Package 'Dbparser'
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Therapeutic Potential of Nicotinamide Adenine Dinucleotide (NAD) T ∗ Marta Arenas-Jala,B, , J.M
European Journal of Pharmacology 879 (2020) 173158 Contents lists available at ScienceDirect European Journal of Pharmacology journal homepage: www.elsevier.com/locate/ejphar Therapeutic potential of nicotinamide adenine dinucleotide (NAD) T ∗ Marta Arenas-Jala,b, , J.M. Suñé-Negrea, Encarna García-Montoyaa a Pharmacy and Pharmaceutical Technology Department (Faculty of Pharmacy and Food Sciences), University of Barcelona, Barcelona, Spain b ICN2 – Catalan Institute of Nanoscience and Nanotechnology (Autonomous University of Barcelona), Bellaterra (Barcelona), Spain ARTICLE INFO ABSTRACT Keywords: Nicotinamide adenine nucleotide (NAD) is a small ubiquitous hydrophilic cofactor that participates in several NAD aspects of cellular metabolism. As a coenzyme it has an essential role in the regulation of energetic metabolism, Metabolism but it is also a cosubstrate for enzymes that regulate fundamental biological processes such as transcriptional Therapeutic potential regulation, signaling and DNA repairing among others. The fluctuation and oxidative state of NAD levels reg- Drug discovery ulate the activity of these enzymes, which is translated into marked effects on cellular function. While alterations Supplementation in NAD homeostasis are a common feature of different conditions and age-associated diseases, in general, in- creased NAD levels have been associated with beneficial health effects. Due to its therapeutic potential, the interest in this molecule has been renewed, and the regulation of NAD metabolism has become an attractive target for drug discovery. In fact, different approaches to replenish or increase NAD levels have been tested, including enhancement of biosynthesis and inhibition of NAD breakdown. Despite further research is needed, this review provides an overview and update on NAD metabolism, including the therapeutic potential of its regulation, as well as pharmacokinetics, safety, precautions and formulation challenges of NAD supplementa- tion. -

Metabolic-Hydroxy and Carboxy Functionalization of Alkyl Moieties in Drug Molecules: Prediction of Structure Influence and Pharmacologic Activity
molecules Review Metabolic-Hydroxy and Carboxy Functionalization of Alkyl Moieties in Drug Molecules: Prediction of Structure Influence and Pharmacologic Activity Babiker M. El-Haj 1,* and Samrein B.M. Ahmed 2 1 Department of Pharmaceutical Sciences, College of Pharmacy and Health Sciences, University of Science and Technology of Fujairah, Fufairah 00971, UAE 2 College of Medicine, Sharjah Institute for Medical Research, University of Sharjah, Sharjah 00971, UAE; [email protected] * Correspondence: [email protected] Received: 6 February 2020; Accepted: 7 April 2020; Published: 22 April 2020 Abstract: Alkyl moieties—open chain or cyclic, linear, or branched—are common in drug molecules. The hydrophobicity of alkyl moieties in drug molecules is modified by metabolic hydroxy functionalization via free-radical intermediates to give primary, secondary, or tertiary alcohols depending on the class of the substrate carbon. The hydroxymethyl groups resulting from the functionalization of methyl groups are mostly oxidized further to carboxyl groups to give carboxy metabolites. As observed from the surveyed cases in this review, hydroxy functionalization leads to loss, attenuation, or retention of pharmacologic activity with respect to the parent drug. On the other hand, carboxy functionalization leads to a loss of activity with the exception of only a few cases in which activity is retained. The exceptions are those groups in which the carboxy functionalization occurs at a position distant from a well-defined primary pharmacophore. Some hydroxy metabolites, which are equiactive with their parent drugs, have been developed into ester prodrugs while carboxy metabolites, which are equiactive to their parent drugs, have been developed into drugs as per se. -

Drugbank 3.0: a Comprehensive Resource for 'Omics' Research On
Published online 8 November 2010 Nucleic Acids Research, 2011, Vol. 39, Database issue D1035–D1041 doi:10.1093/nar/gkq1126 DrugBank 3.0: a comprehensive resource for ‘Omics’ research on drugs Craig Knox1, Vivian Law2, Timothy Jewison1, Philip Liu3, Son Ly2, Alex Frolkis1, Allison Pon1, Kelly Banco2, Christine Mak2, Vanessa Neveu1, Yannick Djoumbou3, Roman Eisner1, An Chi Guo1 and David S. Wishart1,2,3,4,* 1Department of Computing Science, University of Alberta, Edmonton, AB, Canada T6G 2E8, 2Faculty of Pharmacy and Pharmaceutical Sciences, University of Alberta, Edmonton, AB, Canada T6G 2N8, 3Department of Biological Sciences, University of Alberta, Edmonton, AB, Canada T6G 2E8 and 4National Institute for Nanotechnology, 11421 Saskatchewan Drive, Edmonton, AB, Canada T6G 2M9 Received September 15, 2010; Revised October 20, 2010; Accepted October 21, 2010 ABSTRACT drug target, drug description and drug action data. DrugBank (http://www.drugbank.ca) is a richly DrugBank 3.0 represents the result of 2 years annotated database of drug and drug target infor- of manual annotation work aimed at making mation. It contains extensive data on the nomencla- the database much more useful for a wide ture, ontology, chemistry, structure, function, range of ‘omics’ (i.e. pharmacogenomic, action, pharmacology, pharmacokinetics, metabol- pharmacoproteomic, pharmacometabolomic and ism and pharmaceutical properties of both small even pharmacoeconomic) applications. molecule and large molecule (biotech) drugs. It also contains comprehensive information on the INTRODUCTION target diseases, proteins, genes and organisms on which these drugs act. First released in 2006, Historically most of the known information on drugs, DrugBank has become widely used by pharmacists, drug targets and drug action has resided in books, medicinal chemists, pharmaceutical researchers, journals and expensive commercial databases. -

Smart Drugs: a Review
International Journal for Innovation Education and Research www.ijier.net Vol:-8 No-11, 2020 Smart Drugs: A Review Sahjesh Soni, Dr Rashmi Srivastava, Ayush Bhandari Mumbai Educational Trust, India Abstracts Smart drugs can change the way our mind functions. Smart drugs are also known as nootropics, which literally means the ability to bend or shape our mind. Smart drugs are classified into two main categories. They are classified based on their pharmacological action and their availability. The stimulant category of drugs is highly used and misused. There has been a rampant increase in the sale of smart drugs, which could be attributed to the rise in competition all over the world. Two major criteria for selecting a good drug are its mechanism of action and bioavailability. Owing to the short-term benefits of smart drugs, many countries have openly accepted this concept. There is still no concrete scientific evidence backing the safety and efficacy of these drugs. Some believe that this is just a fad that will soon pass, while others believe that this is something that will revolutionize our future. Key Words: Smart drugs, Nootropics, Cognitive enhancers, Stimulants, Uses and Side effects. What are Smart Drugs? "Smart drugs" are a group of compounds that can promote brain performance. They have got a lot of attention due to our stressful lifestyle, and these drugs help to boost our memory, focus, creativity, intelligence, and motivation. The origin of the word comes from the Greek language meaning “to bend or shape the mind”.1 These chemicals have many mechanisms of action. -

Drug Knowledge Bases and Their Applications in Biomedical Informatics Research Yongjun Zhu, Olivier Elemento, Jyotishman Pathak and Fei Wang
Briefings in Bioinformatics, 2018, 1–14 doi: 10.1093/bib/bbx169 Paper Drug knowledge bases and their applications in biomedical informatics research Yongjun Zhu, Olivier Elemento, Jyotishman Pathak and Fei Wang Corresponding author: Fei Wang, Division of Health Informatics, Department of Healthcare Policy and Research at Weill Cornell Medicine at Cornell University, 425 East 61st Street, Suite 301, DV-308, New York, NY 10065, USA. E-mail: [email protected] Abstract Recent advances in biomedical research have generated a large volume of drug-related data. To effectively handle this flood of data, many initiatives have been taken to help researchers make good use of them. As the results of these initiatives, many drug knowledge bases have been constructed. They range from simple ones with specific focuses to comprehensive ones that contain information on almost every aspect of a drug. These curated drug knowledge bases have made significant contributions to the development of efficient and effective health information technologies for better health-care service delivery. Understanding and comparing existing drug knowledge bases and how they are applied in various biomedical studies will help us recognize the state of the art and design better knowledge bases in the future. In addition, researchers can get insights on novel applications of the drug knowledge bases through a review of successful use cases. In this study, we provide a review of existing popular drug knowledge bases and their applications in drug-related studies. We discuss challenges in constructing and using drug knowledge bases as well as future research directions toward a better ecosystem of drug knowledge bases. -
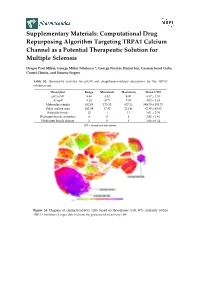
Computational Drug Repurposing Algorithm Targeting TRPA1 Calcium Channel As a Potential Therapeutic Solution for Multiple Sclerosis
Supplementary Materials: Computational Drug Repurposing Algorithm Targeting TRPA1 Calcium Channel as a Potential Therapeutic Solution for Multiple Sclerosis Dragos Paul Mihai, George Mihai Nitulescu *, George Nicolae Daniel Ion, Cosmin Ionut Ciotu, Cornel Chirita, and Simona Negres Table S1. Descriptive statistics for pIC50 and druglikeness-related descriptors for the TRPA1 inhibitors set. Descriptor Range Minimum Maximum Mean ± SD pIC50 (M) 4.48 4.52 9.00 6.57 ± 1.01 ALogP 8.28 −0.71 7.57 4.02 ± 1.34 Molecular weight 482.03 175.10 657.13 389.70 ± 101.73 Polar surface area 193.59 17.82 211.41 82.80 ± 40.83 Rotatable bonds 12 1 13 5.01 ± 2.08 Hydrogen bonds acceptors 8 0 8 2.92 ± 1.46 Hydrogen bonds donors 3 0 3 1.06 ± 0.54 SD – standard deviation. Figure S1. Diagram of similarity/activity cliffs based on flexophores with 80% similarity within TRPA1 inhibitors. Larger dots indicate the presence of an activity cliff. Figure S2. Representative structures for similarity/activity cliffs analysis of TRPA1 inhibitors. Table S2. Highest similarity pairs between TRPA1 inhibitors and screened drugs based on flexophore descriptors data mining procedure. TRPA1 inhibitors Repurposing dataset Entry Similarity (ChEMBL ID) (DrugBank ID) 1 CHEMBL3298238 DB08135 0.9832 2 CHEMBL3220230 DB08561 0.9696 3 CHEMBL3220228 DB08561 0.9614 4 CHEMBL593902 DB07311 0.9553 5 CHEMBL3297780 DB01065 0.9533 6 CHEMBL3220448 DB08561 0.9509 Figure S3. Diagram of similarity/activity cliffs based on flexophores with 80% similarity threshold for merged TRPA1 inhibitors dataset (colored dots) and similar DrugBank entries (grey dots). Table S3. -

Drug Repositioning: New Approaches and Future Prospects for Life-Debilitating Diseases and the COVID-19 Pandemic Outbreak
viruses Review Drug Repositioning: New Approaches and Future Prospects for Life-Debilitating Diseases and the COVID-19 Pandemic Outbreak Zheng Yao Low 1 , Isra Ahmad Farouk 1 and Sunil Kumar Lal 1,2,* 1 School of Science, Monash University, Bandar Sunway, Subang Jaya 47500, Selangor Darul Ehsan, Malaysia; [email protected] (Z.Y.L.); [email protected] (I.A.F.) 2 Tropical Medicine & Biology Platform, Monash University, Subang Jaya 47500, Selangor Darul Ehsan, Malaysia * Correspondence: [email protected] Received: 3 July 2020; Accepted: 21 August 2020; Published: 22 September 2020 Abstract: Traditionally, drug discovery utilises a de novo design approach, which requires high cost and many years of drug development before it reaches the market. Novel drug development does not always account for orphan diseases, which have low demand and hence low-profit margins for drug developers. Recently, drug repositioning has gained recognition as an alternative approach that explores new avenues for pre-existing commercially approved or rejected drugs to treat diseases aside from the intended ones. Drug repositioning results in lower overall developmental expenses and risk assessments, as the efficacy and safety of the original drug have already been well accessed and approved by regulatory authorities. The greatest advantage of drug repositioning is that it breathes new life into the novel, rare, orphan, and resistant diseases, such as Cushing’s syndrome, HIV infection, and pandemic outbreaks such as COVID-19. Repositioning existing drugs such as Hydroxychloroquine, Remdesivir, Ivermectin and Baricitinib shows good potential for COVID-19 treatment. This can crucially aid in resolving outbreaks in urgent times of need. -

Home Browse Drug Browse Pharma Browse Geno Browse Reaction
13/12/13 DrugBank: Asparaginase (DB00023) Home Browse Drug Browse Pharma Browse Geno Browse Reaction Browse Pathway Browse Class Browse Association Browse Search ChemQuery Text Query Interax Interaction Search Sequence Search Data Extractor Downloads About About DrugBank Statistics Other Databases Data Sources News Archive Wishart Research Group Help Citing DrugBank DrugCard Documentation Searching DrugBank Tools Human Metabolome Database T3DB Toxin Database Small Molecule Pathway Database FooDB Food Component Database More Contact Us DrugBank version 4.0 beta is now online for public preview! Take me to the beta site now. Search: Search DrugBank Search Help / Advanced Identification Taxonomy Pharmacology Pharmacoeconomics Properties References Interactions 0 Comments targets (1) Identification Name Asparaginase Accession Number DB00023 (BIOD00011, BTD00011) Type biotech Groups approved Description L-asparagine amidohydrolase from E. coli www.drugbank.ca/drugs/DB00023#identification 1/4 13/12/13 DrugBank: Asparaginase (DB00023) Protein structure Display: 3D Structure Protein chemical C H N O S formula 1377 2208 382 442 17 Protein average 31731.9000 weight >DB00023 sequence QMSLQQELRYIEALSAIVETGQKMLEAGESALDVVTEAVRLLEECPLFNAGIGAVFTRDE THELDACVMDGNTLKAGAVAGVSHLRNPVLAARLVMEQSPHVMMIGEGAENFAFARGMER VSPEIFSTSLRYEQLLAARKEGATVLDHSGAPLDEKQKMGTVGAVALDLDGNLAAATSTG Sequences GMTNKLPGRVGDSPLVGAGCYANNASVAVSCTGTGEVFIRALAAYDIAALMDYGGLSLAE ACERVVMEKLPALGGSGGLIAIDHEGNVALPFNTEGMYRAWGYAGDTPTTGIYREKGDTV ATQ FASTA L-asparagine amidohydrolase Synonyms Putative -

Potential Herb–Drug Interactions in the Management of Age-Related Cognitive Dysfunction
pharmaceutics Review Potential Herb–Drug Interactions in the Management of Age-Related Cognitive Dysfunction Maria D. Auxtero 1, Susana Chalante 1,Mário R. Abade 1 , Rui Jorge 1,2,3 and Ana I. Fernandes 1,* 1 CiiEM, Interdisciplinary Research Centre Egas Moniz, Instituto Universitário Egas Moniz, Quinta da Granja, Monte de Caparica, 2829-511 Caparica, Portugal; [email protected] (M.D.A.); [email protected] (S.C.); [email protected] (M.R.A.); [email protected] (R.J.) 2 Polytechnic Institute of Santarém, School of Agriculture, Quinta do Galinheiro, 2001-904 Santarém, Portugal 3 CIEQV, Life Quality Research Centre, IPSantarém/IPLeiria, Avenida Dr. Mário Soares, 110, 2040-413 Rio Maior, Portugal * Correspondence: [email protected]; Tel.: +35-12-1294-6823 Abstract: Late-life mild cognitive impairment and dementia represent a significant burden on health- care systems and a unique challenge to medicine due to the currently limited treatment options. Plant phytochemicals have been considered in alternative, or complementary, prevention and treat- ment strategies. Herbals are consumed as such, or as food supplements, whose consumption has recently increased. However, these products are not exempt from adverse effects and pharmaco- logical interactions, presenting a special risk in aged, polymedicated individuals. Understanding pharmacokinetic and pharmacodynamic interactions is warranted to avoid undesirable adverse drug reactions, which may result in unwanted side-effects or therapeutic failure. The present study reviews the potential interactions between selected bioactive compounds (170) used by seniors for cognitive enhancement and representative drugs of 10 pharmacotherapeutic classes commonly prescribed to the middle-aged adults, often multimorbid and polymedicated, to anticipate and prevent risks arising from their co-administration. -

A Major Update to the Drugbank Database for 2018 David S
D1074–D1082 Nucleic Acids Research, 2018, Vol. 46, Database issue Published online 8 November 2017 doi: 10.1093/nar/gkx1037 DrugBank 5.0: a major update to the DrugBank database for 2018 David S. Wishart1,2,3,4,*, Yannick D. Feunang1,AnC.Guo1, Elvis J. Lo1,AnaMarcu1, Jason R. Grant1, Tanvir Sajed2, Daniel Johnson1, Carin Li1, Zinat Sayeeda1, Nazanin Assempour1, Ithayavani Iynkkaran1,4, Yifeng Liu2, Adam Maciejewski1, Nicola Gale5, Alex Wilson5, Lucy Chin5, Ryan Cummings5, Diana Le5, Allison Pon1,5,CraigKnox1,5 and Michael Wilson1,5 1Department of Biological Sciences, University of Alberta, Edmonton, AB T6G 2E9, Canada, 2Department of Computing Science, University of Alberta, Edmonton, AB T6G 2E8, Canada, 3Faculty of Pharmacy and Pharmaceutical Sciences, University of Alberta, Edmonton, AB T6G 2N8, Canada, 4Department of Laboratory Medicine and Pathology, University of Alberta, Edmonton, AB T6G 2R3, Canada and 5OMx Personal Health Analytics, Inc., 301-10359 104 St NW, Edmonton, AB T5J 1B9, Canada Received September 15, 2017; Revised October 12, 2017; Editorial Decision October 13, 2017; Accepted November 03, 2017 ABSTRACT content, interface and performance of the DrugBank website have been made and these should greatly DrugBank (www.drugbank.ca) is a web-enabled enhance its ease of use, utility and potential appli- database containing comprehensive molecular infor- cations in many areas of pharmacological research, mation about drugs, their mechanisms, their interac- pharmaceutical science and drug education. tions and their targets. First described in 2006, Drug- Bank has continued to evolve over the past 12 years in response to marked improvements to web stan- INTRODUCTION dards and changing needs for drug research and de- DrugBank is a comprehensive, freely available web resource velopment. -

Product Monograph Including Patient
PRODUCT MONOGRAPH INCLUDING PATIENT MEDICATION INFORMATION N METADOL® Methadone Hydrochloride Tablets 1 mg, 5 mg, 10 mg and 25 mg Methadone Hydrochloride Oral Solution USP 1 mg/mL Methadone Hydrochloride Oral Concentrate USP 10 mg/mL Opioid Analgesic Paladin Labs Inc. Date of Revision: 100 Blvd. Alexis Nihon, Suite 600 August 15, 2018 Saint-Laurent, Québec Version: 12.0 H4M 2P2 Submission Control No: 211399 ® Registered trademark of Paladin Labs Inc. METADOL® Product Monograph Page 1 of 46 TABLE OF CONTENTS PART I: HEALTH PROFESSIONAL INFORMATION .........................................................3 SUMMARY PRODUCT INFORMATION ........................................................................3 INDICATIONS AND CLINICAL USE ..............................................................................3 CONTRAINDICATIONS ...................................................................................................4 WARNINGS AND PRECAUTIONS ..................................................................................4 ADVERSE REACTIONS ..................................................................................................15 DRUG INTERACTIONS ..................................................................................................17 DOSAGE AND ADMINISTRATION ..............................................................................20 OVERDOSAGE ................................................................................................................22 ACTION AND CLINICAL PHARMACOLOGY ............................................................24 -

NMN) from Bacterial Cells Published: Xx Xx Xxxx George Cătălin Marinescu 1,2, Roua-Gabriela Popescu1,2 & Anca Dinischiotu1
www.nature.com/scientificreports OPEN Size Exclusion Chromatography Method for Purifcation of Nicotinamide Mononucleotide Received: 24 July 2017 Accepted: 1 March 2018 (NMN) from Bacterial Cells Published: xx xx xxxx George Cătălin Marinescu 1,2, Roua-Gabriela Popescu1,2 & Anca Dinischiotu1 Over 12% of the world’s health resources are spent on treating diabetes, as high blood glucose is the third cause of mortality worldwide. Insulin resistance is the basis of the most common form of diabetes: type 2 diabetes. Recent animal studies report successful attempts at reversing type 2 diabetes by the administering of the NAD+ precursor nicotinamide mononucleotide (NMN). However, the current high price of this molecule urges for more efcient and cost-efective production methods. This work proposes a method for purifying NMN by Size Exclusion Chromatography (SEC) on silica with a covalently attached coating of poly(2-hydroxyethyl aspartamide) (PolyHEA) stationary phase using an isocratic elution with a denaturing mobile phase (50 mM formic acid) from a complex molecular mixture such as a fermentation broth. The eluted peaks were identifed by UV-Vis analysis and confrmed with ESI+ mass spectrometry and a HPLC reversed-phase method. The proposed SEC method is simple, patent-free, directly applicable for industrial production with a minimum scale up efort. The need for multiple chromatographic steps is eliminated and the lysate fltration and clarifcation steps are simplifed. Substantial reduction in NMN production costs and increased purity of NMN to the level suitable for usage in humans are expected. High blood glucose level is the third highest risk factor for premature mortality worldwide.