Lichenized Fungi and the Evolution of Symbiotic Organization
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
Cuivre Bryophytes
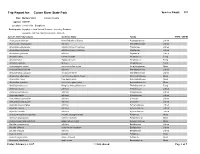
Trip Report for: Cuivre River State Park Species Count: 335 Date: Multiple Visits Lincoln County Agency: MODNR Location: Lincoln Hills - Bryophytes Participants: Bryophytes from Natural Resource Inventory Database Bryophyte List from NRIDS and Bruce Schuette Species Name (Synonym) Common Name Family COFC COFW Acarospora unknown Identified only to Genus Acarosporaceae Lichen Acrocordia megalospora a lichen Monoblastiaceae Lichen Amandinea dakotensis a button lichen (crustose) Physiaceae Lichen Amandinea polyspora a button lichen (crustose) Physiaceae Lichen Amandinea punctata a lichen Physiaceae Lichen Amanita citrina Citron Amanita Amanitaceae Fungi Amanita fulva Tawny Gresette Amanitaceae Fungi Amanita vaginata Grisette Amanitaceae Fungi Amblystegium varium common willow moss Amblystegiaceae Moss Anisomeridium biforme a lichen Monoblastiaceae Lichen Anisomeridium polypori a crustose lichen Monoblastiaceae Lichen Anomodon attenuatus common tree apron moss Anomodontaceae Moss Anomodon minor tree apron moss Anomodontaceae Moss Anomodon rostratus velvet tree apron moss Anomodontaceae Moss Armillaria tabescens Ringless Honey Mushroom Tricholomataceae Fungi Arthonia caesia a lichen Arthoniaceae Lichen Arthonia punctiformis a lichen Arthoniaceae Lichen Arthonia rubella a lichen Arthoniaceae Lichen Arthothelium spectabile a lichen Uncertain Lichen Arthothelium taediosum a lichen Uncertain Lichen Aspicilia caesiocinerea a lichen Hymeneliaceae Lichen Aspicilia cinerea a lichen Hymeneliaceae Lichen Aspicilia contorta a lichen Hymeneliaceae Lichen -

Opuscula Philolichenum, 6: 1-XXXX
Opuscula Philolichenum, 15: 56-81. 2016. *pdf effectively published online 25July2016 via (http://sweetgum.nybg.org/philolichenum/) Lichens, lichenicolous fungi, and allied fungi of Pipestone National Monument, Minnesota, U.S.A., revisited M.K. ADVAITA, CALEB A. MORSE1,2 AND DOUGLAS LADD3 ABSTRACT. – A total of 154 lichens, four lichenicolous fungi, and one allied fungus were collected by the authors from 2004 to 2015 from Pipestone National Monument (PNM), in Pipestone County, on the Prairie Coteau of southwestern Minnesota. Twelve additional species collected by previous researchers, but not found by the authors, bring the total number of taxa known for PNM to 171. This represents a substantial increase over previous reports for PNM, likely due to increased intensity of field work, and also to the marked expansion of corticolous and anthropogenic substrates since the site was first surveyed in 1899. Reexamination of 116 vouchers deposited in MIN and the PNM herbarium led to the exclusion of 48 species previously reported from the site. Crustose lichens are the most common growth form, comprising 65% of the lichen diversity. Sioux Quartzite provided substrate for 43% of the lichen taxa collected. Saxicolous lichen communities were characterized by sampling four transects on cliff faces and low outcrops. An annotated checklist of the lichens of the site is provided, as well as a list of excluded taxa. We report 24 species (including 22 lichens and two lichenicolous fungi) new for Minnesota: Acarospora boulderensis, A. contigua, A. erythrophora, A. strigata, Agonimia opuntiella, Arthonia clemens, A. muscigena, Aspicilia americana, Bacidina delicata, Buellia tyrolensis, Caloplaca flavocitrina, C. lobulata, C. -

Aportes Al Conocimiento De La Biota Liquénica Del Oasis De Neblina De Alto Patache, Desierto De Atacama1
Revista de Geografía Norte Grande, 68: 49-64 (2017) Artículos Aportes al conocimiento de la biota liquénica del oasis de neblina de Alto Patache, Desierto de Atacama1 Reinaldo Vargas Castillo2, Daniel Stanton3 y Peter R. Nelson4 RESUMEN Los denominados oasis de neblina son áreas en las zonas costeras del Desierto de Ataca- ma donde el ingreso habitual de niebla permite el establecimiento y desarrollo de diver- sas poblaciones de plantas vasculares, generando verdaderos hotspots de diversidad. En estas áreas, la biota liquenológica ha sido poco explorada y representa uno de los ele- mentos perennes más importantes que conforman la comunidad. En un estudio previo de la biota del oasis de neblina de Alto Patache se reportaron siete especies. Con el fin de mejorar este conocimiento, se analizó la riqueza de especies presentes en el oasis si- guiendo dos transectos altitudinales en diferentes orientaciones del farellón. Aquí repor- tamos preliminarmente 77 especies de líquenes para el oasis de neblina de Alto Patache. De estas, 61 especies corresponden a nuevos registros para la región de Tarapacá, en tanto que las especies Amandinea eff lorescens, Diploicia canescens, Myriospora smarag- dula y Rhizocarpon simillimum corresponden a nuevos registros para el país. Asimismo, se destaca a Alto Patache como la única localidad conocida para Santessonia cervicornis, una especie endémica y en Peligro Crítico. Palabras clave: Oasis de neblina, Desierto de Atacama, líquenes. ABSTRACT Fog oases are zones along the Atacama Desert where the regular input of fog favors the development of rich communities of vascular plants, becoming biodiversity hotspots. In these areas, the lichen biota has been poorly explored and represents one of the most conspicuous elements among the perennials organisms that form the community. -

Foliicolous Lichens and Their Lichenicolous Fungi Collected During the Smithsonian International Cryptogamic Expedition to Guyana 1996
45 Tropical Bryology 15: 45-76, 1998 Foliicolous lichens and their lichenicolous fungi collected during the Smithsonian International Cryptogamic Expedition to Guyana 1996 Robert Lücking Lehrstuhl für Pflanzensystematik, Universität Bayreuth, D-95447 Bayreuth, Germany Abstract: A total of 233 foliicolous lichen species and 18 lichenicolous fungi are reported from Guyana as a result of the Smithsonian „International Cryptogamic Expedition to Guyana“ 1996. Three lichens and two lichenicolous fungi are new to science: Arthonia grubei sp.n., Badimia subelegans sp.n., Calopadia pauciseptata sp.n., Opegrapha matzeri sp.n. (lichenicolous on Amazonomyces sprucei), and Pyrenidium santessonii sp.n. (lichenicolous on Bacidia psychotriae). The new combination Strigula janeirensis (Bas.: Phylloporina janeirensis; syn.: Raciborskiella janeirensis) is proposed. Apart from Amazonomyces sprucei and Bacidia psychotriae, Arthonia lecythidicola (with the lichenicolous A. pseudopegraphina) and Byssolecania deplanata (with the lichenicolous Opegrapha cf. kalbii) are reported as new hosts for lichenicolous fungi. Arthonia pseudopegraphina growing on A. lecythidicola is the first known case of adelphoparasitism at generic level in foliicolous Arthonia. Arthonia flavoverrucosa, Badimia polillensis, and Byssoloma vezdanum are new records for the Neotropics, and 115 species are new for Guyana, resulting in a total of c. 280 genuine foliicolous species reported for that country, while Porina applanata and P. verruculosa are excluded from its flora. The foliicolous lichen flora of Guyana is representative for the Guianas (Guyana, Suriname, French Guiana) and has great affinities with the Amazon region, while the degree of endemism is low. A characteristic species for this area is Amazonomyces sprucei. Species composition is typical of Neotropical lowland to submontane humid forests, with a dominance of the genera Porina, Strigula, and Mazosia. -

Mycosphere Notes 225–274: Types and Other Specimens of Some Genera of Ascomycota
Mycosphere 9(4): 647–754 (2018) www.mycosphere.org ISSN 2077 7019 Article Doi 10.5943/mycosphere/9/4/3 Copyright © Guizhou Academy of Agricultural Sciences Mycosphere Notes 225–274: types and other specimens of some genera of Ascomycota Doilom M1,2,3, Hyde KD2,3,6, Phookamsak R1,2,3, Dai DQ4,, Tang LZ4,14, Hongsanan S5, Chomnunti P6, Boonmee S6, Dayarathne MC6, Li WJ6, Thambugala KM6, Perera RH 6, Daranagama DA6,13, Norphanphoun C6, Konta S6, Dong W6,7, Ertz D8,9, Phillips AJL10, McKenzie EHC11, Vinit K6,7, Ariyawansa HA12, Jones EBG7, Mortimer PE2, Xu JC2,3, Promputtha I1 1 Department of Biology, Faculty of Science, Chiang Mai University, Chiang Mai 50200, Thailand 2 Key Laboratory for Plant Diversity and Biogeography of East Asia, Kunming Institute of Botany, Chinese Academy of Sciences, 132 Lanhei Road, Kunming 650201, China 3 World Agro Forestry Centre, East and Central Asia, 132 Lanhei Road, Kunming 650201, Yunnan Province, People’s Republic of China 4 Center for Yunnan Plateau Biological Resources Protection and Utilization, College of Biological Resource and Food Engineering, Qujing Normal University, Qujing, Yunnan 655011, China 5 Shenzhen Key Laboratory of Microbial Genetic Engineering, College of Life Sciences and Oceanography, Shenzhen University, Shenzhen 518060, China 6 Center of Excellence in Fungal Research, Mae Fah Luang University, Chiang Rai 57100, Thailand 7 Department of Entomology and Plant Pathology, Faculty of Agriculture, Chiang Mai University, Chiang Mai 50200, Thailand 8 Department Research (BT), Botanic Garden Meise, Nieuwelaan 38, BE-1860 Meise, Belgium 9 Direction Générale de l'Enseignement non obligatoire et de la Recherche scientifique, Fédération Wallonie-Bruxelles, Rue A. -

An Evolving Phylogenetically Based Taxonomy of Lichens and Allied Fungi
Opuscula Philolichenum, 11: 4-10. 2012. *pdf available online 3January2012 via (http://sweetgum.nybg.org/philolichenum/) An evolving phylogenetically based taxonomy of lichens and allied fungi 1 BRENDAN P. HODKINSON ABSTRACT. – A taxonomic scheme for lichens and allied fungi that synthesizes scientific knowledge from a variety of sources is presented. The system put forth here is intended both (1) to provide a skeletal outline of the lichens and allied fungi that can be used as a provisional filing and databasing scheme by lichen herbarium/data managers and (2) to announce the online presence of an official taxonomy that will define the scope of the newly formed International Committee for the Nomenclature of Lichens and Allied Fungi (ICNLAF). The online version of the taxonomy presented here will continue to evolve along with our understanding of the organisms. Additionally, the subfamily Fissurinoideae Rivas Plata, Lücking and Lumbsch is elevated to the rank of family as Fissurinaceae. KEYWORDS. – higher-level taxonomy, lichen-forming fungi, lichenized fungi, phylogeny INTRODUCTION Traditionally, lichen herbaria have been arranged alphabetically, a scheme that stands in stark contrast to the phylogenetic scheme used by nearly all vascular plant herbaria. The justification typically given for this practice is that lichen taxonomy is too unstable to establish a reasonable system of classification. However, recent leaps forward in our understanding of the higher-level classification of fungi, driven primarily by the NSF-funded Assembling the Fungal Tree of Life (AFToL) project (Lutzoni et al. 2004), have caused the taxonomy of lichen-forming and allied fungi to increase significantly in stability. This is especially true within the class Lecanoromycetes, the main group of lichen-forming fungi (Miadlikowska et al. -

The Fungi of Slapton Ley National Nature Reserve and Environs
THE FUNGI OF SLAPTON LEY NATIONAL NATURE RESERVE AND ENVIRONS APRIL 2019 Image © Visit South Devon ASCOMYCOTA Order Family Name Abrothallales Abrothallaceae Abrothallus microspermus CY (IMI 164972 p.p., 296950), DM (IMI 279667, 279668, 362458), N4 (IMI 251260), Wood (IMI 400386), on thalli of Parmelia caperata and P. perlata. Mainly as the anamorph <it Abrothallus parmeliarum C, CY (IMI 164972), DM (IMI 159809, 159865), F1 (IMI 159892), 2, G2, H, I1 (IMI 188770), J2, N4 (IMI 166730), SV, on thalli of Parmelia carporrhizans, P Abrothallus parmotrematis DM, on Parmelia perlata, 1990, D.L. Hawksworth (IMI 400397, as Vouauxiomyces sp.) Abrothallus suecicus DM (IMI 194098); on apothecia of Ramalina fustigiata with st. conid. Phoma ranalinae Nordin; rare. (L2) Abrothallus usneae (as A. parmeliarum p.p.; L2) Acarosporales Acarosporaceae Acarospora fuscata H, on siliceous slabs (L1); CH, 1996, T. Chester. Polysporina simplex CH, 1996, T. Chester. Sarcogyne regularis CH, 1996, T. Chester; N4, on concrete posts; very rare (L1). Trimmatothelopsis B (IMI 152818), on granite memorial (L1) [EXTINCT] smaragdula Acrospermales Acrospermaceae Acrospermum compressum DM (IMI 194111), I1, S (IMI 18286a), on dead Urtica stems (L2); CY, on Urtica dioica stem, 1995, JLT. Acrospermum graminum I1, on Phragmites debris, 1990, M. Marsden (K). Amphisphaeriales Amphisphaeriaceae Beltraniella pirozynskii D1 (IMI 362071a), on Quercus ilex. Ceratosporium fuscescens I1 (IMI 188771c); J1 (IMI 362085), on dead Ulex stems. (L2) Ceriophora palustris F2 (IMI 186857); on dead Carex puniculata leaves. (L2) Lepteutypa cupressi SV (IMI 184280); on dying Thuja leaves. (L2) Monographella cucumerina (IMI 362759), on Myriophyllum spicatum; DM (IMI 192452); isol. ex vole dung. (L2); (IMI 360147, 360148, 361543, 361544, 361546). -

Lichens and Associated Fungi from Glacier Bay National Park, Alaska
The Lichenologist (2020), 52,61–181 doi:10.1017/S0024282920000079 Standard Paper Lichens and associated fungi from Glacier Bay National Park, Alaska Toby Spribille1,2,3 , Alan M. Fryday4 , Sergio Pérez-Ortega5 , Måns Svensson6, Tor Tønsberg7, Stefan Ekman6 , Håkon Holien8,9, Philipp Resl10 , Kevin Schneider11, Edith Stabentheiner2, Holger Thüs12,13 , Jan Vondrák14,15 and Lewis Sharman16 1Department of Biological Sciences, CW405, University of Alberta, Edmonton, Alberta T6G 2R3, Canada; 2Department of Plant Sciences, Institute of Biology, University of Graz, NAWI Graz, Holteigasse 6, 8010 Graz, Austria; 3Division of Biological Sciences, University of Montana, 32 Campus Drive, Missoula, Montana 59812, USA; 4Herbarium, Department of Plant Biology, Michigan State University, East Lansing, Michigan 48824, USA; 5Real Jardín Botánico (CSIC), Departamento de Micología, Calle Claudio Moyano 1, E-28014 Madrid, Spain; 6Museum of Evolution, Uppsala University, Norbyvägen 16, SE-75236 Uppsala, Sweden; 7Department of Natural History, University Museum of Bergen Allégt. 41, P.O. Box 7800, N-5020 Bergen, Norway; 8Faculty of Bioscience and Aquaculture, Nord University, Box 2501, NO-7729 Steinkjer, Norway; 9NTNU University Museum, Norwegian University of Science and Technology, NO-7491 Trondheim, Norway; 10Faculty of Biology, Department I, Systematic Botany and Mycology, University of Munich (LMU), Menzinger Straße 67, 80638 München, Germany; 11Institute of Biodiversity, Animal Health and Comparative Medicine, College of Medical, Veterinary and Life Sciences, University of Glasgow, Glasgow G12 8QQ, UK; 12Botany Department, State Museum of Natural History Stuttgart, Rosenstein 1, 70191 Stuttgart, Germany; 13Natural History Museum, Cromwell Road, London SW7 5BD, UK; 14Institute of Botany of the Czech Academy of Sciences, Zámek 1, 252 43 Průhonice, Czech Republic; 15Department of Botany, Faculty of Science, University of South Bohemia, Branišovská 1760, CZ-370 05 České Budějovice, Czech Republic and 16Glacier Bay National Park & Preserve, P.O. -

Checklist of the Lichens and Allied Fungi of Kathy Stiles Freeland Bibb County Glades Preserve, Alabama, U.S.A
Opuscula Philolichenum, 18: 420–434. 2019. *pdf effectively published online 2December2019 via (http://sweetgum.nybg.org/philolichenum/) Checklist of the lichens and allied fungi of Kathy Stiles Freeland Bibb County Glades Preserve, Alabama, U.S.A. J. KEVIN ENGLAND1, CURTIS J. HANSEN2, JESSICA L. ALLEN3, SEAN Q. BEECHING4, WILLIAM R. BUCK5, VITALY CHARNY6, JOHN G. GUCCION7, RICHARD C. HARRIS8, MALCOLM HODGES9, NATALIE M. HOWE10, JAMES C. LENDEMER11, R. TROY MCMULLIN12, ERIN A. TRIPP13, DENNIS P. WATERS14 ABSTRACT. – The first checklist of lichenized, lichenicolous and lichen-allied fungi from the Kathy Stiles Freeland Bibb County Glades Preserve in Bibb County, Alabama, is presented. Collections made during the 2017 Tuckerman Workshop and additional records from herbaria and online sources are included. Two hundred and thirty-eight taxa in 115 genera are enumerated. Thirty taxa of lichenized, lichenicolous and lichen-allied fungi are newly reported for Alabama: Acarospora fuscata, A. novomexicana, Circinaria contorta, Constrictolumina cinchonae, Dermatocarpon dolomiticum, Didymocyrtis cladoniicola, Graphis anfractuosa, G. rimulosa, Hertelidea pseudobotryosa, Heterodermia pseudospeciosa, Lecania cuprea, Marchandiomyces lignicola, Minutoexcipula miniatoexcipula, Monoblastia rappii, Multiclavula mucida, Ochrolechia trochophora, Parmotrema subsumptum, Phaeographis brasiliensis, Phaeographis inusta, Piccolia nannaria, Placynthiella icmalea, Porina scabrida, Psora decipiens, Pyrenographa irregularis, Ramboldia blochiana, Thyrea confusa, Trichothelium -

Lichens and Allied Fungi of the Indiana Forest Alliance
2017. Proceedings of the Indiana Academy of Science 126(2):129–152 LICHENS AND ALLIED FUNGI OF THE INDIANA FOREST ALLIANCE ECOBLITZ AREA, BROWN AND MONROE COUNTIES, INDIANA INCORPORATED INTO A REVISED CHECKLIST FOR THE STATE OF INDIANA James C. Lendemer: Institute of Systematic Botany, The New York Botanical Garden, Bronx, NY 10458-5126 USA ABSTRACT. Based upon voucher collections, 108 lichen species are reported from the Indiana Forest Alliance Ecoblitz area, a 900 acre unit in Morgan-Monroe and Yellowwood State Forests, Brown and Monroe Counties, Indiana. The lichen biota of the study area was characterized as: i) dominated by species with green coccoid photobionts (80% of taxa); ii) comprised of 49% species that reproduce primarily with lichenized diaspores vs. 44% that reproduce primarily through sexual ascospores; iii) comprised of 65% crustose taxa, 29% foliose taxa, and 6% fruticose taxa; iv) one wherein many species are rare (e.g., 55% of species were collected fewer than three times) and fruticose lichens other than Cladonia were entirely absent; and v) one wherein cyanolichens were poorly represented, comprising only three species. Taxonomic diversity ranged from 21 to 56 species per site, with the lowest diversity sites concentrated in riparian corridors and the highest diversity sites on ridges. Low Gap Nature Preserve, located within the study area, was found to have comparable species richness to areas outside the nature preserve, although many species rare in the study area were found only outside preserve boundaries. Sets of rare species are delimited and discussed, as are observations as to the overall low abundance of lichens on corticolous substrates and the presence of many unhealthy foliose lichens on mature tree boles. -

A Multigene Phylogenetic Synthesis for the Class Lecanoromycetes (Ascomycota): 1307 Fungi Representing 1139 Infrageneric Taxa, 317 Genera and 66 Families
A multigene phylogenetic synthesis for the class Lecanoromycetes (Ascomycota): 1307 fungi representing 1139 infrageneric taxa, 317 genera and 66 families Miadlikowska, J., Kauff, F., Högnabba, F., Oliver, J. C., Molnár, K., Fraker, E., ... & Stenroos, S. (2014). A multigene phylogenetic synthesis for the class Lecanoromycetes (Ascomycota): 1307 fungi representing 1139 infrageneric taxa, 317 genera and 66 families. Molecular Phylogenetics and Evolution, 79, 132-168. doi:10.1016/j.ympev.2014.04.003 10.1016/j.ympev.2014.04.003 Elsevier Version of Record http://cdss.library.oregonstate.edu/sa-termsofuse Molecular Phylogenetics and Evolution 79 (2014) 132–168 Contents lists available at ScienceDirect Molecular Phylogenetics and Evolution journal homepage: www.elsevier.com/locate/ympev A multigene phylogenetic synthesis for the class Lecanoromycetes (Ascomycota): 1307 fungi representing 1139 infrageneric taxa, 317 genera and 66 families ⇑ Jolanta Miadlikowska a, , Frank Kauff b,1, Filip Högnabba c, Jeffrey C. Oliver d,2, Katalin Molnár a,3, Emily Fraker a,4, Ester Gaya a,5, Josef Hafellner e, Valérie Hofstetter a,6, Cécile Gueidan a,7, Mónica A.G. Otálora a,8, Brendan Hodkinson a,9, Martin Kukwa f, Robert Lücking g, Curtis Björk h, Harrie J.M. Sipman i, Ana Rosa Burgaz j, Arne Thell k, Alfredo Passo l, Leena Myllys c, Trevor Goward h, Samantha Fernández-Brime m, Geir Hestmark n, James Lendemer o, H. Thorsten Lumbsch g, Michaela Schmull p, Conrad L. Schoch q, Emmanuël Sérusiaux r, David R. Maddison s, A. Elizabeth Arnold t, François Lutzoni a,10, -

Modelling Lichen Abundance for Woodland Caribou in a Fire-Driven Boreal Landscape
Article Modelling Lichen Abundance for Woodland Caribou in a Fire-Driven Boreal Landscape Joseph A. Silva 1,*, Scott E. Nielsen 2 , Clayton T. Lamb 1 , Christine Hague 3 and Stan Boutin 1 1 Department of Biological Sciences, University of Alberta, Edmonton, AB T6G 2E9, Canada; [email protected] (C.T.L.); [email protected] (S.B.) 2 Department of Renewable Resources, University of Alberta, Edmonton, AB T6G 2H1, Canada; [email protected] 3 Ontario Parks, Ministry of Environment, Conservation and Parks, Red Lake, ON P0V 2M0, Canada; [email protected] * Correspondence: [email protected]; Tel.: +1-519-277-1940 Received: 18 September 2019; Accepted: 25 October 2019; Published: 1 November 2019 Abstract: Woodland caribou (Rangifer tarandus caribou) are reliant on Cladonia spp. ground lichens as a major component of their diet and lichen abundance could be an important indicator of habitat quality, particularly in winter. The boreal forest is typified by large, stand-replacing forest fires that consume ground lichens, which take decades to recover. The large spatial extent of caribou ranges and the mosaic of lichen availability created by fires make it challenging to track the abundance of ground lichens. Researchers have developed various techniques to map lichens across northern boreal and tundra landscapes, but it remains unclear which techniques are best suited for use in the continuous boreal forest, where many of the conflicts amongst caribou and human activities are most acute. In this study, we propose a two-stage regression modelling approach to map the abundance (biomass, kg/ha) of Cladonia spp. ground lichens in the boreal forest.