Transcriptional Programs During Mammalian Cell Prolifération
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Development and Validation of a Novel Immune-Related Prognostic Model
Wang et al. J Transl Med (2020) 18:67 https://doi.org/10.1186/s12967-020-02255-6 Journal of Translational Medicine RESEARCH Open Access Development and validation of a novel immune-related prognostic model in hepatocellular carcinoma Zheng Wang1, Jie Zhu1, Yongjuan Liu3, Changhong Liu2, Wenqi Wang2, Fengzhe Chen1* and Lixian Ma1* Abstract Background: Growing evidence has suggested that immune-related genes play crucial roles in the development and progression of hepatocellular carcinoma (HCC). Nevertheless, the utility of immune-related genes for evaluating the prognosis of HCC patients are still lacking. The study aimed to explore gene signatures and prognostic values of immune-related genes in HCC. Methods: We comprehensively integrated gene expression data acquired from 374 HCC and 50 normal tissues in The Cancer Genome Atlas (TCGA). Diferentially expressed genes (DEGs) analysis and univariate Cox regression analysis were performed to identify DEGs that related to overall survival. An immune prognostic model was constructed using the Lasso and multivariate Cox regression analyses. Furthermore, Cox regression analysis was applied to identify independent prognostic factors in HCC. The correlation analysis between immune-related signature and immune cells infltration were also investigated. Finally, the signature was validated in an external independent dataset. Results: A total of 329 diferentially expressed immune‐related genes were detected. 64 immune‐related genes were identifed to be markedly related to overall survival in HCC patients using univariate Cox regression analysis. Then we established a TF-mediated network for exploring the regulatory mechanisms of these genes. Lasso and multivariate Cox regression analyses were applied to construct the immune-based prognostic model, which consisted of nine immune‐related genes. -

A Curated Database of Experimentally Identified Interaction Proteins Of
International Journal of Molecular Sciences Article OGT Protein Interaction Network (OGT-PIN): A Curated Database of Experimentally Identified Interaction Proteins of OGT Junfeng Ma 1,* , Chunyan Hou 2, Yaoxiang Li 1 , Shufu Chen 3 and Ci Wu 1 1 Lombardi Comprehensive Cancer Center, Department of Oncology, Georgetown University Medical Center, Washington, DC 20057, USA; [email protected] (Y.L.); [email protected] (C.W.) 2 Dalian Institute of Chemical Physics, Chinese Academy of Sciences, Dalian 116023, China; [email protected] 3 School of Engineering, Pennsylvania State University Behrend, Erie, PA 16563, USA; [email protected] * Correspondence: [email protected]; Tel.: +1-202-6873802 Abstract: Interactions between proteins are essential to any cellular process and constitute the basis for molecular networks that determine the functional state of a cell. With the technical advances in recent years, an astonishingly high number of protein–protein interactions has been revealed. However, the interactome of O-linked N-acetylglucosamine transferase (OGT), the sole enzyme adding the O-linked β-N-acetylglucosamine (O-GlcNAc) onto its target proteins, has been largely undefined. To that end, we collated OGT interaction proteins experimentally identified in the past several decades. Rigorous curation of datasets from public repositories and O-GlcNAc-focused publications led to the identification of up to 929 high-stringency OGT interactors from multiple species studied (including Homo sapiens, Mus musculus, Rattus norvegicus, Drosophila melanogaster, Citation: Ma, J.; Hou, C.; Li, Y.; Chen, S.; Wu, C. OGT Protein Interaction Arabidopsis thaliana, and others). Among them, 784 human proteins were found to be interactors Network (OGT-PIN): A Curated of human OGT. -

Mining Microarray Data to Identify Transcription Factors Expressed in Naıve Resting but Not Activated T Lymphocytes
Genes and Immunity (2004) 5, 16–25 & 2004 Nature Publishing Group All rights reserved 1466-4879/04 $25.00 www.nature.com/gene Mining microarray data to identify transcription factors expressed in naı¨ve resting but not activated T lymphocytes C Argyropoulos1,2, GC Nikiforidis2, M Theodoropoulou1, P Adamopoulos1, S Boubali3, TN Georgakopoulos4, F Paliogianni3, AG Papavassiliou4 and A Mouzaki1 1Laboratory of Hematology & Transfusion Medicine, University of Patras, Patras, Greece; 2Laboratory of Medical Physics, University of Patras, Patras, Greece; 3Laboratory of Microbiology, University of Patras, Patras, Greece; 4Laboratory of Biological Chemistry, Medical School, University of Patras, Patras, Greece Transcriptional repressors controlling the expression of cytokine genes have been implicated in a variety of physiological and pathological phenomena. An unknown repressor that binds to the distal NFAT element of the interleukin-2 (IL-2) gene promoter in naive T-helper lymphocytes has been implicated in autoimmune phenomena and has emerged as a potentially important factor controlling the latency of HIV-1. The aim of this paper was the identification of this repressor. We resorted to public microarray databases looking for DNA-binding proteins that are present in naı¨ve resting T cells but are downregulated when the cells are activated. A Bayesian data mining statistical analysis uncovered 25 candidate factors. Of the 25, NFAT4 and the oncogene ets-2 bind to the common motif AAGGAG found in the HIV-1 LTR and IL-2 probes. Ets-2 binding site contains the three G’s that have been shown to be important for binding of the unknown factor; hence, we considered it the likeliest candidate. -

Supplementary Information.Pdf
Supplementary Information Whole transcriptome profiling reveals major cell types in the cellular immune response against acute and chronic active Epstein‐Barr virus infection Huaqing Zhong1, Xinran Hu2, Andrew B. Janowski2, Gregory A. Storch2, Liyun Su1, Lingfeng Cao1, Jinsheng Yu3, and Jin Xu1 Department of Clinical Laboratory1, Children's Hospital of Fudan University, Minhang District, Shanghai 201102, China; Departments of Pediatrics2 and Genetics3, Washington University School of Medicine, Saint Louis, Missouri 63110, United States. Supplementary information includes the following: 1. Supplementary Figure S1: Fold‐change and correlation data for hyperactive and hypoactive genes. 2. Supplementary Table S1: Clinical data and EBV lab results for 110 study subjects. 3. Supplementary Table S2: Differentially expressed genes between AIM vs. Healthy controls. 4. Supplementary Table S3: Differentially expressed genes between CAEBV vs. Healthy controls. 5. Supplementary Table S4: Fold‐change data for 303 immune mediators. 6. Supplementary Table S5: Primers used in qPCR assays. Supplementary Figure S1. Fold‐change (a) and Pearson correlation data (b) for 10 cell markers and 61 hypoactive and hyperactive genes identified in subjects with acute EBV infection (AIM) in the primary cohort. Note: 23 up‐regulated hyperactive genes were highly correlated positively with cytotoxic T cell (Tc) marker CD8A and NK cell marker CD94 (KLRD1), and 38 down‐regulated hypoactive genes were highly correlated positively with B cell, conventional dendritic cell -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -

Emerging Multifaceted Roles of BAP1 Complexes in Biological Processes Aileen Patricia Szczepanski1 and Lu Wang1
Szczepanski and Wang Cell Death Discovery (2021) 7:20 https://doi.org/10.1038/s41420-021-00406-2 Cell Death Discovery REVIEW ARTICLE Open Access Emerging multifaceted roles of BAP1 complexes in biological processes Aileen Patricia Szczepanski1 and Lu Wang1 Abstract Histone H2AK119 mono-ubiquitination (H2AK119Ub) is a relatively abundant histone modification, mainly catalyzed by the Polycomb Repressive Complex 1 (PRC1) to regulate Polycomb-mediated transcriptional repression of downstream target genes. Consequently, H2AK119Ub can also be dynamically reversed by the BAP1 complex, an evolutionarily conserved multiprotein complex that functions as a general transcriptional activator. In previous studies, it has been reported that the BAP1 complex consists of important biological roles in development, metabolism, and cancer. However, identifying the BAP1 complex’s regulatory mechanisms remains to be elucidated due to its various complex forms and its ability to target non-histone substrates. In this review, we will summarize recent findings that have contributed to the diverse functional role of the BAP1 complex and further discuss the potential in targeting BAP1 for therapeutic use. Facts levels for catalytic activity and accurate cellular localization, which are critical for the determinant of ● cell fate and transformation. The BAP1 complex has 1234567890():,; 1234567890():,; 1234567890():,; 1234567890():,; The BAP1 complex is an evolutionary conserved, emerged as an ideal therapeutic target for treatments multiprotein complex that functions -

Modulating BAP1 Expression Affects ROS Homeostasis, Cell Motility And
Modulating BAP1 expression affects ROS homeostasis, cell motility and mitochondrial function Lucie Hébert, Dorine Bellanger, Chloé Guillas, Antoine Campagne, Florent Dingli, Damarys Loew, Alice Fievet, Virginie Jacquemin, Tatiana Popova, Didier Jean, et al. To cite this version: Lucie Hébert, Dorine Bellanger, Chloé Guillas, Antoine Campagne, Florent Dingli, et al.. Modulating BAP1 expression affects ROS homeostasis, cell motility and mitochondrial function. Oncotarget, Impact journals, 2017, 8 (42), pp.72513-72527. 10.18632/oncotarget.19872. inserm-02478605 HAL Id: inserm-02478605 https://www.hal.inserm.fr/inserm-02478605 Submitted on 14 Feb 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 42), pp: 72513-72527 Research Paper Modulating BAP1 expression affects ROS homeostasis, cell motility and mitochondrial function Lucie Hebert1, Dorine Bellanger1, Chloé Guillas1, Antoine Campagne2, Florent Dingli3, Damarys Loew3, Alice Fievet1,4, Virginie Jacquemin1, Tatiana Popova1, Didier -

Identification of Genomic Targets of Krüppel-Like Factor 9 in Mouse Hippocampal
Identification of Genomic Targets of Krüppel-like Factor 9 in Mouse Hippocampal Neurons: Evidence for a role in modulating peripheral circadian clocks by Joseph R. Knoedler A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Neuroscience) in the University of Michigan 2016 Doctoral Committee: Professor Robert J. Denver, Chair Professor Daniel Goldman Professor Diane Robins Professor Audrey Seasholtz Associate Professor Bing Ye ©Joseph R. Knoedler All Rights Reserved 2016 To my parents, who never once questioned my decision to become the other kind of doctor, And to Lucy, who has pushed me to be a better person from day one. ii Acknowledgements I have a huge number of people to thank for having made it to this point, so in no particular order: -I would like to thank my adviser, Dr. Robert J. Denver, for his guidance, encouragement, and patience over the last seven years; his mentorship has been indispensable for my growth as a scientist -I would also like to thank my committee members, Drs. Audrey Seasholtz, Dan Goldman, Diane Robins and Bing Ye, for their constructive feedback and their willingness to meet in a frequently cold, windowless room across campus from where they work -I am hugely indebted to Pia Bagamasbad and Yasuhiro Kyono for teaching me almost everything I know about molecular biology and bioinformatics, and to Arasakumar Subramani for his tireless work during the home stretch to my dissertation -I am grateful for the Neuroscience Program leadership and staff, in particular -

Is Regulated by O-Glcnac Transferase
UC Irvine UC Irvine Previously Published Works Title Nuclear factor-erythroid-2 related transcription factor-1 (Nrf1) is regulated by O-GlcNAc transferase. Permalink https://escholarship.org/uc/item/5mp8q65s Journal Free radical biology & medicine, 110 ISSN 0891-5849 Authors Han, Jeong Woo Valdez, Joshua L Ho, Daniel V et al. Publication Date 2017-09-01 DOI 10.1016/j.freeradbiomed.2017.06.008 License https://creativecommons.org/licenses/by/4.0/ 4.0 Peer reviewed eScholarship.org Powered by the California Digital Library University of California Free Radical Biology and Medicine 110 (2017) 196–205 Contents lists available at ScienceDirect Free Radical Biology and Medicine journal homepage: www.elsevier.com/locate/freeradbiomed Original article Nuclear factor-erythroid-2 related transcription factor-1 (Nrf1) is regulated MARK by O-GlcNAc transferase Jeong Woo Hana,1, Joshua L. Valdeza,1, Daniel V. Hoa, Candy S. Leea, Hyun Min Kima, ⁎ Xiaorong Wangb, Lan Huangb,Jefferson Y. Chana, a Department of Laboratory Medicine and Pathology, University of California, Irvine, D440 Medical Sciences, Irvine, CA 92697, USA b Departments of Physiology and Biophysics, University of California, Irvine, D440 Medical Sciences, Irvine, CA 92697, USA ARTICLE INFO ABSTRACT Keywords: The Nrf1 (Nuclear factor E2-related factor 1) transcription factor performs a critical role in regulating cellular Oxidative stress homeostasis. Using a proteomic approach, we identified Host Cell Factor-1 (HCF1), a co-regulator of tran- O-GlcNAcylation scription, and O-GlcNAc transferase (OGT), the enzyme that mediates protein O-GlcNAcylation, as cellular OGT partners of Nrf1a, an isoform of Nrf1. Nrf1a directly interacts with HCF1 through the HCF1 binding motif Transcription factor (HBM), while interaction with OGT is mediated through HCF1. -

(12) Patent Application Publication (10) Pub. No.: US 2009/0269772 A1 Califano Et Al
US 20090269772A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2009/0269772 A1 Califano et al. (43) Pub. Date: Oct. 29, 2009 (54) SYSTEMS AND METHODS FOR Publication Classification IDENTIFYING COMBINATIONS OF (51) Int. Cl. COMPOUNDS OF THERAPEUTIC INTEREST CI2O I/68 (2006.01) CI2O 1/02 (2006.01) (76) Inventors: Andrea Califano, New York, NY G06N 5/02 (2006.01) (US); Riccardo Dalla-Favera, New (52) U.S. Cl. ........... 435/6: 435/29: 706/54; 707/E17.014 York, NY (US); Owen A. (57) ABSTRACT O'Connor, New York, NY (US) Systems, methods, and apparatus for searching for a combi nation of compounds of therapeutic interest are provided. Correspondence Address: Cell-based assays are performed, each cell-based assay JONES DAY exposing a different sample of cells to a different compound 222 EAST 41ST ST in a plurality of compounds. From the cell-based assays, a NEW YORK, NY 10017 (US) Subset of the tested compounds is selected. For each respec tive compound in the Subset, a molecular abundance profile from cells exposed to the respective compound is measured. (21) Appl. No.: 12/432,579 Targets of transcription factors and post-translational modu lators of transcription factor activity are inferred from the (22) Filed: Apr. 29, 2009 molecular abundance profile data using information theoretic measures. This data is used to construct an interaction net Related U.S. Application Data work. Variances in edges in the interaction network are used to determine the drug activity profile of compounds in the (60) Provisional application No. 61/048.875, filed on Apr. -

A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression
Supplementary Materials A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression Table S1. Statistically significant DEGs (Adj. p-value < 0.01) derived from meta-analysis for samples irradiated with high doses of HZE particles, collected 6-24 h post-IR not common with any other meta- analysis group. This meta-analysis group consists of 3 DEG lists obtained from DGEA, using a total of 11 control and 11 irradiated samples [Data Series: E-MTAB-5761 and E-MTAB-5754]. Ensembl ID Gene Symbol Gene Description Up-Regulated Genes ↑ (2425) ENSG00000000938 FGR FGR proto-oncogene, Src family tyrosine kinase ENSG00000001036 FUCA2 alpha-L-fucosidase 2 ENSG00000001084 GCLC glutamate-cysteine ligase catalytic subunit ENSG00000001631 KRIT1 KRIT1 ankyrin repeat containing ENSG00000002079 MYH16 myosin heavy chain 16 pseudogene ENSG00000002587 HS3ST1 heparan sulfate-glucosamine 3-sulfotransferase 1 ENSG00000003056 M6PR mannose-6-phosphate receptor, cation dependent ENSG00000004059 ARF5 ADP ribosylation factor 5 ENSG00000004777 ARHGAP33 Rho GTPase activating protein 33 ENSG00000004799 PDK4 pyruvate dehydrogenase kinase 4 ENSG00000004848 ARX aristaless related homeobox ENSG00000005022 SLC25A5 solute carrier family 25 member 5 ENSG00000005108 THSD7A thrombospondin type 1 domain containing 7A ENSG00000005194 CIAPIN1 cytokine induced apoptosis inhibitor 1 ENSG00000005381 MPO myeloperoxidase ENSG00000005486 RHBDD2 rhomboid domain containing 2 ENSG00000005884 ITGA3 integrin subunit alpha 3 ENSG00000006016 CRLF1 cytokine receptor like -
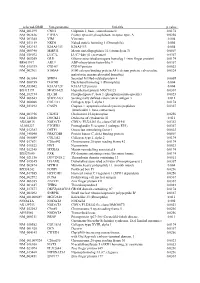
Supplementary Figures S1-S3
selected-GBID Uni-genename Uni-title p value NM_001299 CNN1 Calponin 1, basic, smooth muscle 0.0174 NM_002836 PTPRA Protein tyrosine phosphatase, receptor type, A 0.0256 NM_003380 VIM Vimentin 0.004 NM_033119 NKD1 Naked cuticle homolog 1 (Drosophila) 0.004 NM_052913 KIAA1913 KIAA1913 0.004 NM_005940 MMP11 Matrix metallopeptidase 11 (stromelysin 3) 0.0069 NM_018032 LUC7L LUC7-like (S. cerevisiae) 0.0367 NM_005269 GLI1 Glioma-associated oncogene homolog 1 (zinc finger protein) 0.0174 BE463997 ARL9 ADP-ribosylation factor-like 9 0.0367 NM_015939 CGI-09 CGI-09 protein 0.0023 NM_002961 S100A4 S100 calcium binding protein A4 (calcium protein, calvasculin, 0.0324 metastasin, murine placental homolog) NM_003014 SFRP4 Secreted frizzled-related protein 4 0.0005 NM_080759 DACH1 Dachshund homolog 1 (Drosophila) 0.004 NM_053042 KIAA1729 KIAA1729 protein 0.004 BX415194 MGC16121 Hypothetical protein MGC16121 0.0367 NM_182734 PLCB1 Phospholipase C, beta 1 (phosphoinositide-specific) 0.0023 NM_006643 SDCCAG3 Serologically defined colon cancer antigen 3 0.011 NM_000088 COL1A1 Collagen, type I, alpha 1 0.0174 NM_033292 CASP1 Caspase 1, apoptosis-related cysteine peptidase 0.0367 (interleukin 1, beta, convertase) NM_003956 CH25H Cholesterol 25-hydroxylase 0.0256 NM_144658 DOCK11 Dedicator of cytokinesis 11 0.011 AK024935 NODATA CDNA: FLJ21283 fis, clone COL01910 0.0363 AL050227 PTGER3 Prostaglandin E receptor 3 (subtype EP3) 0.0367 NM_012383 OSTF1 Osteoclast stimulating factor 1 0.0023 NM_145040 PRKCDBP Protein kinase C, delta binding protein 0.0069 NM_000089