate reprogramming of the responsive TFs aligns well with the robustness aspect of the otherwise inefcient and stochastic OSKM reprogramming. OCT4, SOX2, KLF4, and MYC (collectively OSKM) can convert human fbroblasts into induced pluripotent stem cells (iPSCs), which are the man-made version of embryonic stem cells (ESCs)1–3. -

Gli3 Utilizes Hand2 to Synergistically Regulate Tissue-Specific
RESEARCH ARTICLE Gli3 utilizes Hand2 to synergistically regulate tissue-specific transcriptional networks Kelsey H Elliott1,2,3, Xiaoting Chen4, Joseph Salomone1,3,5, Praneet Chaturvedi1, Preston A Schultz1,2, Sai K Balchand1,2, Jeffrey D Servetas6, Aime´ e Zuniga7, Rolf Zeller7, Brian Gebelein1, Matthew T Weirauch1,4, Kevin A Peterson6*, Samantha A Brugmann1,2,8* 1Division of Developmental Biology, Cincinnati Children’s Hospital Medical Center, Cincinnati, United States; 2Division of Plastic Surgery, Department of Surgery, Cincinnati Children’s Hospital Medical Center, Cincinnati, United States; 3Graduate Program in Molecular and Developmental Biology, Cincinnati Children’s Hospital Research Foundation, Cincinnati, United States; 4Center for Autoimmune Genomics and Etiology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, United States; 5Medical-Scientist Training Program, University of Cincinnati College of Medicine, Cincinnati, United States; 6Jackson Laboratory, Bar Harbor, United States; 7Developmental Genetics, Department of Biomedicine, University of Basel, Basel, Switzerland; 8Shriners Children’s Hospital, Cincinnati, United States Abstract Despite a common understanding that Gli TFs are utilized to convey a Hh morphogen gradient, genetic analyses suggest craniofacial development does not completely fit this paradigm. Using the mouse model (Mus musculus), we demonstrated that rather than being driven by a Hh *For correspondence: threshold, robust Gli3 transcriptional activity during skeletal and glossal development required [email protected] (KAP); interaction with the basic helix-loop-helix TF Hand2. Not only did genetic and expression data [email protected] support a co-factorial relationship, but genomic analysis revealed that Gli3 and Hand2 were (SAB) enriched at regulatory elements for genes essential for mandibular patterning and development. -

Jnsclcase20102 1..5
J Neurosurg Case Lessons 1(3):CASE20102, 2021 DOI: 10.3171/CASE20102 Bilateral squamosal synostosis: unusual presentation of chromosome 1p12–1p13.3 deletion. Illustrative case Sarut Chaisrisawadisuk, MD, FRCST,1,4 Nithiwat Vatanavicharn, MD,2 Verayuth Praphanphoj, MD,3 Peter J. Anderson, DSc, FRACS,4,5 and Mark H. Moore, MBChB, FRACS4 1Division of Plastic Surgery, Department of Surgery, and 2Division of Medical Genetics, Department of Paediatrics, Faculty of Medicine Siriraj Hospital, Mahidol University, Bangkok, Thailand; 3Medical Genetics Center, Sai Mai District, Bangkok, Thailand; 4Cleft and Craniofacial South Australia, Women’s and Children’s Hospital, North Adelaide, South Australia, Australia; and 5Faculty of Health and Medical Sciences, University of Adelaide, Adelaide, South Australia, Australia BACKGROUND Squamosal sutures are minor sutures of the human skull. Early isolated fusion of the sutures (squamosal synostosis) is rarely found. OBSERVATIONS The authors report a case of a girl who presented with an abnormal head shape and bilateral squamosal synostosis. Genetic testing revealed a chromosome 1p12–1p13.3 deletion. She has been managed with conservative treatment of the synostosis. She has global developmental delay and multiple anomalies due to the chromosome abnormality. LESSONS Isolated squamosal suture synostosis could be an uncommon feature of chromosome 1p12–1p13.3 deletion. https://thejns.org/doi/abs/10.3171/CASE20102 KEYWORDS cranial sutures; craniosynostosis; chromosome 1p12–1p13.3 deletion The squamosal suture, one of the minor sutures of the human revealed chromosome 46,XX. She was born at 38 weeks of gestation. calvaria, sits between the temporal and parietal bones and extends Her physical examination at 1 year of age showed a widening of both from the pterion, curving around the temporal squama.1 The suture is parietal bones and an open anterior fontanelle.