Pushing the Envelope
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Herpes Simplex Virus Type 1 Oril Is Not Required for Virus Infection In
JOURNAL OF VIROLOGY, Nov. 1987, p. 3528-3535 Vol. 61, No. 11 0022-538X/87/113528-08$02.00/0 Copyright C 1987, American Society for Microbiology Herpes Simplex Virus Type 1 oriL Is Not Required for Virus Replication or for the Establishment and Reactivation of Latent Infection in Mice MARYELLEN POLVINO-BODNAR, PAULO K. ORBERG, AND PRISCILLA A. SCHAFFER* Laboratory of Tumor Virus Genetics, Dana-Farber Cancer Institute, and Department of Microbiology and Molecular Genetics, Harvard Medical School, Boston, Massachusetts 02115 Received 11 May 1987/Accepted 31 July 1987 During the course of experiments designed to isolate deletion mutants of herpes simplex virus type 1 in the gene encoding the major DNA-binding protein, ICP8, a mutant, d61, that grew efficiently in ICP8-expressing Vero cells but not in normal Vero cells was isolated (P. K. Orberg and P. A. Schaffer, J. Virol. 61:1136-1146, 1987). d61 was derived by cotransfection of ICP8-expressing Vero cells with infectious wild-type viral DNA and a plasmid, pDX, that contains an engineered 780-base-pair (bp) deletion in the ICP8 gene, as well as a spontaneous -55-bp deletion in OriL. Gel electrophoresis and Southern blot analysis indicated that d61 DNA carried both deletions present in pDX. The ability of d61 to replicate despite the deletion in OriL suggested that a functional OriL is not essential for virus replication in vitro. Because d61 harbored two mutations, a second mutant, ts+7, with a deletion in oriL-associated sequences and an intact ICP8 gene was constructed. Both d61 and ts+7 replicated efficiently in their respective permissive host cells, although their yields were slightly lower than those of control viruses with intact oriL sequences. -

HERV-K(HML7) Integrations in the Human Genome: Comprehensive Characterization and Comparative Analysis in Non-Human Primates
biology Article HERV-K(HML7) Integrations in the Human Genome: Comprehensive Characterization and Comparative Analysis in Non-Human Primates Nicole Grandi 1,* , Maria Paola Pisano 1 , Eleonora Pessiu 1, Sante Scognamiglio 1 and Enzo Tramontano 1,2 1 Laboratory of Molecular Virology, Department of Life and Environmental Sciences, University of Cagliari, 09042 Monserrato, Cagliari, Italy; [email protected] (M.P.P.); [email protected] (E.P.); [email protected] (S.S.); [email protected] (E.T.) 2 Istituto di Ricerca Genetica e Biomedica, Consiglio Nazionale delle Ricerche (CNR), 09042 Monserrato, Cagliari, Italy * Correspondence: [email protected] Simple Summary: The human genome is not human at all, but it includes a multitude of sequences inherited from ancient viral infections that affected primates’ germ line. These elements can be seen as the fossils of now-extinct retroviruses, and are called Human Endogenous Retroviruses (HERVs). View as “junk DNA” for a long time, HERVs constitute 4 times the amount of DNA needed to produce all cellular proteins, and growing evidence indicates their crucial role in primate brain evolution, placenta development, and innate immunity shaping. HERVs are also intensively studied for a pathological role, even if the incomplete knowledge about their exact number and genomic position has thus far prevented any causal association. Among possible relevant HERVs, the HERV-K Citation: Grandi, N.; Pisano, M.P.; supergroup is of particular interest, including some of the oldest (HML5) as well as youngest (HML2) Pessiu, E.; Scognamiglio, S.; integrations. Among HERV-Ks, the HML7 group still lack a detailed description, and the present Tramontano, E. -

Topological Analysis of the Gp41 MPER on Lipid Bilayers Relevant to the Metastable HIV-1 Envelope Prefusion State
Topological analysis of the gp41 MPER on lipid bilayers relevant to the metastable HIV-1 envelope prefusion state Yi Wanga,b, Pavanjeet Kaurc,d, Zhen-Yu J. Sune,1, Mostafa A. Elbahnasawya,b,2, Zahra Hayatic,d, Zhi-Song Qiaoa,b,3, Nhat N. Buic, Camila Chilea,b,4, Mahmoud L. Nasre,5, Gerhard Wagnere, Jia-Huai Wanga,f, Likai Songc, Ellis L. Reinherza,b,6, and Mikyung Kima,g,6 aLaboratory of Immunobiology, Department of Medical Oncology, Dana-Farber Cancer Institute, Boston, MA 02115; bDepartment of Medicine, Harvard Medical School, Boston, MA 02115; cNational High Magnetic Field Laboratory, Florida State University, Tallahassee, FL 32306; dDepartment of Physics, Florida State University, Tallahassee, FL 32306; eDepartment of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115; fDepartment of Cancer Biology, Dana-Farber Cancer Institute, Boston, MA 02215; and gDepartment of Dermatology, Harvard Medical School, Boston, MA 02215 Edited by Peter Palese, Icahn School of Medicine at Mount Sinai, New York, NY, and approved September 23, 2019 (received for review July 18, 2019) The membrane proximal external region (MPER) of HIV-1 envelope immunologically vulnerable epitopes targeted by several of the most glycoprotein (gp) 41 is an attractive vaccine target for elicitation of broadly neutralizing antibodies (bNAbs) developed during the broadly neutralizing antibodies (bNAbs) by vaccination. However, course of natural HIV-1 infection (10–13). Insertion, deletion, current details regarding the quaternary structural organization of and mutations of residues in the MPER defined the functional the MPER within the native prefusion trimer [(gp120/41)3] are elu- importance of the MPER in Env incorporation, viral fusion, and sive and even contradictory, hindering rational MPER immunogen infectivity (14–16). -

Viroporins: Structures and Functions Beyond Cell Membrane Permeabilization
Editorial Viroporins: Structures and Functions beyond Cell Membrane Permeabilization José Luis Nieva 1,* and Luis Carrasco 2,* Received: 17 September 2015 ; Accepted: 21 September 2015 ; Published: 29 September 2015 Academic Editor: Eric O. Freed 1 Biophysics Unit (CSIC, UPV/EHU) and Biochemistry and Molecular Biology Department, University of the Basque Country (UPV/EHU), P.O. Box 644, 48080 Bilbao, Spain 2 Centro de Biología Molecular Severo Ochoa (CSIC, UAM), c/Nicolás Cabrera, 1, Universidad Autónoma de Madrid, Cantoblanco, 28049 Madrid, Spain * Correspondence: [email protected] (J.L.N.); [email protected] (L.C.); Tel.: +34-94-601-3353 (J.L.N.); +34-91-497-8450 (L.C.) Viroporins represent an interesting group of viral proteins that exhibit two sets of functions. First, they participate in several viral processes that are necessary for efficient production of virus progeny. Besides, viroporins interfere with a number of cellular functions, thus contributing to viral cytopathogenicity. Twenty years have elapsed from the first review on viroporins [1]; since then several reviews have covered the advances on viroporin structure and functioning [2–8]. This Special Issue updates and revises new emerging roles of viroporins, highlighting their potential use as antiviral targets and in vaccine development. Viroporin structure. Viroporins are usually short proteins with at least one hydrophobic amphipathic helix. Homo-oligomerization is achieved by helix–helix interactions in membranes rendering higher order structures, forming aqueous pores. Progress in viroporin structures during the last 2–3 years has in some instances provided a detailed knowledge of their functional architecture, including the fine definition of binding sites for effective inhibitors. -

How Influenza Virus Uses Host Cell Pathways During Uncoating
cells Review How Influenza Virus Uses Host Cell Pathways during Uncoating Etori Aguiar Moreira 1 , Yohei Yamauchi 2 and Patrick Matthias 1,3,* 1 Friedrich Miescher Institute for Biomedical Research, 4058 Basel, Switzerland; [email protected] 2 Faculty of Life Sciences, School of Cellular and Molecular Medicine, University of Bristol, Bristol BS8 1TD, UK; [email protected] 3 Faculty of Sciences, University of Basel, 4031 Basel, Switzerland * Correspondence: [email protected] Abstract: Influenza is a zoonotic respiratory disease of major public health interest due to its pan- demic potential, and a threat to animals and the human population. The influenza A virus genome consists of eight single-stranded RNA segments sequestered within a protein capsid and a lipid bilayer envelope. During host cell entry, cellular cues contribute to viral conformational changes that promote critical events such as fusion with late endosomes, capsid uncoating and viral genome release into the cytosol. In this focused review, we concisely describe the virus infection cycle and highlight the recent findings of host cell pathways and cytosolic proteins that assist influenza uncoating during host cell entry. Keywords: influenza; capsid uncoating; HDAC6; ubiquitin; EPS8; TNPO1; pandemic; M1; virus– host interaction Citation: Moreira, E.A.; Yamauchi, Y.; Matthias, P. How Influenza Virus Uses Host Cell Pathways during 1. Introduction Uncoating. Cells 2021, 10, 1722. Viruses are microscopic parasites that, unable to self-replicate, subvert a host cell https://doi.org/10.3390/ for their replication and propagation. Despite their apparent simplicity, they can cause cells10071722 severe diseases and even pose pandemic threats [1–3]. -

Assembly Lecture 11 Biology W3310/4310 Virology Spring 2014
Assembly Lecture 11 Biology W3310/4310 Virology Spring 2014 “Anatomy is des.ny.” --SIGMUND FREUD All virions complete a common set of assembly reac3ons * common to all viruses common to many viruses ©Principles of Virology, ASM Press The structure of a virus parcle determines how it is formed ©Principles of Virology, ASM Press Assembly is dependent on host cell machinery • Cellular chaperones • Transport systems • Secretory pathway • Nuclear import and export machinery Concentrang components for assembly: Nothing happens fast in dilute solu1ons • Viral components oSen visible by light microscopy (‘factories’ or ‘inclusions’) • Concentrate proteins on internal membranes (poliovirus) • Negri bodies (rabies virus) Viral proteins have ‘addresses’ built into their structure • Membrane targeYng: Signal sequences, fa\y acid modificaons • Membrane retenYon signals • Nuclear localizaYon sequences (NLS) • Nuclear export signals 414 Localiza3on of viral proteins to the nucleus CHAPTER 12 Golgi apparatus Ribosome Rough endoplasmic reticulum Plasma membrane Py(VP1) + VP2/3 Ad hexon + 5 100 kDa Nuclear envelope: Outer nuclear membrane Inner nuclear membrane Nucleus Nuclear pore complex Mitochondrion Cytoskeleton: Influenza virus NP Intermediate filament Microtubule Actin filament bundle Extracellular matrix ©Principles of Virology, ASM Press Figure 12.1 Localization of viral proteins to the nucleus. The nucleus and major membrane-bound compartments of the cytoplasm, as well as components of the cytoskeleton, are illustrated schematically and not to scale. Viral proteins destined for the nucleus are synthesized by cytoplasmic polyribosomes, as illustrated for the infl uenza virus NP protein. They engage with the cytoplasmic face of the nuclear pore complex and are translocated into the nucleus by the protein import machinery of the host cell. -

Paleovirology of 'Syncytins', Retroviral Env Genes Exapted for a Role in Placentation
Paleovirology of ‘syncytins’, retroviral env genes exapted for a role in placentation Christian Lavialle1,2, Guillaume Cornelis1,2, Anne Dupressoir1,2, Ce´cile Esnault1,2, Odile Heidmann1,2,Ce´cile Vernochet1,2 1,2 rstb.royalsocietypublishing.org and Thierry Heidmann 1UMR 8122, Unite´ des Re´trovirus Endoge`nes et E´le´ments Re´troı¨des des Eucaryotes Supe´rieurs, CNRS, Institut Gustave Roussy, 94805 Villejuif, France 2Universite´ Paris-Sud XI, 91405 Orsay, France Review The development of the emerging field of ‘paleovirology’ allows biologists to reconstruct the evolutionary history of fossil endogenous retroviral sequences Cite this article: Lavialle C, Cornelis G, integrated within the genome of living organisms and has led to the retrieval of conserved, ancient retroviral genes ‘exapted’ by ancestral hosts to fulfil Dupressoir A, Esnault C, Heidmann O, essential physiological roles, syncytin genes being undoubtedly among the Vernochet C, Heidmann T. 2013 Paleovirology most remarkable examples of such a phenomenon. Indeed, syncytins are of ‘syncytins’, retroviral env genes exapted for a ‘new’ genes encoding proteins derived from the envelope protein of endogen- role in placentation. Phil Trans R Soc B 368: ous retroviral elements that have been captured and domesticated on multiple 20120507. occasions and independently in diverse mammalian species, through a pro- http://dx.doi.org/10.1098/rstb.2012.0507 cess of convergent evolution. Knockout of syncytin genes in mice provided evidence for their absolute requirement for placenta development and embryo survival, via formation by cell–cell fusion of syncytial cell layers at One contribution of 13 to a Theme Issue the fetal–maternal interface. -

Developing a Single-Cycle Infectious System to Study an ERV-K Retroviral Envelope
Developing a Single-Cycle Infectious System to Study an ERV-K Retroviral Envelope Author: Rana Elias Akleh Persistent link: http://hdl.handle.net/2345/bc-ir:107695 This work is posted on eScholarship@BC, Boston College University Libraries. Boston College Electronic Thesis or Dissertation, 2017 Copyright is held by the author. This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License (http:// creativecommons.org/licenses/by-nc-sa/4.0). Developing a Single-Cycle Infectious System to Study an ERV-K Retroviral Envelope Rana Elias Akleh A thesis submitted to the faculty of the department of Biology in partial fulfillment of the requirements for the degree of Master of Science Boston College Morrissey College of Arts and Sciences Graduate School September 2017 © 2017 Rana Elias Akleh Developing a Single-Cycle Infectious System to Study an ERV-K Retroviral Envelope Rana Elias Akleh Advisor: Welkin Johnson, Ph.D. Endogenous Retroviruses (ERVs) are “fossilized” retroviruses of a once exogenous retrovirus located in the genome of extant vertebrates. Retroviral infection results in a provirus integration into the host genome. An infection of a germline cell could lead to the provirus potentially being inherited by the offspring of the infected individual. Once in the genome, the provirus becomes subject to evolutionary processes and can become either lost or fixed in a population, remaining as “fossils” long after the exogenous retrovirus has gone extinct23. Notably, 8% of the human genome consists of ERVs30. Human Endogenous Retrovirus Type K (HERV-K)(HML-2) family is of particular interest. HERV-K integrations are as old as 30-35 million years, endogenizing before the separation of humans and Old World Monkeys. -

Ube1a Suppresses Herpes Simplex Virus-1 Replication
viruses Article UBE1a Suppresses Herpes Simplex Virus-1 Replication Marina Ikeda 1 , Akihiro Ito 2, Yuichi Sekine 1 and Masahiro Fujimuro 1,* 1 Department of Cell Biology, Kyoto Pharmaceutical University, Kyoto 607-8412, Japan; [email protected] (M.I.); [email protected] (Y.S.) 2 Laboratory of Cell Signaling, School of Life Sciences, Tokyo University of Pharmacy and Life Sciences, Tokyo 192-0392, Japan; [email protected] * Correspondence: [email protected]; Tel.: +81-75-595-4717 Academic Editors: Magdalena Weidner-Glunde and Andrea Lipi´nska Received: 7 November 2020; Accepted: 1 December 2020; Published: 4 December 2020 Abstract: Herpes simplex virus-1 (HSV-1) is the causative agent of cold sores, keratitis, meningitis, and encephalitis. HSV-1-encoded ICP5, the major capsid protein, is essential for capsid assembly during viral replication. Ubiquitination is a post-translational modification that plays a critical role in the regulation of cellular events such as proteasomal degradation, protein trafficking, and the antiviral response and viral events such as the establishment of infection and viral replication. Ub-activating enzyme (E1, also named UBE1) is involved in the first step in the ubiquitination. However, it is still unknown whether UBE1 contributes to viral infection or the cellular antiviral response. Here, we found that UBE1a suppressed HSV-1 replication and contributed to the antiviral response. The UBE1a inhibitor PYR-41 increased HSV-1 production. Immunofluorescence analysis revealed that UBE1a highly expressing cells presented low ICP5 expression, and vice versa. UBE1a inhibition by PYR-41 and shRNA increased ICP5 expression in HSV-1-infected cells. -

Researchers Unveil New Monkey Model for HIV 2 March 2009
Researchers unveil new monkey model for HIV 2 March 2009 By altering just one gene in HIV-1, scientists have evolving genes, APOBEC3 and TRIM5, which succeeded in infecting pig-tailed macaque produce unusual classes of defensive proteins with monkeys with a human version of the virus that distinctive capabilities to fight retroviruses such as has until now been impossible to study directly in HIV. These genes, shared by humans and their animals. The new strain of HIV has already been simian forebears, have evolved mutations specific used to demonstrate one method for preventing to each species' unique history of retroviral battles. infection and, with a little tweaking, could be a In most simians, the APOBEC3 and TRIM5 valuable model for vetting vaccine candidates. proteins actually kill HIV on sight, making it impossible for researchers to study the virus in an A team of researchers led by Paul Bieniasz and animal model. Instead, they have studied HIV's Theodora Hatziioannou at The Rockefeller cousin, simian immunodeficiency virus (SIV), which University showed that two pig-tailed macaques, causes an AIDS-like disease in certain monkey given a common antiretroviral treatment one week species. But SIV shares only about half of its amino before and one week after being exposed to the acid sequence with HIV, making it a very imperfect newly engineered HIV, had no signs of infection. substitute for testing anti-HIV drugs and vaccines. "We're not saying we can save the world with Several labs have engineered hybrids called SHIVs antiretroviral pills. But this model will allow us to — SIVs that contain pieces of HIV DNA — but these start studying the best way to administer have problematic differences, too. -

Identification of Small Endogenous Viral Elements Within Host
IDENTIFICATION OF SMALL ENDOGENOUS VIRAL ELEMENTS WITHIN HOST GENOMES by Edward C. Davis, Jr. A thesis submitted in partial fulfillment of the requirements for the degree of Master of Science in Computer Science Boise State University May 2016 c 2016 Edward C. Davis, Jr. ALL RIGHTS RESERVED BOISE STATE UNIVERSITY GRADUATE COLLEGE DEFENSE COMMITTEE AND FINAL READING APPROVALS of the thesis submitted by Edward C. Davis, Jr. Thesis Title: Identification of Small Endogenous Viral Elements within Host Genomes Date of Final Oral Examination: 04 March 2016 The following individuals read and discussed the thesis submitted by student Edward C. Davis, Jr., and they evaluated his presentation and response to questions during the final oral examination. They found that the student passed the final oral examination. Timothy Andersen, Ph.D. Chair, Supervisory Committee Amit Jain, Ph.D. Member, Supervisory Committee Gregory Hampikian, Ph.D. Member, Supervisory Committee The final reading approval of the thesis was granted by Timothy Andersen, Ph.D., Chair, Supervisory Committee. The thesis was approved for the Graduate College by John R. Pelton, Ph.D., Dean of the Graduate College. Dedicated to Elaina, Arianna, and Zora. iv ACKNOWLEDGMENTS The author wishes to express gratitude to the members of the supervisory com- mittee for providing guidance and patience. v ABSTRACT A parallel string matching software architecture has been developed (incorpo- rating several algorithms) to identify small genetic sequences in large genomes. En- dogenous viral elements (EVEs) are sequences originating in the genomes of viruses that have become integrated into the chromosomes of sperm or egg cells of infected hosts, and passed to subsequent generations. -
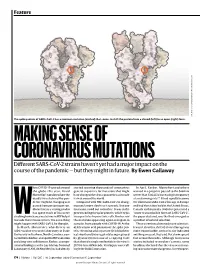
MAKING SENSE of CORONAVIRUS MUTATIONS Different SARS-Cov-2 Strains Haven’T Yet Had a Major Impact on the Course of the Pandemic — but They Might in Future
Feature SOURCE: STRUCTURAL DATA FROM K. SHEN & J. LUBAN K. SHEN FROM DATA STRUCTURAL SOURCE: The spike protein of SARS-CoV-2 has a common mutation (circled) that seems to shift the protein from a closed (left) to an open (right) form. MAKING SENSE OF CORONAVIRUS MUTATIONS Different SARS-CoV-2 strains haven’t yet had a major impact on the course of the pandemic — but they might in future. By Ewen Callaway hen COVID-19 spread around started scouring thousands of coronavirus In April, Korber, Montefiori and others the globe this year, David genetic sequences for mutations that might warned in a preprint posted to the bioRxiv Montefiori wondered how the have changed the virus’s properties as it made server that “D614G is increasing in frequency deadly virus behind the pan- its way around the world. at an alarming rate”1. It had rapidly become demic might be changing as it Compared with HIV, SARS-CoV-2 is chang- the dominant SARS-CoV-2 lineage in Europe passed from person to person. ing much more slowly as it spreads. But one and had then taken hold in the United States, Montefiori is a virologist who mutation stood out to Korber. It was in the Canada and Australia. D614G represented a has spent much of his career gene encoding the spike protein, which helps “more transmissible form of SARS-CoV-2”, Wstudying how chance mutations in HIV help it virus particles to penetrate cells. Korber saw the paper declared, one that had emerged as to evade the immune system.