SLURP1 Gene Secreted LY6/PLAUR Domain Containing 1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Hippo and Sonic Hedgehog Signalling Pathway Modulation of Human Urothelial Tissue Homeostasis
Hippo and Sonic Hedgehog signalling pathway modulation of human urothelial tissue homeostasis Thomas Crighton PhD University of York Department of Biology November 2020 Abstract The urinary tract is lined by a barrier-forming, mitotically-quiescent urothelium, which retains the ability to regenerate following injury. Regulation of tissue homeostasis by Hippo and Sonic Hedgehog signalling has previously been implicated in various mammalian epithelia, but limited evidence exists as to their role in adult human urothelial physiology. Focussing on the Hippo pathway, the aims of this thesis were to characterise expression of said pathways in urothelium, determine what role the pathways have in regulating urothelial phenotype, and investigate whether the pathways are implicated in muscle-invasive bladder cancer (MIBC). These aims were assessed using a cell culture paradigm of Normal Human Urothelial (NHU) cells that can be manipulated in vitro to represent different differentiated phenotypes, alongside MIBC cell lines and The Cancer Genome Atlas resource. Transcriptomic analysis of NHU cells identified a significant induction of VGLL1, a poorly understood regulator of Hippo signalling, in differentiated cells. Activation of upstream transcription factors PPARγ and GATA3 and/or blockade of active EGFR/RAS/RAF/MEK/ERK signalling were identified as mechanisms which induce VGLL1 expression in NHU cells. Ectopic overexpression of VGLL1 in undifferentiated NHU cells and MIBC cell line T24 resulted in significantly reduced proliferation. Conversely, knockdown of VGLL1 in differentiated NHU cells significantly reduced barrier tightness in an unwounded state, while inhibiting regeneration and increasing cell cycle activation in scratch-wounded cultures. A signalling pathway previously observed to be inhibited by VGLL1 function, YAP/TAZ, was unaffected by VGLL1 manipulation. -

Organization, Evolution and Functions of the Human and Mouse Ly6/Upar Family Genes Chelsea L
Loughner et al. Human Genomics (2016) 10:10 DOI 10.1186/s40246-016-0074-2 GENE FAMILY UPDATE Open Access Organization, evolution and functions of the human and mouse Ly6/uPAR family genes Chelsea L. Loughner1, Elspeth A. Bruford2, Monica S. McAndrews3, Emili E. Delp1, Sudha Swamynathan1 and Shivalingappa K. Swamynathan1,4,5,6,7* Abstract Members of the lymphocyte antigen-6 (Ly6)/urokinase-type plasminogen activator receptor (uPAR) superfamily of proteins are cysteine-rich proteins characterized by a distinct disulfide bridge pattern that creates the three-finger Ly6/uPAR (LU) domain. Although the Ly6/uPAR family proteins share a common structure, their expression patterns and functions vary. To date, 35 human and 61 mouse Ly6/uPAR family members have been identified. Based on their subcellular localization, these proteins are further classified as GPI-anchored on the cell membrane, or secreted. The genes encoding Ly6/uPAR family proteins are conserved across different species and are clustered in syntenic regions on human chromosomes 8, 19, 6 and 11, and mouse Chromosomes 15, 7, 17, and 9, respectively. Here, we review the human and mouse Ly6/uPAR family gene and protein structure and genomic organization, expression, functions, and evolution, and introduce new names for novel family members. Keywords: Ly6/uPAR family, LU domain, Three-finger domain, uPAR, Lymphocytes, Neutrophils Introduction an overview of the Ly6/uPAR gene family and their gen- The lymphocyte antigen-6 (Ly6)/urokinase-type plas- omic organization, evolution, as well as functions, and minogen activator receptor (uPAR) superfamily of struc- provide a nomenclature system for the newly identified turally related proteins is characterized by the LU members of this family. -

UNIVERSITY of CALIFORNIA, IRVINE Gene Regulatory
UNIVERSITY OF CALIFORNIA, IRVINE Gene Regulatory Mechanisms in Epithelial Specification and Function DISSERTATION submitted in partial satisfaction of the requirements for the degree of DOCTOR OF PHILOSOPHY in Biomedical Sciences by Rachel Herndon Klein Dissertation Committee: Professor Bogi Andersen, M.D., Chair Professor Xing Dai, Ph.D. Professor Anand Ganesan, M.D. Professor Ali Mortazavi, Ph.D Professor Kyoko Yokomori, Ph.D 2015 © 2015 Rachel Herndon Klein DEDICATION To My parents, my sisters, my husband, and my friends for your love and support, and to Ben with all my love. ii TABLE OF CONTENTS Page LIST OF FIGURES iv LIST OF TABLES vi ACKNOWLEDGMENTS vii CURRICULUM VITAE viii-ix ABSTRACT OF THE DISSERTATION x-xi CHAPTER 1: INTRODUCTION 1 CHAPTER 2: Cofactors of LIM domain (CLIM) proteins regulate corneal epithelial progenitor cell function through noncoding RNA H19 22 CHAPTER 3: KLF7 regulates the corneal epithelial progenitor cell state acting antagonistically to KLF4 49 CHAPTER 4: GRHL3 interacts with super enhancers and the neuronal repressor REST to regulate keratinocyte differentiation and migration 77 CHAPTER 5: Methods 103 CHAPTER 6: Summary and Conclusions 111 REFERENCES 115 iii LIST OF FIGURES Page Figure 1-1. Structure and organization of the epidermis. 3 Figure 1-2. Structure of the limbus, and cornea epithelium. 4 Figure 1-3. Comparison of H3K4 methylating SET enzymes between S. cerevisiae, D. melanogaster, and H. sapiens. 18 Figure 1-4. The WRAD complex associates with Trithorax SET enzymes. 18 Figure 1-5. Model for GRHL3, PcG, and TrX –mediated regulation of epidermal differentiation genes. 19 Figure 2-1. Microarray gene expression analysis of postnatal day 3 (P3) whole mouse corneas reveals genes and pathways with altered expression in K14-DN-Clim mice. -

High Mrna Expression of LY6 Gene Family Is Associated with Overall Survival Outcome in Pancreatic Ductal Adenocarcinoma
www.oncotarget.com Oncotarget, 2021, Vol. 12, (No. 3), pp: 145-159 Research Paper High mRNA expression of LY6 gene family is associated with overall survival outcome in pancreatic ductal adenocarcinoma Eric Russ1, Krithika Bhuvaneshwar2, Guisong Wang3,4, Benjamin Jin1,4, Michele M. Gage3,5, Subha Madhavan2, Yuriy Gusev2 and Geeta Upadhyay1,3 1Department of Pathology, Uniformed Services University, Bethesda, MD, USA 2Innovation Center for Biomedical Informatics, Georgetown University Medical Center, Washington DC, USA 3Murtha Cancer Center/Research Program, Department of Surgery, Uniformed Services University of the Health Sciences, Bethesda, MD, USA 4The Henry M. Jackson Foundation for the Advancement of Military Medicine Inc, Bethesda, MD, USA 5Walter Reed Navy Military Medical Center, Department of Surgery, Uniformed Services University, Bethesda, MD, USA Correspondence to: Geeta Upadhyay, email: [email protected] Keywords: LY6 genes; pancreatic cancer; immune cells; survival outcome Received: November 21, 2020 Accepted: January 19, 2021 Published: February 02, 2021 Copyright: © 2021 Russ et al. This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Pancreatic cancer ranks one of the worst in overall survival outcome with a 5 year survival rate being less than 10%. Pancreatic cancer faces unique challenges in its diagnosis and treatment, such as the lack of clinically validated biomarkers and the immensely immunosuppressive tumor microenvironment. Recently, the LY6 gene family has received increasing attention for its multi-faceted roles in cancer development, stem cell maintenance, immunomodulation, and association with more aggressive and hard-to-treat cancers. -

Structural Variant Selection for High-Altitude Adaptation Using Single-Molecule 2 Long-Read Sequencing
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.27.436702; this version posted March 27, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Structural variant selection for high-altitude adaptation using single-molecule 2 long-read sequencing 3 Jinlong Shi1,2,4*, Zhilong Jia1,2,3*, Xiaojing Zhao2*, Jinxiu Sun4*, Fan Liang5*, Minsung Park5*, Chenghui Zhao1, 4 Xiaoreng Wang2, Qi Chen4, Xinyu Song2,3, Kang Yu1, Qian Jia2, Depeng Wang5, Yuhui Xiao5, Yinzhe Liu5, Shijing 5 Wu1, Qin Zhong2, Jue Wu2, Saijia Cui2, Xiaochen Bo6, Zhenzhou Wu7, Manolis Kellis8,9, Kunlun He1,2# 6 1. Key Laboratory of Biomedical Engineering and Translational Medicine, Ministry of Industry and Information 7 Technology, Chinese PLA General Hospital, Beijing, China. 8 2. Beijing Key Laboratory for Precision Medicine of Chronic Heart Failure, Chinese PLA General Hospital, Beijing, 9 China. 10 3. Research Center of Medical Artificial Intelligence, Chinese PLA General Hospital, Beijing, China. 11 4. Research Center of Medical Big Data, Chinese PLA General Hospital, Beijing, China. 12 5. GrandOmics Biosciences Inc, Beijing, China. 13 6. Beijing Institute of Radiation Medicine, Beijing, China. 14 7. BioMind Inc, Beijing, China. 15 8. Computer Science and Artificial Intelligence Laboratory, Massachusetts Institute of Technology, Cambridge, 16 MA, USA. 17 9. Broad Institute of MIT and Harvard, Cambridge, MA, USA. 18 *These authors contributed equally to this work. 19 #Corresponding author: [email protected] 20 21 Abstract: (150 words) 22 Structural variants (SVs) can be important drivers of human adaptation with strong effects, but previous studies 23 have focused primarily on common variants with weak effects. -

The Hypothalamus As a Hub for SARS-Cov-2 Brain Infection and Pathogenesis
bioRxiv preprint doi: https://doi.org/10.1101/2020.06.08.139329; this version posted June 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. The hypothalamus as a hub for SARS-CoV-2 brain infection and pathogenesis Sreekala Nampoothiri1,2#, Florent Sauve1,2#, Gaëtan Ternier1,2ƒ, Daniela Fernandois1,2 ƒ, Caio Coelho1,2, Monica ImBernon1,2, Eleonora Deligia1,2, Romain PerBet1, Vincent Florent1,2,3, Marc Baroncini1,2, Florence Pasquier1,4, François Trottein5, Claude-Alain Maurage1,2, Virginie Mattot1,2‡, Paolo GiacoBini1,2‡, S. Rasika1,2‡*, Vincent Prevot1,2‡* 1 Univ. Lille, Inserm, CHU Lille, Lille Neuroscience & Cognition, DistAlz, UMR-S 1172, Lille, France 2 LaBoratorY of Development and PlasticitY of the Neuroendocrine Brain, FHU 1000 daYs for health, EGID, School of Medicine, Lille, France 3 Nutrition, Arras General Hospital, Arras, France 4 Centre mémoire ressources et recherche, CHU Lille, LiCEND, Lille, France 5 Univ. Lille, CNRS, INSERM, CHU Lille, Institut Pasteur de Lille, U1019 - UMR 8204 - CIIL - Center for Infection and ImmunitY of Lille (CIIL), Lille, France. # and ƒ These authors contriButed equallY to this work. ‡ These authors directed this work *Correspondence to: [email protected] and [email protected] Short title: Covid-19: the hypothalamic hypothesis 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.06.08.139329; this version posted June 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Gnomad Lof Supplement
1 gnomAD supplement gnomAD supplement 1 Data processing 4 Alignment and read processing 4 Variant Calling 4 Coverage information 5 Data processing 5 Sample QC 7 Hard filters 7 Supplementary Table 1 | Sample counts before and after hard and release filters 8 Supplementary Table 2 | Counts by data type and hard filter 9 Platform imputation for exomes 9 Supplementary Table 3 | Exome platform assignments 10 Supplementary Table 4 | Confusion matrix for exome samples with Known platform labels 11 Relatedness filters 11 Supplementary Table 5 | Pair counts by degree of relatedness 12 Supplementary Table 6 | Sample counts by relatedness status 13 Population and subpopulation inference 13 Supplementary Figure 1 | Continental ancestry principal components. 14 Supplementary Table 7 | Population and subpopulation counts 16 Population- and platform-specific filters 16 Supplementary Table 8 | Summary of outliers per population and platform grouping 17 Finalizing samples in the gnomAD v2.1 release 18 Supplementary Table 9 | Sample counts by filtering stage 18 Supplementary Table 10 | Sample counts for genomes and exomes in gnomAD subsets 19 Variant QC 20 Hard filters 20 Random Forest model 20 Features 21 Supplementary Table 11 | Features used in final random forest model 21 Training 22 Supplementary Table 12 | Random forest training examples 22 Evaluation and threshold selection 22 Final variant counts 24 Supplementary Table 13 | Variant counts by filtering status 25 Comparison of whole-exome and whole-genome coverage in coding regions 25 Variant annotation 30 Frequency and context annotation 30 2 Functional annotation 31 Supplementary Table 14 | Variants observed by category in 125,748 exomes 32 Supplementary Figure 5 | Percent observed by methylation. -

Novel and Highly Recurrent Chromosomal Alterations in Se´Zary Syndrome
Research Article Novel and Highly Recurrent Chromosomal Alterations in Se´zary Syndrome Maarten H. Vermeer,1 Remco van Doorn,1 Remco Dijkman,1 Xin Mao,3 Sean Whittaker,3 Pieter C. van Voorst Vader,4 Marie-Jeanne P. Gerritsen,5 Marie-Louise Geerts,6 Sylke Gellrich,7 Ola So¨derberg,8 Karl-Johan Leuchowius,8 Ulf Landegren,8 Jacoba J. Out-Luiting,1 Jeroen Knijnenburg,2 Marije IJszenga,2 Karoly Szuhai,2 Rein Willemze,1 and Cornelis P. Tensen1 Departments of 1Dermatology and 2Molecular Cell Biology, Leiden University Medical Center, Leiden, the Netherlands; 3Department of Dermatology, St Thomas’ Hospital, King’s College, London, United Kingdom; 4Department of Dermatology, University Medical Center Groningen, Groningen, the Netherlands; 5Department of Dermatology, Radboud University Nijmegen Medical Center, Nijmegen, the Netherlands; 6Department of Dermatology, Gent University Hospital, Gent, Belgium; 7Department of Dermatology, Charite, Berlin, Germany; and 8Department of Genetics and Pathology, Rudbeck Laboratory, University of Uppsala, Uppsala, Sweden Abstract Introduction This study was designed to identify highly recurrent genetic Se´zary syndrome (Sz) is an aggressive type of cutaneous T-cell alterations typical of Se´zary syndrome (Sz), an aggressive lymphoma/leukemia of skin-homing, CD4+ memory T cells and is cutaneous T-cell lymphoma/leukemia, possibly revealing characterized by erythroderma, generalized lymphadenopathy, and pathogenetic mechanisms and novel therapeutic targets. the presence of neoplastic T cells (Se´zary cells) in the skin, lymph High-resolution array-based comparative genomic hybridiza- nodes, and peripheral blood (1). Sz has a poor prognosis, with a tion was done on malignant T cells from 20 patients. disease-specific 5-year survival of f24% (1). -
Downloaded from the Mouse Lysosome Gene Database, Mlgdb
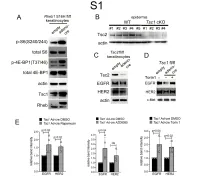
1 Supplemental Figure Legends 2 3 Supplemental Figure S1: Epidermal-specific mTORC1 gain-of-function models show 4 increased mTORC1 activation and down-regulate EGFR and HER2 protein expression in a 5 mTORC1-sensitive manner. (A) Immunoblotting of Rheb1 S16H flox/flox keratinocyte cultures 6 infected with empty or adenoviral cre recombinase for markers of mTORC1 (p-S6, p-4E-BP1) 7 activity. (B) Tsc1 cKO epidermal lysates also show decreased expression of TSC2 by 8 immunoblotting of the same experiment as in Figure 2A. (C) Immunoblotting of Tsc2 flox/flox 9 keratinocyte cultures infected with empty or adenoviral cre recombinase showing decreased EGFR 10 and HER2 protein expression. (D) Expression of EGFR and HER2 was decreased in Tsc1 cre 11 keratinocytes compared to empty controls, and up-regulated in response to Torin1 (1µM, 24 hrs), 12 by immunoblot analyses. Immunoblots are contemporaneous and parallel from the same biological 13 replicate and represent the same experiment as depicted in Figure 7B. (E) Densitometry 14 quantification of representative immunoblot experiments shown in Figures 2E and S1D (r≥3; error 15 bars represent STDEV; p-values by Student’s T-test). 16 17 18 19 20 21 22 23 Supplemental Figure S2: EGFR and HER2 transcription are unchanged with epidermal/ 24 keratinocyte Tsc1 or Rptor loss. Egfr and Her2 mRNA levels in (A) Tsc1 cKO epidermal lysates, 25 (B) Tsc1 cKO keratinocyte lysates and(C) Tsc1 cre keratinocyte lysates are minimally altered 26 compared to their respective controls. (r≥3; error bars represent STDEV; p-values by Student’s T- 27 test). -

Cytokines Mapping for Tissue-Specific Expression, Eqtls and GWAS Traits
www.nature.com/scientificreports OPEN Cytokines mapping for tissue‑specifc expression, eQTLs and GWAS traits Lyubov E. Salnikova1,2*, Maryam B. Khadzhieva1,2, Dmitry S. Kolobkov1,3,4, Alesya S. Gracheva1,2, Artem N. Kuzovlev2 & Serikbay K. Abilev1 Dysregulation in cytokine production has been linked to the pathogenesis of various immune‑ mediated traits, in which genetic variability contributes to the etiopathogenesis. GWA studies have identifed many genetic variants in or near cytokine genes, nonetheless, the translation of these fndings into knowledge of functional determinants of complex traits remains a fundamental challenge. In this study we aimed at collection, analysis and interpretation of data on cytokines focused on their tissue‑specifc expression, eQTLs and GWAS traits. Using GO annotations, we generated a list of 314 cytokines and analyzed them with the GTEx resource. Cytokines were highly tissue‑specifc, 82.3% of cytokines had Tau expression metrics ≥ 0.8. In total, 3077 associations for 1760 unique SNPs in or near 244 cytokines were mapped in the NHGRI‑EBI GWAS Catalog. According to the Experimental Factor Ontology resource, the largest numbers of disease associations were related to ‘Infammatory disease’, ‘Immune system disease’ and ‘Asthma’. The GTEx‑based analysis revealed that among GWAS SNPs, 1142 SNPs had eQTL efects and infuenced expression levels of 999 eGenes, among them 178 cytokines. Several types of enrichment analysis showed that it was cytokines expression variability that fundamentally contributed to the molecular origins of considered immune-mediated conditions. Cytokines are regulatory proteins and glycoproteins that are synthesized and secreted by immune system cells and other cell types. Tey regulate innate and acquired immunity, embryogenesis, hematopoiesis, infammation and regeneration processes, and proliferation. -

Targeted Disruption of Lynx2 Reveals Distinct Functions for Lynx Homologues in Learning and Behavior Ayse Begum Tekinay
Rockefeller University Digital Commons @ RU Student Theses and Dissertations 2007 Targeted Disruption of Lynx2 Reveals Distinct Functions for Lynx Homologues in Learning and Behavior Ayse Begum Tekinay Follow this and additional works at: http://digitalcommons.rockefeller.edu/ student_theses_and_dissertations Part of the Life Sciences Commons Recommended Citation Tekinay, Ayse Begum, "Targeted Disruption of Lynx2 Reveals Distinct Functions for Lynx Homologues in Learning and Behavior" (2007). Student Theses and Dissertations. Paper 28. This Thesis is brought to you for free and open access by Digital Commons @ RU. It has been accepted for inclusion in Student Theses and Dissertations by an authorized administrator of Digital Commons @ RU. For more information, please contact [email protected]. TARGETED DISRUPTION OF LYNX2 REVEALS DISTINCT FUNCTIONS FOR LYNX HOMOLOGUES IN LEARNING AND BEHAVIOR A Thesis Presented to the Faculty of The Rockefeller University In Partial Fulfillment of the Requirements for the degree of Doctor of Philosophy by Ayse Begum Tekinay June 2007 © Copyright by Ayse Begum Tekinay 2007 TARGETED DISRUPTION OF LYNX2 REVEALS DISTINCT FUNCTIONS FOR LYNX HOMOLOGUES IN LEARNING AND BEHAVIOR Ayse Begum Tekinay, Ph.D. The Rockefeller University 2007 Endogenous short peptide modulators of ion channels provide a new level of regulation of nervous function. Lynx1 was identified as an endogenous mammalian homologue of snake venom peptide neurotoxins capable of binding to and functionally modulating nicotinic acetylcholine receptors (nAChR). Lynx1 is a member of the Ly6- α−neurotoxin superfamily (Ly6SF) of genes. Through extensive database searches, I identified 85 members of this superfamily including previously unidentified vertebrate and invertebrate family members. I show that these proteins are very divergent in their sequences, and identify two conserved subfamilies, snake toxins and immune system expressed Ly6 genes through phylogenetic inference. -

Study on the Selection of the Targets of Esophageal Carcinoma and Interventions of Ginsenosides Based on Network Pharmacology and Bioinformatics
Hindawi Evidence-Based Complementary and Alternative Medicine Volume 2020, Article ID 4821056, 10 pages https://doi.org/10.1155/2020/4821056 Research Article Study on the Selection of the Targets of Esophageal Carcinoma and Interventions of Ginsenosides Based on Network Pharmacology and Bioinformatics Xin Yang , Yahui Li , and Haibing Qian Guizhou University of Traditional Chinese Medicine, Guiyang 550025, China Correspondence should be addressed to Haibing Qian; [email protected] Received 16 January 2020; Revised 3 May 2020; Accepted 27 May 2020; Published 24 June 2020 Academic Editor: Armando Zarrelli Copyright © 2020 Xin Yang et al. ,is is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Background. Esophageal carcinoma (ESCA) is not only a threat to people’s health but also the sixth most common cause of cancer- related mortality worldwide. Methods. In this study, the key targets of ESCA are screened through GeneCards and DisGeNET databases combined with the Gene Expression Omnibus (GEO) database (GSE1420 and GSE20347). ,en, data associated with ESCA samples are downloaded from ,e Cancer Genome Atlas (TCGA) database for integrated analysis. Moreover, the effect of epithelial cell adhesion molecule (EpCAM) expression on the survival of patients with ESCA is evaluated by Kaplan–Meier and Cox analyses. ,e virtual screening is carried out using a Suflex-Dock molecular docking module. ,e chemical components, which have been well bound to EpCAM, are screened out based on a total score >5 as a threshold. Ginsenosides and EpCAM are analyzed by LigPlot + v.2.2 software to identify the binding sites.