Temperature-Sensitive Mutation of PEX6 in Peroxisome Biogenesis Disorders in Complementation Group C (CG-C): Comparative Study of PEX6 and PEX1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Ophthalmic Manifestations of Heimler Syndrome Due to PEX6 Mutations
Thomas Jefferson University Jefferson Digital Commons Wills Eye Hospital Papers Wills Eye Hospital 5-4-2018 Ophthalmic manifestations of Heimler syndrome due to PEX6 mutations. Nutsuchar Wangtiraumnuay Wills Eye Hospital; Queen Sirikit National Institute of Child Health Waleed Abed Alnabi Wills Eye Hospital Mai Tsukikawa Thomas Jefferson University Avrey Thau Wills Eye Hosptial; Thomas Jefferson University Jenina Capasso Wills Eye Hospital Follow this and additional works at: https://jdc.jefferson.edu/willsfp Part of the Ophthalmology Commons LetSee next us page know for additional how authors access to this document benefits ouy Recommended Citation Wangtiraumnuay, Nutsuchar; Alnabi, Waleed Abed; Tsukikawa, Mai; Thau, Avrey; Capasso, Jenina; Sharony, Reuven; Inglehearn, Chris F.; and Levin, Alex V., "Ophthalmic manifestations of Heimler syndrome due to PEX6 mutations." (2018). Wills Eye Hospital Papers. Paper 83. https://jdc.jefferson.edu/willsfp/83 This Article is brought to you for free and open access by the Jefferson Digital Commons. The Jefferson Digital Commons is a service of Thomas Jefferson University's Center for Teaching and Learning (CTL). The Commons is a showcase for Jefferson books and journals, peer-reviewed scholarly publications, unique historical collections from the University archives, and teaching tools. The Jefferson Digital Commons allows researchers and interested readers anywhere in the world to learn about and keep up to date with Jefferson scholarship. This article has been accepted for inclusion in Wills Eye Hospital Papers by an authorized administrator of the Jefferson Digital Commons. For more information, please contact: [email protected]. Authors Nutsuchar Wangtiraumnuay, Waleed Abed Alnabi, Mai Tsukikawa, Avrey Thau, Jenina Capasso, Reuven Sharony, Chris F. -

Propranolol-Mediated Attenuation of MMP-9 Excretion in Infants with Hemangiomas
Supplementary Online Content Thaivalappil S, Bauman N, Saieg A, Movius E, Brown KJ, Preciado D. Propranolol-mediated attenuation of MMP-9 excretion in infants with hemangiomas. JAMA Otolaryngol Head Neck Surg. doi:10.1001/jamaoto.2013.4773 eTable. List of All of the Proteins Identified by Proteomics This supplementary material has been provided by the authors to give readers additional information about their work. © 2013 American Medical Association. All rights reserved. Downloaded From: https://jamanetwork.com/ on 10/01/2021 eTable. List of All of the Proteins Identified by Proteomics Protein Name Prop 12 mo/4 Pred 12 mo/4 Δ Prop to Pred mo mo Myeloperoxidase OS=Homo sapiens GN=MPO 26.00 143.00 ‐117.00 Lactotransferrin OS=Homo sapiens GN=LTF 114.00 205.50 ‐91.50 Matrix metalloproteinase‐9 OS=Homo sapiens GN=MMP9 5.00 36.00 ‐31.00 Neutrophil elastase OS=Homo sapiens GN=ELANE 24.00 48.00 ‐24.00 Bleomycin hydrolase OS=Homo sapiens GN=BLMH 3.00 25.00 ‐22.00 CAP7_HUMAN Azurocidin OS=Homo sapiens GN=AZU1 PE=1 SV=3 4.00 26.00 ‐22.00 S10A8_HUMAN Protein S100‐A8 OS=Homo sapiens GN=S100A8 PE=1 14.67 30.50 ‐15.83 SV=1 IL1F9_HUMAN Interleukin‐1 family member 9 OS=Homo sapiens 1.00 15.00 ‐14.00 GN=IL1F9 PE=1 SV=1 MUC5B_HUMAN Mucin‐5B OS=Homo sapiens GN=MUC5B PE=1 SV=3 2.00 14.00 ‐12.00 MUC4_HUMAN Mucin‐4 OS=Homo sapiens GN=MUC4 PE=1 SV=3 1.00 12.00 ‐11.00 HRG_HUMAN Histidine‐rich glycoprotein OS=Homo sapiens GN=HRG 1.00 12.00 ‐11.00 PE=1 SV=1 TKT_HUMAN Transketolase OS=Homo sapiens GN=TKT PE=1 SV=3 17.00 28.00 ‐11.00 CATG_HUMAN Cathepsin G OS=Homo -
The Peroxisomal PTS1-Import Defect of PEX1- Deficient Cells Is Independent of Pexophagy in Saccharomyces Cerevisiae
International Journal of Molecular Sciences Communication The Peroxisomal PTS1-Import Defect of PEX1- Deficient Cells Is Independent of Pexophagy in Saccharomyces cerevisiae Thomas Mastalski, Rebecca Brinkmeier and Harald W. Platta * Biochemie Intrazellulärer Transportprozesse, Ruhr-Universität Bochum, Universitätsstr. 150, 44801 Bochum, Germany; [email protected] (T.M.); [email protected] (R.B.) * Correspondence: [email protected]; Tel.: +49-234-3224-968 Received: 10 January 2020; Accepted: 27 January 2020; Published: 29 January 2020 Abstract: The important physiologic role of peroxisomes is shown by the occurrence of peroxisomal biogenesis disorders (PBDs) in humans. This spectrum of autosomal recessive metabolic disorders is characterized by defective peroxisome assembly and impaired peroxisomal functions. PBDs are caused by mutations in the peroxisomal biogenesis factors, which are required for the correct compartmentalization of peroxisomal matrix enzymes. Recent work from patient cells that contain the Pex1(G843D) point mutant suggested that the inhibition of the lysosome, and therefore the block of pexophagy, was beneficial for peroxisomal function. The resulting working model proposed that Pex1 may not be essential for matrix protein import at all, but rather for the prevention of pexophagy. Thus, the observed matrix protein import defect would not be caused by a lack of Pex1 activity, but rather by enhanced removal of peroxisomal membranes via pexophagy. In the present study, we can show that the specific block of PEX1 deletion-induced pexophagy does not restore peroxisomal matrix protein import or the peroxisomal function in beta-oxidation in yeast. Therefore, we conclude that Pex1 is directly and essentially involved in peroxisomal matrix protein import, and that the PEX1 deletion-induced pexophagy is not responsible for the defect in peroxisomal function. -

Spectrum of PEX1 and PEX6 Variants in Heimler Syndrome
European Journal of Human Genetics (2016) 24, 1565–1571 Official Journal of The European Society of Human Genetics www.nature.com/ejhg ARTICLE Spectrum of PEX1 and PEX6 variants in Heimler syndrome Claire EL Smith1, James A Poulter1, Alex V Levin2,3,4, Jenina E Capasso4, Susan Price5, Tamar Ben-Yosef6, Reuven Sharony7, William G Newman8,9, Roger C Shore10, Steven J Brookes10, Alan J Mighell1,11,12 and Chris F Inglehearn*,1,12 Heimler syndrome (HS) consists of recessively inherited sensorineural hearing loss, amelogenesis imperfecta (AI) and nail abnormalities, with or without visual defects. Recently HS was shown to result from hypomorphic mutations in PEX1 or PEX6,both previously implicated in Zellweger Syndrome Spectrum Disorders (ZSSD). ZSSD are a group of conditions consisting of craniofacial and neurological abnormalities, sensory defects and multi-organ dysfunction. The finding of HS-causing mutations in PEX1 and PEX6 shows that HS represents the mild end of the ZSSD spectrum, though these conditions were previously thought to be distinct nosological entities. Here, we present six further HS families, five with PEX6 variants and one with PEX1 variants, and show the patterns of Pex1, Pex14 and Pex6 immunoreactivity in the mouse retina. While Ratbi et al. found more HS-causing mutations in PEX1 than in PEX6, as is the case for ZSSD, in this cohort PEX6 variants predominate, suggesting both genes play a significant role in HS. The PEX6 variant c.1802G4A, p.(R601Q), reported previously in compound heterozygous state in one HS and three ZSSD cases, was found in compound heterozygous state in three HS families. -

Common Variants in SOX-2 and Congenital Cataract Genes Contribute to Age-Related Nuclear Cataract
ARTICLE https://doi.org/10.1038/s42003-020-01421-2 OPEN Common variants in SOX-2 and congenital cataract genes contribute to age-related nuclear cataract Ekaterina Yonova-Doing et al.# 1234567890():,; Nuclear cataract is the most common type of age-related cataract and a leading cause of blindness worldwide. Age-related nuclear cataract is heritable (h2 = 0.48), but little is known about specific genetic factors underlying this condition. Here we report findings from the largest to date multi-ethnic meta-analysis of genome-wide association studies (discovery cohort N = 14,151 and replication N = 5299) of the International Cataract Genetics Consortium. We confirmed the known genetic association of CRYAA (rs7278468, P = 2.8 × 10−16) with nuclear cataract and identified five new loci associated with this dis- ease: SOX2-OT (rs9842371, P = 1.7 × 10−19), TMPRSS5 (rs4936279, P = 2.5 × 10−10), LINC01412 (rs16823886, P = 1.3 × 10−9), GLTSCR1 (rs1005911, P = 9.8 × 10−9), and COMMD1 (rs62149908, P = 1.2 × 10−8). The results suggest a strong link of age-related nuclear cat- aract with congenital cataract and eye development genes, and the importance of common genetic variants in maintaining crystalline lens integrity in the aging eye. #A list of authors and their affiliations appears at the end of the paper. COMMUNICATIONS BIOLOGY | (2020) 3:755 | https://doi.org/10.1038/s42003-020-01421-2 | www.nature.com/commsbio 1 ARTICLE COMMUNICATIONS BIOLOGY | https://doi.org/10.1038/s42003-020-01421-2 ge-related cataract is the leading cause of blindness, structure (meta-analysis genomic inflation factor λ = 1.009, accounting for more than one-third of blindness Supplementary Table 4 and Supplementary Fig. -

Peroxisomal Disorders and Their Mouse Models Point to Essential Roles of Peroxisomes for Retinal Integrity
International Journal of Molecular Sciences Review Peroxisomal Disorders and Their Mouse Models Point to Essential Roles of Peroxisomes for Retinal Integrity Yannick Das, Daniëlle Swinkels and Myriam Baes * Lab of Cell Metabolism, Department of Pharmaceutical and Pharmacological Sciences, KU Leuven, 3000 Leuven, Belgium; [email protected] (Y.D.); [email protected] (D.S.) * Correspondence: [email protected] Abstract: Peroxisomes are multifunctional organelles, well known for their role in cellular lipid homeostasis. Their importance is highlighted by the life-threatening diseases caused by peroxisomal dysfunction. Importantly, most patients suffering from peroxisomal biogenesis disorders, even those with a milder disease course, present with a number of ocular symptoms, including retinopathy. Patients with a selective defect in either peroxisomal α- or β-oxidation or ether lipid synthesis also suffer from vision problems. In this review, we thoroughly discuss the ophthalmological pathology in peroxisomal disorder patients and, where possible, the corresponding animal models, with a special emphasis on the retina. In addition, we attempt to link the observed retinal phenotype to the underlying biochemical alterations. It appears that the retinal pathology is highly variable and the lack of histopathological descriptions in patients hampers the translation of the findings in the mouse models. Furthermore, it becomes clear that there are still large gaps in the current knowledge on the contribution of the different metabolic disturbances to the retinopathy, but branched chain fatty acid accumulation and impaired retinal PUFA homeostasis are likely important factors. Citation: Das, Y.; Swinkels, D.; Baes, Keywords: peroxisome; Zellweger; metabolism; fatty acid; retina M. Peroxisomal Disorders and Their Mouse Models Point to Essential Roles of Peroxisomes for Retinal Integrity. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Full Disclosure Forms
Expanding genotype/phenotype of neuromuscular diseases by comprehensive target capture/NGS Xia Tian, PhD* ABSTRACT * Wen-Chen Liang, MD Objective: To establish and evaluate the effectiveness of a comprehensive next-generation * Yanming Feng, PhD sequencing (NGS) approach to simultaneously analyze all genes known to be responsible for Jing Wang, MD the most clinically and genetically heterogeneous neuromuscular diseases (NMDs) involving spi- Victor Wei Zhang, PhD nal motoneurons, neuromuscular junctions, nerves, and muscles. Chih-Hung Chou, MS Methods: All coding exons and at least 20 bp of flanking intronic sequences of 236 genes causing Hsien-Da Huang, PhD NMDs were enriched by using SeqCap EZ solution-based capture and enrichment method fol- Ching Wan Lam, PhD lowed by massively parallel sequencing on Illumina HiSeq2000. Ya-Yun Hsu, PhD ; 3 Thy-Sheng Lin, MD Results: The target gene capture/deep sequencing provides an average coverage of 1,000 per Wan-Tzu Chen, MS nucleotide. Thirty-five unrelated NMD families (38 patients) with clinical and/or muscle pathologic Lee-Jun Wong, PhD diagnoses but without identified causative genetic defects were analyzed. Deleterious mutations Yuh-Jyh Jong, MD were found in 29 families (83%). Definitive causative mutations were identified in 21 families (60%) and likely diagnoses were established in 8 families (23%). Six families were left without diagnosis due to uncertainty in phenotype/genotype correlation and/or unidentified causative Correspondence to genes. Using this comprehensive panel, we not only identified mutations in expected genes but Dr. Wong: also expanded phenotype/genotype among different subcategories of NMDs. [email protected] or Dr. Jong: Conclusions: Target gene capture/deep sequencing approach can greatly improve the genetic [email protected] diagnosis of NMDs. -

Perkinelmer Genomics to Request the Saliva Swab Collection Kit for Patients That Cannot Provide a Blood Sample As Whole Blood Is the Preferred Sample
Zellweger Spectrum Disorder Panel Test Code D4611 Test Summary This panel analyzes 15 genes that have been associated with Zellweger Spectrum Disorders. Turn-Around-Time (TAT)* 3 - 5 weeks Acceptable Sample Types Whole Blood (EDTA) (Preferred sample type) DNA, Isolated Dried Blood Spots Saliva Acceptable Billing Types Self (patient) Payment Institutional Billing Commercial Insurance Indications for Testing This panel may be appropriate for individuals with signs and symptoms of a peroxisomal biogenesis disfuction and/or Zellweger syndrome spectrum disorder and/or a family history of these conditions. Test Description This panel analyzes 15 genes that have been associated with Zellweger Spectrum Disorders. Both sequencing and deletion/duplication (CNV) analysis will be performed on the coding regions of all genes included (unless otherwise marked). All analysis is performed utilizing Next Generation Sequencing (NGS) technology. CNV analysis is designed to detect the majority of deletions and duplications of three exons or greater in size. Smaller CNV events may also be detected and reported, but additional follow-up testing is recommended if a smaller CNV is suspected. All variants are classified according to ACMG guidelines. Condition Description Zellweger syndrome spectrum disorders include Zellweger syndrome (ZS), neonatal adrenoleukodystrophy (NALD), an intermediate form and infantile Refsum disease (IRD). Symptoms may include hypotonia, feeding problems, hearing and vision loss, seizures, distinctive facial characteristics and skeletal abnormalities. Zellweger spectrum disorder is estimated to occur in 1/50,000 individuals. Genes ACOX1, AMACR, HSD17B4, PEX1, PEX10, PEX12, PEX13, PEX14, PEX16, PEX19, PEX2, PEX26, PEX3, PEX5, PEX6 Test Methods and Limitations Sequencing is performed on genomic DNA using an Agilent targeted sequence capture method to enrich for the genes on this panel. -

Perkinelmer Genomics
Adrenoleukodystrophy Panel with ABCD1 Test Code D4043 Test Summary This test analyzes 15 genes that have been associated with X-linked adrenoleukodystophy. Turn-Around-Time (TAT)* 3 - 5 weeks Acceptable Sample Types DNA, Isolated Dried Blood Spots Saliva Whole Blood (EDTA) Acceptable Billing Types Self (patient) Payment Institutional Billing Commercial Insurance Indications for Testing Children with symptoms of adrenal gland dysfunction, behavioral problems, learning difficulties, vision problems, poor coordination, and difficulty swallowing. Individuals with adrenocortical insufficiency, leg stiffness, behavioral changes, and urinary and genital tract disorders. Individuals with a family history a X-linked adrenoleukodystrophy. Test Description This panel analyzes 15 genes that have been associated with X-linked adrenoleukodystophy. Both sequencing and deletion/duplication (CNV) analysis will be performed on the coding regions of all genes included (unless otherwise marked). All analysis is performed utilizing Next Generation Sequencing (NGS) technology. CNV analysis is designed to detect the majority of deletions and duplications of three exons or greater in size. Smaller CNV events may also be detected and reported, but additional follow-up testing is recommended if a smaller CNV is suspected. All variants are classified according to ACMG guidelines. Condition Description X-linked adrenoleukodystrophy is a disease that primarily affects the nervous system and adrenal glands. There are 3 forms of X-linked adrenoleukodystrophy and the disease occurs mainly in males. The cerebral form of X-linked adrenoleukodystrophy typically has an age of onset in childhood with symptoms of learning difficulty, behavioral problems, vision problems, difficulty swallowing, poor coordination, and impaired adrenal function. Death usually occurs a few years after symptoms begin. -

Geneseq PLUS
GeneSeq® PLUS Specimen ID: 00000000050 Container ID: H0655 Control ID: Acct #: LCA-BN Phone: SAMPLE REPORT, F-630068 Patient Details Specimen Details Physician Details DOB: 05/05/1995 Date Collected: 08/05/2019 12:00 (Local) Ordering: Age (yyy/mm/dd): 024/07/04 Date Received: 08/06/2019 Referring: Gender: Female Date Entered: 08/06/2019 ID: Patient ID: 00000000050 Date Reported: 08/16/2019 13:21 (Local) NPI: Ethnicity: Unknown Specimen Type: Blood Lab ID: MNEGA Indication: MAN2B1 carrier testing Genetic Counselor: SUMMARY: POSITIVE POSITIVE RESULTS DISORDER (GENE) RESULTS INTERPRETATION Alpha-mannosidosis POSITIVE Predicted to be a carrier. Genetic counseling is (MAN2B1) Heterozygous for c.2248C>T recommended. NMID: NM_000528 (p.Arg750Trp), pathogenic Chr19:12760746 (GRCh37) Risk: AT INCREASED RISK FOR AFFECTED PREGNANCY. See Additional Clinical Information. Genetic counseling is recommended to discuss the potential clinical and/or reproductive implications of positive results, as well as recommendations for testing family members and, when applicable, this individual's partner. Genetic counseling services are available. To access Integrated Genetics Genetic Counselors please visit www.integratedgenetics.com/genetic-counseling or call (855) GC-CALLS (855-422-2557). ADDITIONAL CLINICAL INFORMATION Alpha-mannosidosis: Alpha-mannosidosis is an autosomal recessive lysosomal storage disorder with variable severity and age at onset. Signs and symptoms may include progressive neurological deterioration, intellectual disability, skeletal and facial abnormalities, immunodeficiency, and hearing impairment. Bone marrow transplant may be an option for some individuals. Treatment is otherwise supportive. (Malm, PMID:18651971). If this individual's reproductive partner is also a carrier of a pathogenic variant in the same gene the risk for an affected fetus is 25%. -
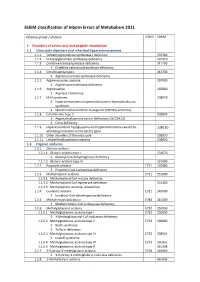
SSIEM Classification of Inborn Errors of Metabolism 2011
SSIEM classification of Inborn Errors of Metabolism 2011 Disease group / disease ICD10 OMIM 1. Disorders of amino acid and peptide metabolism 1.1. Urea cycle disorders and inherited hyperammonaemias 1.1.1. Carbamoylphosphate synthetase I deficiency 237300 1.1.2. N-Acetylglutamate synthetase deficiency 237310 1.1.3. Ornithine transcarbamylase deficiency 311250 S Ornithine carbamoyltransferase deficiency 1.1.4. Citrullinaemia type1 215700 S Argininosuccinate synthetase deficiency 1.1.5. Argininosuccinic aciduria 207900 S Argininosuccinate lyase deficiency 1.1.6. Argininaemia 207800 S Arginase I deficiency 1.1.7. HHH syndrome 238970 S Hyperammonaemia-hyperornithinaemia-homocitrullinuria syndrome S Mitochondrial ornithine transporter (ORNT1) deficiency 1.1.8. Citrullinemia Type 2 603859 S Aspartate glutamate carrier deficiency ( SLC25A13) S Citrin deficiency 1.1.9. Hyperinsulinemic hypoglycemia and hyperammonemia caused by 138130 activating mutations in the GLUD1 gene 1.1.10. Other disorders of the urea cycle 238970 1.1.11. Unspecified hyperammonaemia 238970 1.2. Organic acidurias 1.2.1. Glutaric aciduria 1.2.1.1. Glutaric aciduria type I 231670 S Glutaryl-CoA dehydrogenase deficiency 1.2.1.2. Glutaric aciduria type III 231690 1.2.2. Propionic aciduria E711 232000 S Propionyl-CoA-Carboxylase deficiency 1.2.3. Methylmalonic aciduria E711 251000 1.2.3.1. Methylmalonyl-CoA mutase deficiency 1.2.3.2. Methylmalonyl-CoA epimerase deficiency 251120 1.2.3.3. Methylmalonic aciduria, unspecified 1.2.4. Isovaleric aciduria E711 243500 S Isovaleryl-CoA dehydrogenase deficiency 1.2.5. Methylcrotonylglycinuria E744 210200 S Methylcrotonyl-CoA carboxylase deficiency 1.2.6. Methylglutaconic aciduria E712 250950 1.2.6.1. Methylglutaconic aciduria type I E712 250950 S 3-Methylglutaconyl-CoA hydratase deficiency 1.2.6.2.