Using a Seed-Network to Query Multiple Large-Scale Gene
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

TRANSCRIPTIONAL REGULATION of Hur in RENAL STRESS
TRANSCRIPTIONAL REGULATION OF HuR IN RENAL STRESS DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Sudha Suman Govindaraju Graduate Program in Biochemistry The Ohio State University 2014 Dissertation Committee: Dr. Beth S. Lee, Ph.D., Advisor Dr. Kathleen Boris-Lawrie, Ph.D. Dr. Sissy M. Jhiang, Ph.D. Dr. Arthur R. Strauch, Ph.D Abstract HuR is a ubiquitously expressed RNA-binding protein that affects the post- transcriptional life of thousands of cellular mRNAs by regulating transcript stability and translation. HuR can post-transcriptionally regulate gene expression and modulate cellular responses to stress, differentiation, proliferation, apoptosis, senescence, inflammation, and the immune response. It is an important mediator of survival during cellular stress, but when inappropriately expressed, can promote oncogenic transformation. Not surprisingly, the expression of HuR itself is tightly regulated at multiple transcriptional and post-transcriptional levels. Previous studies demonstrated the existence of two alternate HuR transcripts that differ in their 5’ untranslated regions and have markedly different translatabilities. These forms were also found to be reciprocally expressed following cellular stress in kidney proximal tubule cell lines, and the shorter, more readily translatable variant was shown to be regulated by Smad 1/5/8 pathway and bone morphogenetic protein-7 (BMP-7) signaling. In this study, the factors that promote transcription of the longer alternate form were identified. NF-κB was shown to be important for expression of the long HuR mRNA, as was a newly identified region with potential for binding the Sp/KLF families of transcription factors. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Hypermethylation-Mediated Silencing of NDRG4 Promotes Pancreatic Ductal Adenocarcinoma by Regulating Mitochondrial Function
BMB Rep. 2020; 53(12): 658-663 BMB www.bmbreports.org Reports Hypermethylation-mediated silencing of NDRG4 promotes pancreatic ductal adenocarcinoma by regulating mitochondrial function Hao-Hong Shi1, Hai-E Liu1 & Xing-Jing Luo1,2,* 1Department of Anesthesia, Children’s Hospital of Fudan University, Shanghai 201102, 2Department of Anesthesia, Anhui Provincial Children’s Hospital, Hefei, Anhui 230022, China The N-myc downstream regulated gene (NDRG) family members are diagnosed in the advanced stage with a 5-year overall sur- are dysregulated in several tumors. Functionally, NDRGs play vival rate of less than 6%. Thus, PDAC is internationally called an important role in the malignant progression of cancer cells. the “king of cancer” (2). Unfortunately, even with the advent of However, little is known about the potential implications of diverse detection technologies and advanced treatment methods, NDRG4 in pancreatic ductal adenocarcinoma (PDAC). The aim the incidence and mortality rate of PDAC are still on the rise of the current study was to elucidate the expression pattern of year by year (3, 4). Due to the lack of in-depth understanding NDRG4 in PDAC and evaluate its potential cellular biological of molecular characteristics of malignant development, compre- effects. Here, we firstly report that epigenetic-mediated silencing hensive and in-depth research studies are urgently needed to of NDRG4 promotes PDAC by regulating mitochondrial func- better prevent and treat PDAC. tion. Data mining demonstrated that NDRG4 was significantly N-Myc downstream-regulated gene 4 (NDRG4) is a member down-regulated in PDAC tissues and cells. PDAC patients with of the NDRG family proteins (NDRG1-4) known to share 57%- low NDRG4 expression showed poor prognosis. -

Supplementary Table S5. Differentially Expressed Gene Lists of PD-1High CD39+ CD8 Tils According to 4-1BB Expression Compared to PD-1+ CD39- CD8 Tils
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Immunother Cancer Supplementary Table S5. Differentially expressed gene lists of PD-1high CD39+ CD8 TILs according to 4-1BB expression compared to PD-1+ CD39- CD8 TILs Up- or down- regulated genes in Up- or down- regulated genes Up- or down- regulated genes only PD-1high CD39+ CD8 TILs only in 4-1BBneg PD-1high CD39+ in 4-1BBpos PD-1high CD39+ CD8 compared to PD-1+ CD39- CD8 CD8 TILs compared to PD-1+ TILs compared to PD-1+ CD39- TILs CD39- CD8 TILs CD8 TILs IL7R KLRG1 TNFSF4 ENTPD1 DHRS3 LEF1 ITGA5 MKI67 PZP KLF3 RYR2 SIK1B ANK3 LYST PPP1R3B ETV1 ADAM28 H2AC13 CCR7 GFOD1 RASGRP2 ITGAX MAST4 RAD51AP1 MYO1E CLCF1 NEBL S1PR5 VCL MPP7 MS4A6A PHLDB1 GFPT2 TNF RPL3 SPRY4 VCAM1 B4GALT5 TIPARP TNS3 PDCD1 POLQ AKAP5 IL6ST LY9 PLXND1 PLEKHA1 NEU1 DGKH SPRY2 PLEKHG3 IKZF4 MTX3 PARK7 ATP8B4 SYT11 PTGER4 SORL1 RAB11FIP5 BRCA1 MAP4K3 NCR1 CCR4 S1PR1 PDE8A IFIT2 EPHA4 ARHGEF12 PAICS PELI2 LAT2 GPRASP1 TTN RPLP0 IL4I1 AUTS2 RPS3 CDCA3 NHS LONRF2 CDC42EP3 SLCO3A1 RRM2 ADAMTSL4 INPP5F ARHGAP31 ESCO2 ADRB2 CSF1 WDHD1 GOLIM4 CDK5RAP1 CD69 GLUL HJURP SHC4 GNLY TTC9 HELLS DPP4 IL23A PITPNC1 TOX ARHGEF9 EXO1 SLC4A4 CKAP4 CARMIL3 NHSL2 DZIP3 GINS1 FUT8 UBASH3B CDCA5 PDE7B SOGA1 CDC45 NR3C2 TRIB1 KIF14 TRAF5 LIMS1 PPP1R2C TNFRSF9 KLRC2 POLA1 CD80 ATP10D CDCA8 SETD7 IER2 PATL2 CCDC141 CD84 HSPA6 CYB561 MPHOSPH9 CLSPN KLRC1 PTMS SCML4 ZBTB10 CCL3 CA5B PIP5K1B WNT9A CCNH GEM IL18RAP GGH SARDH B3GNT7 C13orf46 SBF2 IKZF3 ZMAT1 TCF7 NECTIN1 H3C7 FOS PAG1 HECA SLC4A10 SLC35G2 PER1 P2RY1 NFKBIA WDR76 PLAUR KDM1A H1-5 TSHZ2 FAM102B HMMR GPR132 CCRL2 PARP8 A2M ST8SIA1 NUF2 IL5RA RBPMS UBE2T USP53 EEF1A1 PLAC8 LGR6 TMEM123 NEK2 SNAP47 PTGIS SH2B3 P2RY8 S100PBP PLEKHA7 CLNK CRIM1 MGAT5 YBX3 TP53INP1 DTL CFH FEZ1 MYB FRMD4B TSPAN5 STIL ITGA2 GOLGA6L10 MYBL2 AHI1 CAND2 GZMB RBPJ PELI1 HSPA1B KCNK5 GOLGA6L9 TICRR TPRG1 UBE2C AURKA Leem G, et al. -

Neuronal Ndrg4 Is Essential for Nodes of Ranvier Organization in Zebrafish
RESEARCH ARTICLE Neuronal Ndrg4 Is Essential for Nodes of Ranvier Organization in Zebrafish Laura Fontenas1¤a, Flavia De Santis2☯, Vincenzo Di Donato2☯, Cindy Degerny1, BeÂatrice Chambraud1, Filippo Del Bene2, Marcel Tawk1* 1 U1195, Inserm, University Paris Sud, University Paris-Saclay, Kremlin-Bicêtre, France, 2 Institut Curie, PSL Research University, Paris, France ☯ These authors contributed equally to this work. ¤a Current address: Department of Biology, University of Virginia, Charlottesville, Virginia, United States of America * [email protected] a11111 Abstract Axon ensheathment by specialized glial cells is an important process for fast propagation of action potentials. The rapid electrical conduction along myelinated axons is mainly due to its saltatory nature characterized by the accumulation of ion channels at the nodes of Ranvier. However, how these ion channels are transported and anchored along axons is not fully OPEN ACCESS understood. We have identified N-myc downstream-regulated gene 4, ndrg4, as a novel fac- Citation: Fontenas L, De Santis F, Di Donato V, tor that regulates sodium channel clustering in zebrafish. Analysis of chimeric larvae indi- Degerny C, Chambraud B, Del Bene F, et al. (2016) Neuronal Ndrg4 Is Essential for Nodes of Ranvier cates that ndrg4 functions autonomously within neurons for sodium channel clustering at Organization in Zebrafish. PLoS Genet 12(11): the nodes. Molecular analysis of ndrg4 mutants shows that expression of snap25 and nsf e1006459. doi:10.1371/journal.pgen.1006459 are sharply decreased, revealing a role of ndrg4 in controlling vesicle exocytosis. This Editor: David Lyons, Centre for Neuroregeneration, uncovers a previously unknown function of ndrg4 in regulating vesicle docking and nodes of Edinburgh, UNITED KINGDOM Ranvier organization, at least through its ability to finely tune the expression of the t-SNARE/ Received: May 3, 2016 NSF machinery. -

Supplementary Table 1. Pain and PTSS Associated Genes (N = 604
Supplementary Table 1. Pain and PTSS associated genes (n = 604) compiled from three established pain gene databases (PainNetworks,[61] Algynomics,[52] and PainGenes[42]) and one PTSS gene database (PTSDgene[88]). These genes were used in in silico analyses aimed at identifying miRNA that are predicted to preferentially target this list genes vs. a random set of genes (of the same length). ABCC4 ACE2 ACHE ACPP ACSL1 ADAM11 ADAMTS5 ADCY5 ADCYAP1 ADCYAP1R1 ADM ADORA2A ADORA2B ADRA1A ADRA1B ADRA1D ADRA2A ADRA2C ADRB1 ADRB2 ADRB3 ADRBK1 ADRBK2 AGTR2 ALOX12 ANO1 ANO3 APOE APP AQP1 AQP4 ARL5B ARRB1 ARRB2 ASIC1 ASIC2 ATF1 ATF3 ATF6B ATP1A1 ATP1B3 ATP2B1 ATP6V1A ATP6V1B2 ATP6V1G2 AVPR1A AVPR2 BACE1 BAMBI BDKRB2 BDNF BHLHE22 BTG2 CA8 CACNA1A CACNA1B CACNA1C CACNA1E CACNA1G CACNA1H CACNA2D1 CACNA2D2 CACNA2D3 CACNB3 CACNG2 CALB1 CALCRL CALM2 CAMK2A CAMK2B CAMK4 CAT CCK CCKAR CCKBR CCL2 CCL3 CCL4 CCR1 CCR7 CD274 CD38 CD4 CD40 CDH11 CDK5 CDK5R1 CDKN1A CHRM1 CHRM2 CHRM3 CHRM5 CHRNA5 CHRNA7 CHRNB2 CHRNB4 CHUK CLCN6 CLOCK CNGA3 CNR1 COL11A2 COL9A1 COMT COQ10A CPN1 CPS1 CREB1 CRH CRHBP CRHR1 CRHR2 CRIP2 CRYAA CSF2 CSF2RB CSK CSMD1 CSNK1A1 CSNK1E CTSB CTSS CX3CL1 CXCL5 CXCR3 CXCR4 CYBB CYP19A1 CYP2D6 CYP3A4 DAB1 DAO DBH DBI DICER1 DISC1 DLG2 DLG4 DPCR1 DPP4 DRD1 DRD2 DRD3 DRD4 DRGX DTNBP1 DUSP6 ECE2 EDN1 EDNRA EDNRB EFNB1 EFNB2 EGF EGFR EGR1 EGR3 ENPP2 EPB41L2 EPHB1 EPHB2 EPHB3 EPHB4 EPHB6 EPHX2 ERBB2 ERBB4 EREG ESR1 ESR2 ETV1 EZR F2R F2RL1 F2RL2 FAAH FAM19A4 FGF2 FKBP5 FLOT1 FMR1 FOS FOSB FOSL2 FOXN1 FRMPD4 FSTL1 FYN GABARAPL1 GABBR1 GABBR2 GABRA2 GABRA4 -

Transcriptome Analysis of Complex I-Deficient Patients Reveals Distinct
van der Lee et al. BMC Genomics (2015) 16:691 DOI 10.1186/s12864-015-1883-8 RESEARCH ARTICLE Open Access Transcriptome analysis of complex I-deficient patients reveals distinct expression programs for subunits and assembly factors of the oxidative phosphorylation system Robin van der Lee1†, Radek Szklarczyk1,2†, Jan Smeitink3,HubertJMSmeets4, Martijn A. Huynen1 and Rutger Vogel3* Abstract Background: Transcriptional control of mitochondrial metabolism is essential for cellular function. A better understanding of this process will aid the elucidation of mitochondrial disorders, in particular of the many genetically unsolved cases of oxidative phosphorylation (OXPHOS) deficiency. Yet, to date only few studies have investigated nuclear gene regulation in the context of OXPHOS deficiency. In this study we performed RNA sequencing of two control and two complex I-deficient patient cell lines cultured in the presence of compounds that perturb mitochondrial metabolism: chloramphenicol, AICAR, or resveratrol. We combined this with a comprehensive analysis of mitochondrial and nuclear gene expression patterns, co-expression calculations and transcription factor binding sites. Results: Our analyses show that subsets of mitochondrial OXPHOS genes respond opposingly to chloramphenicol and AICAR, whereas the response of nuclear OXPHOS genes is less consistent between cell lines and treatments. Across all samples nuclear OXPHOS genes have a significantly higher co-expression with each other than with other genes, including those encoding mitochondrial proteins. We found no evidence for complex-specific mRNA expression regulation: subunits of different OXPHOS complexes are similarly (co-)expressed and regulated by a common set of transcription factors. However, we did observe significant differences between the expression of nuclear genes for OXPHOS subunits versus assembly factors, suggesting divergent transcription programs. -

Whole Exome Sequencing in Families at High Risk for Hodgkin Lymphoma: Identification of a Predisposing Mutation in the KDR Gene
Hodgkin Lymphoma SUPPLEMENTARY APPENDIX Whole exome sequencing in families at high risk for Hodgkin lymphoma: identification of a predisposing mutation in the KDR gene Melissa Rotunno, 1 Mary L. McMaster, 1 Joseph Boland, 2 Sara Bass, 2 Xijun Zhang, 2 Laurie Burdett, 2 Belynda Hicks, 2 Sarangan Ravichandran, 3 Brian T. Luke, 3 Meredith Yeager, 2 Laura Fontaine, 4 Paula L. Hyland, 1 Alisa M. Goldstein, 1 NCI DCEG Cancer Sequencing Working Group, NCI DCEG Cancer Genomics Research Laboratory, Stephen J. Chanock, 5 Neil E. Caporaso, 1 Margaret A. Tucker, 6 and Lynn R. Goldin 1 1Genetic Epidemiology Branch, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; 2Cancer Genomics Research Laboratory, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; 3Ad - vanced Biomedical Computing Center, Leidos Biomedical Research Inc.; Frederick National Laboratory for Cancer Research, Frederick, MD; 4Westat, Inc., Rockville MD; 5Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; and 6Human Genetics Program, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD, USA ©2016 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol.2015.135475 Received: August 19, 2015. Accepted: January 7, 2016. Pre-published: June 13, 2016. Correspondence: [email protected] Supplemental Author Information: NCI DCEG Cancer Sequencing Working Group: Mark H. Greene, Allan Hildesheim, Nan Hu, Maria Theresa Landi, Jennifer Loud, Phuong Mai, Lisa Mirabello, Lindsay Morton, Dilys Parry, Anand Pathak, Douglas R. Stewart, Philip R. Taylor, Geoffrey S. Tobias, Xiaohong R. Yang, Guoqin Yu NCI DCEG Cancer Genomics Research Laboratory: Salma Chowdhury, Michael Cullen, Casey Dagnall, Herbert Higson, Amy A. -

Fabbri Et Al. Whole Genome Analysis and Micrornas Regulation in Hepg2 Cells Exposed to Cadmium Supplementary Data
Fabbri et al. Whole Genome Analysis and MicroRNAs Regulation in HepG2 Cells Exposed to Cadmium Supplementary Data Tab. S1: KEGG enrichment for downregulated genes Genes identified in Figure 1 were analyzed by DAVID for associations with particular KEGG pathways. KEGG Entry is KEGG identifier, Name is name of the KEGG pathway, Genes shows the number of genes associated with the specific pathway, the PValue refers to how significant an association a particular KEGG pathway has with the gene list. KEGG Entry Name Genes PValue hsa04610 Complement and coagulation cascades 22 1.11E-14 hsa00260 Glycine, serine and threonine metabolism 11 8.50E-08 hsa00071 Fatty acid metabolism 11 9.41E-07 hsa00650 Butanoate metabolism 9 1.89E-05 hsa00100 Steroid biosynthesis 7 2.09E-05 hsa00280 Valine, leucine and isoleucine degradation 10 2.47E-05 hsa00380 Tryptophan metabolism 9 8.40E-05 hsa00330 Arginine and proline metabolism 10 1.16E-04 hsa00900 Terpenoid backbone biosynthesis 6 1.46E-04 hsa00980 Metabolism of xenobiotics by cytochrome P450 10 2.71E-04 hsa00010 Glycolysis / Gluconeogenesis 10 2.71E-04 hsa00982 Drug metabolism 10 3.98E-04 hsa03320 PPAR signaling pathway 10 7.98E-04 hsa00620 Pyruvate metabolism 7 0.003185725 hsa00561 Glycerolipid metabolism 7 0.005184764 hsa00640 Propanoate metabolism 6 0.005876295 hsa00910 Nitrogen metabolism 5 0.009266837 hsa00480 Glutathione metabolism 7 0.009722623 hsa04950 Maturity onset diabetes of the young 5 0.012498995 hsa00903 Limonene and pinene degradation 4 0.013441968 hsa00680 Methane metabolism 3 0.018538005 hsa00120 Primary bile acid biosynthesis 4 0.01958794 hsa00340 Histidine metabolism 5 0.020928876 hsa00310 Lysine degradation 6 0.022199526 hsa00250 Alanine, aspartate and glutamate metabolism 5 0.026189764 hsa00410 beta-Alanine metabolism 4 0.04583419 hsa01040 Biosynthesis of unsaturated fatty acids 4 0.04583419 ALTEX, 2/12 SUPPL., 1 FABBRI ET AL . -
The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas
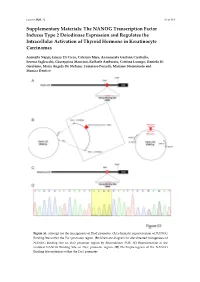
Cancers 2020, 12 S1 of S18 Supplementary Materials: The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas Annarita Nappi, Emery Di Cicco, Caterina Miro, Annunziata Gaetana Cicatiello, Serena Sagliocchi, Giuseppina Mancino, Raffaele Ambrosio, Cristina Luongo, Daniela Di Girolamo, Maria Angela De Stefano, Tommaso Porcelli, Mariano Stornaiuolo and Monica Dentice Figure S1. Strategy for the mutagenesis of Dio2 promoter. (A) Schematic representation of NANOG Binding Site within the Dio2 promoter region. (B) Schematic diagram for site‐directed mutagenesis of NANOG Binding Site on Dio2 promoter region by Recombinant PCR. (C) Representation of the mutated NANOG Binding Site on Dio2 promoter region. (D) Electropherogram of the NANOG Binding Site mutation within the Dio2 promoter. Cancers 2020, 12 S2 of S18 Figure S2. Strategy for the silencing of NANOG expression. (A) Cloning strategies for the generation of NANOG shRNA expression vectors. (B) Electropherograms of the NANOG shRNA sequences cloned into pcDNA3.1 vector. (C) Validation of effective NANOG down-modulation by two different NANOG shRNA vectors was assessed by Western Blot analysis of NANOG expression in BCC cells. (D) Quantification of NANOG protein levels versus Tubulin levels in the same experiment as in C is represented by histograms. Cancers 2020, 12 S3 of S18 Figure S3. The CD34+ cells are characterized by the expression of typical epithelial stemness genes. The mRNA levels of a panel of indicated stemness markers of epidermis were measured by Real Time PCR in the same experiment indicated in figure 3F and G. Cancers 2020, 12 S4 of S18 Figure S4. -

Evolving Principles Underlying Neural Lineage Conversion
F1000Research 2019, 8(F1000 Faculty Rev):1548 Last updated: 16 SEP 2019 REVIEW Evolving principles underlying neural lineage conversion and their relevance for biomedical translation [version 1; peer review: 4 approved] Lea Jessica Flitsch, Oliver Brüstle Institute of Reconstructive Neurobiology, University of Bonn School of Medicine & University Hospital Bonn, Bonn, North Rhine Wesphalia, 53127, Germany First published: 30 Aug 2019, 8(F1000 Faculty Rev):1548 ( Open Peer Review v1 https://doi.org/10.12688/f1000research.18926.1) Latest published: 30 Aug 2019, 8(F1000 Faculty Rev):1548 ( https://doi.org/10.12688/f1000research.18926.1) Reviewer Status Abstract Invited Reviewers Scientific and technological advances of the past decade have shed light 1 2 3 4 on the mechanisms underlying cell fate acquisition, including its transcriptional and epigenetic regulation during embryonic development. version 1 This knowledge has enabled us to purposefully engineer cell fates in vitro published by manipulating expression levels of lineage-instructing transcription 30 Aug 2019 factors. Here, we review the state of the art in the cell programming field with a focus on the derivation of neural cells. We reflect on what we know about the mechanisms underlying fate changes in general and on the F1000 Faculty Reviews are written by members of degree of epigenetic remodeling conveyed by the distinct reprogramming the prestigious F1000 Faculty. They are and direct conversion strategies available. Moreover, we discuss the commissioned and are peer reviewed before implications of residual epigenetic memory for biomedical applications such publication to ensure that the final, published version as disease modeling and neuroregeneration. Finally, we cover recent developments approaching cell fate conversion in the living brain and is comprehensive and accessible. -

A Missense Mutation in the KLF7 Gene Is a Potential Candidate Variant for Congenital Deafness in Australian Stumpy Tail Cattle Dogs
G C A T T A C G G C A T genes Article A Missense Mutation in the KLF7 Gene Is a Potential Candidate Variant for Congenital Deafness in Australian Stumpy Tail Cattle Dogs Fangzheng Xu 1,† , Shuwen Shan 1,†, Susan Sommerlad 2, Jennifer M. Seddon 2 and Bertram Brenig 1,* 1 Institute of Veterinary Medicine, University of Goettingen, 37077 Göttingen, Germany; [email protected] (F.X.); [email protected] (S.S.) 2 School of Veterinary Science, The University of Queensland, Gatton, QLD 4343, Australia; [email protected] (S.S.); [email protected] (J.M.S.) * Correspondence: [email protected]; Tel.: +49-551-3928383; Fax: +49-551-3933392 † These authors have equally contributed. Abstract: Congenital deafness is prevalent among modern dog breeds, including Australian Stumpy Tail Cattle Dogs (ASCD). However, in ASCD, no causative gene has been identified so far. Therefore, we performed a genome-wide association study (GWAS) and whole genome sequencing (WGS) of affected and normal individuals. For GWAS, 3 bilateral deaf ASCDs, 43 herding dogs, and one unaffected ASCD were used, resulting in 13 significantly associated loci on 6 chromosomes, i.e., CFA3, 8, 17, 23, 28, and 37. CFA37 harbored a region with the most significant association −21 (−log10(9.54 × 10 ) = 20.02) as well as 7 of the 13 associated loci. For whole genome sequencing, the same three affected ASCDs and one unaffected ASCD were used. The WGS data were compared with 722 canine controls and filtered for protein coding and non-synonymous variants, resulting in Citation: Xu, F.; Shan, S.; Sommerlad, four missense variants present only in the affected dogs.